

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.11 + phase: 0
(308 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
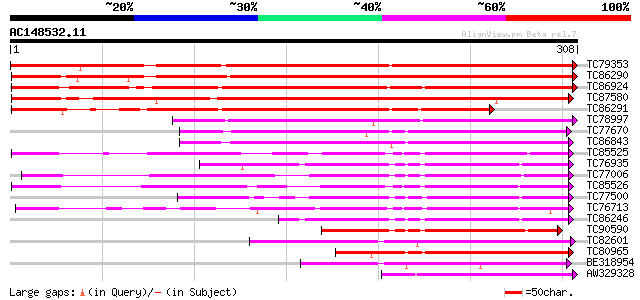
Score E
Sequences producing significant alignments: (bits) Value
TC79353 similar to GP|16588826|gb|AAL26909.1 dehydration-respons... 463 e-131
TC86290 weakly similar to GP|14422444|dbj|BAB60848. BURP domain-... 409 e-115
TC86924 weakly similar to GP|14422444|dbj|BAB60848. BURP domain-... 361 e-100
TC87580 similar to GP|18447761|gb|AAL67991.1 dehydration-induced... 314 3e-86
TC86291 weakly similar to GP|16588826|gb|AAL26909.1 dehydration-... 278 1e-75
TC78997 weakly similar to PIR|D96529|D96529 probable RAB7 GTP-bi... 174 4e-44
TC77670 similar to PIR|D96529|D96529 probable RAB7 GTP-binding P... 157 6e-39
TC86843 138 3e-33
TC85525 similar to PIR|T08896|T08896 Sali3-2 protein aluminium-... 130 5e-31
TC76935 similar to SP|P21747|EA92_VICFA Embryonic abundant prote... 124 3e-29
TC77006 similar to SP|P21745|EA30_VICFA Embryonic abundant prote... 122 1e-28
TC85526 similar to PIR|T08896|T08896 Sali3-2 protein aluminium-... 120 7e-28
TC77500 weakly similar to SP|P21747|EA92_VICFA Embryonic abundan... 119 2e-27
TC76713 similar to PIR|S33622|S33622 ADR6 protein - soybean, par... 117 4e-27
TC86246 weakly similar to PIR|S33622|S33622 ADR6 protein - soybe... 117 7e-27
TC90590 similar to GP|20914|emb|CAA38756.1|| unknown seed protei... 115 2e-26
TC82601 similar to GP|1762584|gb|AAB39546.1| polygalacturonase i... 113 8e-26
TC80965 similar to PIR|D96529|D96529 probable RAB7 GTP-binding P... 113 1e-25
BE318954 similar to PIR|T07587|T07 probable polygalacturonase (E... 105 2e-23
AW329328 similar to PIR|D96529|D965 probable RAB7 GTP-binding Pr... 100 5e-22
>TC79353 similar to GP|16588826|gb|AAL26909.1 dehydration-responsive protein
RD22 {Prunus persica}, partial (64%)
Length = 1324
Score = 463 bits (1192), Expect = e-131
Identities = 234/312 (75%), Positives = 256/312 (82%), Gaps = 4/312 (1%)
Frame = +2
Query: 1 MLPPQLYWQSMLPNSPMPKAFTNLLHPAGYWSKEKAR--DASNGGLDVGVRKGY-EGGGT 57
MLPPQLYW+SMLPNSPMPKA TNLLHPAGYWS+EK D GG+DVGVRKGY EGGGT
Sbjct: 131 MLPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGT 310
Query: 58 YLN-DEKIIPLIYFYPIPIPLNESQIQLDDKQNVTLFFLKKDLHHGTKLNLQFKETTSNN 116
+N P IY Y S+ QL DK NV LFFL+KDLHHGTKLNLQF +TTSN
Sbjct: 311 DVNVGVGRSPFIYNYAA------SETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSN- 469
Query: 117 NGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCV 176
FLPR+VANSIPFSSNKME I+NK +IK+GSK +IVK TISECE GIKGEEK+CV
Sbjct: 470 -AATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCV 646
Query: 177 TSLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPY 236
TSLESMVDFT SKLG NVEAVSTEV+K SN LQQY IA GVKKLGEKNK +VCHKENYPY
Sbjct: 647 TSLESMVDFTTSKLGKNVEAVSTEVNKESN-LQQYTIASGVKKLGEKNKAVVCHKENYPY 823
Query: 237 AVFYCHKTDSTEVYSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICH 296
AVFYCHKTD+T+ YSVPLEG DG+ VK IAVCHTDTSEWNPKHLAF VLKVQPGTVP+CH
Sbjct: 824 AVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCH 1003
Query: 297 ILPQDHVVWVSK 308
+LP+DHVVW+ K
Sbjct: 1004LLPEDHVVWIRK 1039
>TC86290 weakly similar to GP|14422444|dbj|BAB60848. BURP domain-containing
protein {Bruguiera gymnorrhiza}, partial (87%)
Length = 1302
Score = 409 bits (1052), Expect = e-115
Identities = 220/372 (59%), Positives = 252/372 (67%), Gaps = 65/372 (17%)
Frame = +2
Query: 2 LPPQLYWQSMLPNSPMPKAFTNLLHPAGYWSKEK-------------------------- 35
LPP++YW+S LP + MPKA T+LLH W++EK
Sbjct: 140 LPPEVYWKSKLPTTQMPKAITDLLHLD--WTEEKSTSVAVGKGGVNVDSGKTKPGGTSVN 313
Query: 36 ------------------ARDASNGGLDVGVRKGYEGGGTYLNDEK-------------- 63
A + GG+ V KG GGGT +N K
Sbjct: 314 VGKGGVHVDAGKTKPGGTAVNVGKGGVHVNAGKGKPGGGTRVNVGKGGVNVHAGKGRGKP 493
Query: 64 -------IIPLIYFYPIPIPLNESQIQLDDKQNVTLFFLKKDLHHGTKLNLQFKETTSNN 116
P +Y Y S+ QL DK NV LFFL+KDLH GTKLNLQF +T SN+
Sbjct: 494 VHVSVGNKSPFLYNYAA------SETQLHDKPNVALFFLEKDLHQGTKLNLQFTKTNSND 655
Query: 117 NGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCV 176
+ KFLP+EVANSIPFSS+K+ENILN FSIKEG++E+EIVK TISECE NGIKGEEKLCV
Sbjct: 656 DA-KFLPKEVANSIPFSSSKVENILNLFSIKEGTEESEIVKNTISECEENGIKGEEKLCV 832
Query: 177 TSLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPY 236
TSLESMVDFT SKLGNNVEAVSTEV K S+ LQ+YV+AKGVKKLGEKNK +VCHKE+YPY
Sbjct: 833 TSLESMVDFTTSKLGNNVEAVSTEVKKESSDLQEYVMAKGVKKLGEKNKAVVCHKESYPY 1012
Query: 237 AVFYCHKTDSTEVYSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICH 296
AVFYCHKTDST+VYSVPLEGVDG+ VK +AVCHTDTS+WNPKHLAF VL VQPGTVP+CH
Sbjct: 1013AVFYCHKTDSTKVYSVPLEGVDGSRVKAVAVCHTDTSQWNPKHLAFQVLNVQPGTVPVCH 1192
Query: 297 ILPQDHVVWVSK 308
LPQDHVVWVSK
Sbjct: 1193FLPQDHVVWVSK 1228
>TC86924 weakly similar to GP|14422444|dbj|BAB60848. BURP domain-containing
protein {Bruguiera gymnorrhiza}, partial (52%)
Length = 1202
Score = 361 bits (927), Expect = e-100
Identities = 185/307 (60%), Positives = 228/307 (74%)
Frame = +1
Query: 2 LPPQLYWQSMLPNSPMPKAFTNLLHPAGYWSKEKARDASNGGLDVGVRKGYEGGGTYLND 61
LPP+LYW+S LP + +PKA T++LHP ++ +G + V + G + D
Sbjct: 130 LPPELYWKSKLPTTQIPKAITDILHP-----RKGVTSVDDGKISVDGKISVNGKTRIVYD 294
Query: 62 EKIIPLIYFYPIPIPLNESQIQLDDKQNVTLFFLKKDLHHGTKLNLQFKETTSNNNGTKF 121
I YFY + ++ +L D +N+ +F L+KDLHHGTK N+QF +T+ ++G F
Sbjct: 295 AAPI---YFYEN----DANEAELHDNRNLAMFLLEKDLHHGTKFNIQFTKTS--DHGPTF 447
Query: 122 LPREVANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCVTSLES 181
LP +VANSIPFSSNK+ENILN FSIK+GS E+EIVK TISECEA GIKGEEKLCVTSLES
Sbjct: 448 LPGDVANSIPFSSNKLENILNYFSIKQGSTESEIVKNTISECEAYGIKGEEKLCVTSLES 627
Query: 182 MVDFTISKLGNNVEAVSTEVDKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPYAVFYC 241
M+DFT KLGNNV+ VSTEV+ S GLQQYVIA GVKK+GE N +VCHK NYPYAVFYC
Sbjct: 628 MIDFTTLKLGNNVDTVSTEVNGES-GLQQYVIANGVKKMGENN-LVVCHKRNYPYAVFYC 801
Query: 242 HKTDSTEVYSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQD 301
HKTD+T+VYSVPLEG DG+ VK +A+CH+DTS+W+ KHLAF VLKVQ GT P+CHIL Q
Sbjct: 802 HKTDATKVYSVPLEGADGSRVKAVAICHSDTSQWSLKHLAFQVLKVQQGTFPVCHILQQG 981
Query: 302 HVVWVSK 308
VVW SK
Sbjct: 982 QVVWFSK 1002
>TC87580 similar to GP|18447761|gb|AAL67991.1 dehydration-induced protein
RD22-like protein {Gossypium hirsutum}, partial (76%)
Length = 1290
Score = 314 bits (804), Expect = 3e-86
Identities = 157/310 (50%), Positives = 210/310 (67%), Gaps = 5/310 (1%)
Frame = +2
Query: 2 LPPQLYWQSMLPNSPMPKAFTNLLHPAGYWSKEKARDASNGGLDVGVRKGYEGGGTYLND 61
LPP+LYW+S+LP +PMPKA T++LHP W ++K+ + VGV KG +
Sbjct: 143 LPPELYWKSVLPTTPMPKAITDILHPG--WMEDKSTN-------VGVGKGVSVNAGHKRK 295
Query: 62 EKIIPLIYFYPIPIPLN--ESQIQLDDKQNVTLFFLKKDLHHGTKLNLQFKETTSNNNGT 119
K ++ P N ++ QL D LFFL+KDLH GTK++L F ++N
Sbjct: 296 GKPATVVVGPHSPFAYNYAATETQLHDDPRAALFFLEKDLHPGTKMDLNF---IRSSNVA 466
Query: 120 KFLPREVANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCVTSL 179
FLPR+VA++ PFSS K+ +I NKFS+K S+EA ++K TI ECE I+GEEK C TSL
Sbjct: 467 TFLPRQVASATPFSSEKLSHIFNKFSVKPESEEAHVMKNTIEECEDAAIQGEEKYCATSL 646
Query: 180 ESMVDFTISKLGNNVEAVSTE-VDKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPYAV 238
ESMVDF+ISKLG V+ VST VD S GL++Y + G+ KL ++K +VCHK+NYPYA+
Sbjct: 647 ESMVDFSISKLGKRVKVVSTTVVDNKSAGLKKYTLKSGLMKLTAEDKAVVCHKQNYPYAI 826
Query: 239 FYCHKTDSTEVYSVPLEGVDGNMVK--TIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICH 296
FYCHKT+ST Y VPLE +G VK + +CHTDTS+WNP+H+AF VLK++PGT+ +CH
Sbjct: 827 FYCHKTESTRAYFVPLEADNGTRVKADAVVICHTDTSKWNPEHIAFQVLKIKPGTLSVCH 1006
Query: 297 ILPQDHVVWV 306
LP DH+VWV
Sbjct: 1007FLPVDHIVWV 1036
>TC86291 weakly similar to GP|16588826|gb|AAL26909.1 dehydration-responsive
protein RD22 {Prunus persica}, partial (42%)
Length = 829
Score = 278 bits (712), Expect = 1e-75
Identities = 152/264 (57%), Positives = 186/264 (69%), Gaps = 2/264 (0%)
Frame = +1
Query: 2 LPPQLYWQSMLPNSPMPKAFTNLLHP--AGYWSKEKARDASNGGLDVGVRKGYEGGGTYL 59
LPP+LYW+S LP + +PKA T++LHP G W +DVG
Sbjct: 130 LPPELYWKSKLPTTQIPKAITDILHPRKGGTW------------VDVG------------ 237
Query: 60 NDEKIIPLIYFYPIPIPLNESQIQLDDKQNVTLFFLKKDLHHGTKLNLQFKETTSNNNGT 119
N ++ L++ Y P + S QL D +N+ LFF +KDLH+GTKLN+QF +T+ + G
Sbjct: 238 NAGRVQQLLFIY---WPYDASDTQLHDYRNLALFFFEKDLHNGTKLNMQFTKTS--DYGA 402
Query: 120 KFLPREVANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCVTSL 179
FLPREVANSIPFSSNK+ENILN FSIK+GS E EIVK TI CE I+GEEK CVTSL
Sbjct: 403 TFLPREVANSIPFSSNKVENILNYFSIKQGSAEYEIVKNTIGSCEMPAIEGEEKSCVTSL 582
Query: 180 ESMVDFTISKLGNNVEAVSTEVDKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPYAVF 239
ESMVDFT SKLGNNVEAVSTEV+K S+ +QQY+IAKGVKKLGE NK +VCH +YPY VF
Sbjct: 583 ESMVDFTTSKLGNNVEAVSTEVNKESD-IQQYIIAKGVKKLGE-NKIVVCHPMDYPYTVF 756
Query: 240 YCHKTDSTEVYSVPLEGVDGNMVK 263
YCHK +T+VY +P+EGVDG +K
Sbjct: 757 YCHKLRATKVYYLPMEGVDGPKIK 828
>TC78997 weakly similar to PIR|D96529|D96529 probable RAB7 GTP-binding
Protein [imported] - Arabidopsis thaliana, partial (27%)
Length = 1039
Score = 174 bits (441), Expect = 4e-44
Identities = 92/224 (41%), Positives = 134/224 (59%), Gaps = 4/224 (1%)
Frame = +2
Query: 89 NVTLFFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILNKFSIKE 148
++ +FF KDL G + + F + + + K ++ A S+PFSSN++ +L FS +
Sbjct: 164 SLMVFFTLKDLKVGNIMQIYFPKRDPSTS-PKLWSKQKAESLPFSSNQLSYLLKFFSFSQ 340
Query: 149 GSKEAEIVKRTISECEANGIKGEEKLCVTSLESMVDFTISKLGNNVEA---VSTEVDKNS 205
S +A +K T+ ECE+ IKGE KLC TS ESM++FT + LG+ E + K+S
Sbjct: 341 DSPQAMAMKNTLRECESKPIKGEVKLCATSFESMLEFTQNVLGSKYEIQGYATLHKTKSS 520
Query: 206 NGLQQYVIAKGVKKLGEKNKTIVCHKENYPYAVFYCHKTDS-TEVYSVPLEGVDGNMVKT 264
LQ Y I + +K++ K + CH YPYAVFYCH +S VY L G +G+ V+
Sbjct: 521 VTLQNYTIVEILKEILAP-KMVACHTVPYPYAVFYCHGQESDNRVYRASLVGENGDKVEA 697
Query: 265 IAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVWVSK 308
+AVCH DTS+W P H++F VL+V PGT +CH P D+ +WV K
Sbjct: 698 MAVCHMDTSQWAPSHVSFQVLEVTPGTSSVCHFFPADNYIWVPK 829
>TC77670 similar to PIR|D96529|D96529 probable RAB7 GTP-binding Protein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 1649
Score = 157 bits (396), Expect = 6e-39
Identities = 85/218 (38%), Positives = 123/218 (55%), Gaps = 5/218 (2%)
Frame = +2
Query: 93 FFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKE 152
FF DL+ G + LQF N FLP++VA+ IPFS +++ ++L FS+ + S +
Sbjct: 677 FFTMDDLYVGNVMTLQFPVREYAN----FLPKKVADYIPFSKSQLPSLLQLFSLTKDSPQ 844
Query: 153 AEIVKRTISECEANGIKGEEKLCVTSLESMVDFTISKLGN----NVEAVSTEVDKNSNGL 208
E +K I +CE KGE K C TSLESM++F S LG N+ S + L
Sbjct: 845 GEDMKDIIDQCEFEPTKGETKACPTSLESMLEFVHSVLGTEARYNIHTTSYPTTSGAR-L 1021
Query: 209 QQYVIAKGVKKLGEKNKTIVCHKENYPYAVFYCHKTD-STEVYSVPLEGVDGNMVKTIAV 267
Q Y I K + K K + CH YPYA++YCH D ++V+ V L+G G+++ + +
Sbjct: 1022QNYTILK-ISKDIYAPKWVACHPRPYPYALYYCHYLDIGSKVFKVLLKGQYGDIMDALGI 1198
Query: 268 CHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVW 305
CH DTS+ NP H F +L ++PG P+CH P HV+W
Sbjct: 1199CHLDTSDMNPNHFIFQLLGIKPGEAPMCHFFPVKHVLW 1312
>TC86843
Length = 1157
Score = 138 bits (347), Expect = 3e-33
Identities = 77/220 (35%), Positives = 125/220 (56%), Gaps = 6/220 (2%)
Frame = +1
Query: 93 FFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKE 152
FF DL+ G + LQF + FL ++ A+SIP + +++ ++L SI E S +
Sbjct: 304 FFNLDDLYVGNVMTLQFPVQVVS----PFLSKKEADSIPLAMSQLPSVLQLLSIPEDSPQ 471
Query: 153 AEIVKRTISECEANGIKGEEKLCVTSLESMVDFTISKLGNNVEAVSTEVDKNSN---GLQ 209
A+ + ++ ECE+ I GE K C SLESM++F + G++ + +K S LQ
Sbjct: 472 AKSMINSLGECESETITGETKTCANSLESMLEFVNTIFGSDAKISILTTNKPSPTAVPLQ 651
Query: 210 QYVIAKGVKKLGEKNKTIVCHKENYPYAVFYCHK-TDSTEVYSVPLEG-VDGNMVKTIAV 267
+Y I + V + K + CH +YPYA+++CH T V+ V L G +G+ ++ + +
Sbjct: 652 KYTILE-VSHDIDAPKWVACHPLSYPYAIYFCHYIATGTRVFKVSLVGDENGDKMEAVGI 828
Query: 268 CHTDTSEWNPKHLAFYVLKVQPG-TVPICHILPQDHVVWV 306
CH DTS+WNP H+ F L ++ G P+CH LP +H++WV
Sbjct: 829 CHLDTSDWNPDHIIFRQLSIKAGKNSPVCHFLPVNHLLWV 948
>TC85525 similar to PIR|T08896|T08896 Sali3-2 protein aluminium-induced -
soybean, partial (89%)
Length = 1076
Score = 130 bits (328), Expect = 5e-31
Identities = 93/308 (30%), Positives = 146/308 (47%), Gaps = 3/308 (0%)
Frame = +3
Query: 2 LPPQLYWQSMLPNSPMPKAFTNLLHPAGYWSKEKARDASNGGLDVGVRKGYEGGGTYLND 61
LP + YW+++ PN+P+P + LL P G EG
Sbjct: 117 LPEEEYWEAVWPNTPIPSSLRELLKP-----------------------GPEG------- 206
Query: 62 EKIIPLIYFYPIPIPLNESQIQLDDKQNVTLFFLKKDLHHGTKLNLQF-KETTSNNNGTK 120
+ +++ +++DD Q FF + +L+ G + +QF K + G
Sbjct: 207 -------------VEIDDLPMEVDDTQYPKTFFYEHELYPGKTMKVQFSKRPFAQPYGVY 347
Query: 121 FLPREVANSIPFSSNKMENILNKFSIKEGSKEAEI-VKRTISECEANGIKGEEKLCVTSL 179
RE+ K KEG E+ VK+ +E GE+K C SL
Sbjct: 348 TWMREI----------------KDIEKEGYTFNEVCVKKAAAE-------GEQKFCAKSL 458
Query: 180 ESMVDFTISKLGNNVEAVSTE-VDKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPYAV 238
+++ F+ISKLG N++A+S+ +DK+ +QY I + V+ LGEK ++CH+ N+ V
Sbjct: 459 GTLIGFSISKLGKNIQALSSSFIDKH----EQYKI-ESVQNLGEK--AVMCHRLNFQKVV 617
Query: 239 FYCHKTDSTEVYSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHIL 298
FYCH+ T + VPL DG + +AVCHTDTS N + L ++K PG+ P+CH L
Sbjct: 618 FYCHEIHGTTAFMVPLVANDGRKTQALAVCHTDTSGMNHEMLQ-QIMKADPGSKPVCHFL 794
Query: 299 PQDHVVWV 306
++WV
Sbjct: 795 GNKAILWV 818
>TC76935 similar to SP|P21747|EA92_VICFA Embryonic abundant protein USP92
precursor. [Broad bean] {Vicia faba}, partial (40%)
Length = 1840
Score = 124 bits (312), Expect = 3e-29
Identities = 73/205 (35%), Positives = 111/205 (53%), Gaps = 2/205 (0%)
Frame = +1
Query: 104 KLNLQFKETTSNNNGTKFLPRE--VANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTIS 161
K N F+ T N+ ++ LP +I + K + + EA I++
Sbjct: 967 KANEPFETRTWNDKTSQPLPAHKWTDKAIAKETEKTSQHQHFVTHTSDENEAHILE---D 1137
Query: 162 ECEANGIKGEEKLCVTSLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQYVIAKGVKKLG 221
C GE+K C SLESM+DF ISKLG N++ +S+ KN + QYV+ + V K+G
Sbjct: 1138 YCGRPSAIGEDKHCAPSLESMMDFAISKLGKNIKVMSSSFSKNHD---QYVVEE-VNKIG 1305
Query: 222 EKNKTIVCHKENYPYAVFYCHKTDSTEVYSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLA 281
+ ++CH+ N+ +FYCH+ ++T Y VPL DG K + +CH DT +P ++
Sbjct: 1306 DN--AVMCHRLNFKKVLFYCHQVNATTTYMVPLVASDGTKSKALTICHHDTRGMDP-NVL 1476
Query: 282 FYVLKVQPGTVPICHILPQDHVVWV 306
+ VLKV+PGTVP+CH + V WV
Sbjct: 1477 YDVLKVKPGTVPVCHFIGNKAVAWV 1551
Score = 28.1 bits (61), Expect = 4.5
Identities = 10/19 (52%), Positives = 16/19 (83%)
Frame = +1
Query: 7 YWQSMLPNSPMPKAFTNLL 25
YW+S+ PN+ MPK+ ++LL
Sbjct: 109 YWKSVWPNTLMPKSLSDLL 165
>TC77006 similar to SP|P21745|EA30_VICFA Embryonic abundant protein VF30.1
precursor. [Broad bean] {Vicia faba}, complete
Length = 1136
Score = 122 bits (307), Expect = 1e-28
Identities = 87/301 (28%), Positives = 130/301 (42%), Gaps = 1/301 (0%)
Frame = +2
Query: 7 YWQSMLPNSPMPKAFTNLLHPAGYWSKEKARDASNGGLDVGVRKGYEGGGTYLNDEKIIP 66
YWQ++ PN+P+PKAF++LL P G
Sbjct: 113 YWQTIWPNTPLPKAFSDLLLPYG------------------------------------- 181
Query: 67 LIYFYPIPIPLNESQIQLDDKQNVTLFFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREV 126
N I+ ++ + F + DL+ G K+ L + N PR+
Sbjct: 182 ---------KTNNLPIKTEELNQYSTLFFEHDLYPGKKMILGNNNSLRNTVRPFGKPRQG 334
Query: 127 ANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCVTSLESMVDFT 186
+ +NK L+ F C + GE K CV+SLESMV+
Sbjct: 335 ITDSIWLANKERQSLDDF------------------CNSPTATGERKHCVSSLESMVEHV 460
Query: 187 ISKLGNN-VEAVSTEVDKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPYAVFYCHKTD 245
IS G + ++A+S+ N + QYV+ + VKK+G+ ++CH+ N+ VF CH+
Sbjct: 461 ISHFGTSKIKAISSTFGVNQD---QYVVEE-VKKVGDN--AVMCHRLNFEKVVFNCHQVQ 622
Query: 246 STEVYSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVW 305
+T Y V L DG K + VCH DT N + L + LKV+PGT+PICH + W
Sbjct: 623 ATTAYVVSLVAPDGGKAKALTVCHHDTRGMNAE-LLYEALKVEPGTIPICHFIGNKAAAW 799
Query: 306 V 306
V
Sbjct: 800 V 802
>TC85526 similar to PIR|T08896|T08896 Sali3-2 protein aluminium-induced -
soybean, partial (89%)
Length = 1095
Score = 120 bits (301), Expect = 7e-28
Identities = 82/307 (26%), Positives = 137/307 (43%), Gaps = 2/307 (0%)
Frame = +2
Query: 2 LPPQLYWQSMLPNSPMPKAFTNLLHPAGYWSKEKARDASNGGLDVGVRKGYEGGGTYLND 61
+P + YW+++ PN+P+P + LL P
Sbjct: 113 VPEEEYWEAVWPNTPIPTSLLELLKPG--------------------------------- 193
Query: 62 EKIIPLIYFYPIPIPLNESQIQLDDKQNVTLFFLKKDLHHGTKLNLQF-KETTSNNNGTK 120
P + +++ ++DD Q T FF + +L+ G +N+QF K + G
Sbjct: 194 ----------PKGVEIDDLPTEIDDTQFPTNFFYEHELYPGKTMNMQFSKRPLAQPYGVY 343
Query: 121 FLPREVANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCVTSLE 180
F ++ + K +++ +K K K EEK C SL
Sbjct: 344 FWMHDIKDL-----QKEGYTIDEMCVKNKPK-----------------KVEEKFCAKSLG 457
Query: 181 SMVDFTISKLGNNVEAVSTE-VDKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPYAVF 239
+++ F ISKLG N++++S+ +DK+ +QY I + V+ LG+K ++CH+ N+ VF
Sbjct: 458 TLIGFAISKLGKNIQSLSSSFIDKH----EQYKI-ESVQNLGDK--AVMCHRLNFQKVVF 616
Query: 240 YCHKTDSTEVYSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILP 299
YCH+ T + VPL DG IA CH D S N +H+ ++K PG+ +CH L
Sbjct: 617 YCHEVHGTTAFKVPLVANDGTKTHAIATCHADISGMN-QHMLHQIMKGDPGSNHVCHFLG 793
Query: 300 QDHVVWV 306
++WV
Sbjct: 794 NKAILWV 814
>TC77500 weakly similar to SP|P21747|EA92_VICFA Embryonic abundant protein
USP92 precursor. [Broad bean] {Vicia faba}, partial
(40%)
Length = 1128
Score = 119 bits (297), Expect = 2e-27
Identities = 80/216 (37%), Positives = 113/216 (52%), Gaps = 1/216 (0%)
Frame = +3
Query: 92 LFFLKKDLHHGTKLN-LQFKETTSNNNGTKFLPREVANSIPFSSNKMENILNKFSIKEGS 150
L FL+ DL+ G K++ LQ +LP E A S N +E KE
Sbjct: 120 LLFLENDLYPGKKMSGLQKLSNIQPFRTIGWLPIEKATQQ--SGNVVE--------KESY 269
Query: 151 KEAEIVKRTISECEANGIKGEEKLCVTSLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQ 210
EI C GE+K C TS ESM +F ISKLG +++ S KN + Q
Sbjct: 270 SLDEI-------CGGPPAIGEDKFCATSSESMKEFAISKLGAKIKSYSGYFAKNQD---Q 419
Query: 211 YVIAKGVKKLGEKNKTIVCHKENYPYAVFYCHKTDSTEVYSVPLEGVDGNMVKTIAVCHT 270
YV+ + V+K+ +K ++CH+ N+ VFYCH+ +++ Y VPL DG+ V +A CH
Sbjct: 420 YVVEE-VRKIADKG--VMCHRMNFEKVVFYCHQVNASTTYMVPLLASDGSKVNALAACHH 590
Query: 271 DTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVWV 306
DT NP +L VLKV+PGT+P+CH + V W+
Sbjct: 591 DTRGMNP-NLLDEVLKVKPGTIPVCHFVGNKAVAWL 695
>TC76713 similar to PIR|S33622|S33622 ADR6 protein - soybean, partial (67%)
Length = 1790
Score = 117 bits (294), Expect = 4e-27
Identities = 88/307 (28%), Positives = 140/307 (44%), Gaps = 4/307 (1%)
Frame = +2
Query: 4 PQLYWQSMLPNSPMPKAFTNLLHPAGYWSKEKARDASNGGLDVGVRKGYEGGGTYLNDEK 63
PQ YW+++ PN+P+P A +LL P E G + D
Sbjct: 767 PQKYWEAVWPNTPIPIAMQDLLKP-------------------------EHEGDEIED-- 865
Query: 64 IIPLIYFYPIPIPLNESQIQLDDKQNVTLFFLKKDLHHGTKLNLQFKETTSNNNGTKFLP 123
+P+ + Q D F + +L+ G +NLQF ++
Sbjct: 866 ---------LPMEFEKQQYPED-------LFFQNELYPGNTMNLQFDKS----------- 964
Query: 124 REVANSIPFS--SNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCVTSLES 181
PF+ + ++ + IK KEA V + A GE+K C SL S
Sbjct: 965 -------PFAQPAGLIKYLGANDDIKNTEKEAYNVDELCVKKRAT--IGEQKHCAKSLGS 1117
Query: 182 MVDFTISKLGNNVEAVSTEVDKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPYAVFYC 241
+++F ISKLGNN++A+S+ + N +QY + + V+ LG+K ++CH+ N+ V+YC
Sbjct: 1118 LIEFAISKLGNNIQALSSSLIDNQ---EQYTV-ESVQNLGDKG--VMCHRLNFQKVVYYC 1279
Query: 242 HKTDSTEVYSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTV--PICHILP 299
H+ +T + VPL DG + IAVCH DTS N + + LK+ + P+CH L
Sbjct: 1280 HRIRATTAFMVPLVAGDGTKTQAIAVCHADTSGMN-QQMLHDALKLDSADIKYPVCHFLG 1456
Query: 300 QDHVVWV 306
++WV
Sbjct: 1457 NKAIMWV 1477
>TC86246 weakly similar to PIR|S33622|S33622 ADR6 protein - soybean, partial
(56%)
Length = 1324
Score = 117 bits (292), Expect = 7e-27
Identities = 62/160 (38%), Positives = 94/160 (58%)
Frame = +2
Query: 147 KEGSKEAEIVKRTISECEANGIKGEEKLCVTSLESMVDFTISKLGNNVEAVSTEVDKNSN 206
K KE I+ S C GEEK C SLESM+DF ISKLG N++ +S+ ++ +
Sbjct: 629 KLDEKETHILH---SYCGNPSAIGEEKHCAYSLESMMDFAISKLGKNIKVMSSSFSQSQD 799
Query: 207 GLQQYVIAKGVKKLGEKNKTIVCHKENYPYAVFYCHKTDSTEVYSVPLEGVDGNMVKTIA 266
QY++ + V+K+G+ ++CH+ N FYCH+ ++T Y VPL DG K +
Sbjct: 800 ---QYMVEE-VRKIGDN--AVMCHRMNLKKVGFYCHQINATTTYMVPLVASDGTKSKALT 961
Query: 267 VCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVWV 306
+CH DT +P ++ + VL+V+PGTVP+CH + + WV
Sbjct: 962 ICHHDTRGMDP-NMLYEVLQVKPGTVPVCHFIGNKAIAWV 1078
Score = 31.6 bits (70), Expect = 0.40
Identities = 11/21 (52%), Positives = 17/21 (80%)
Frame = +2
Query: 7 YWQSMLPNSPMPKAFTNLLHP 27
YW+S+ PN+PMP+ ++LL P
Sbjct: 110 YWKSVWPNTPMPEVLSDLLLP 172
>TC90590 similar to GP|20914|emb|CAA38756.1|| unknown seed protein {Pisum
sativum}, partial (44%)
Length = 675
Score = 115 bits (289), Expect = 2e-26
Identities = 59/131 (45%), Positives = 86/131 (65%)
Frame = +1
Query: 170 GEEKLCVTSLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQYVIAKGVKKLGEKNKTIVC 229
GEEK C SL+SM++F ISKLG N++ +S+ +N + QY + + VKK+G+K ++C
Sbjct: 238 GEEKYCALSLKSMMNFAISKLGTNIKVISSSFAQNQD---QYKVDE-VKKIGDK--AVMC 399
Query: 230 HKENYPYAVFYCHKTDSTEVYSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQP 289
H+ N+ VFYCH+ ++T VY V L DG K + +CH DT NP L + +LKV+P
Sbjct: 400 HRLNFKDMVFYCHQVNATTVYMVLLVASDGTKAKALTICHHDTRGMNPDVL-YELLKVKP 576
Query: 290 GTVPICHILPQ 300
GT+PICH + Q
Sbjct: 577 GTIPICHFVWQ 609
>TC82601 similar to GP|1762584|gb|AAB39546.1| polygalacturonase isoenzyme 1
beta subunit homolog {Arabidopsis thaliana}, partial
(32%)
Length = 1072
Score = 113 bits (283), Expect = 8e-26
Identities = 62/181 (34%), Positives = 97/181 (53%), Gaps = 4/181 (2%)
Frame = +2
Query: 131 PFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCVTSLESMVDFTISKL 190
P S+K+ + F + E S +++ +SECE GE K CV SLE M+DF S L
Sbjct: 281 PLHSSKLNEMRQVFKVSENSSMNKMMVDALSECERAPSVGETKRCVGSLEDMIDFATSVL 460
Query: 191 GNNVEAVSTEVDKNSNGLQQYVIAKGVKKL--GEKNKTIVCHKENYPYAVFYCHKTDSTE 248
G++V STE N NG + V+ VK + G+ +++ CH+ +PY ++YCH +
Sbjct: 461 GHSVTVRSTE---NVNGAGKDVMVGKVKGINGGKVTESVSCHQSLFPYLLYYCHSVPNVH 631
Query: 249 VYSVP-LEGVDGNMVK-TIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVWV 306
VY L+ V + +A+CH DT+ W+ H AF L PG + +CH + ++ + W
Sbjct: 632 VYEADLLDPVSKAKINHGVAICHLDTTAWSSTHGAFMALGSAPGQIEVCHWIFENDMTWS 811
Query: 307 S 307
S
Sbjct: 812 S 814
>TC80965 similar to PIR|D96529|D96529 probable RAB7 GTP-binding Protein
[imported] - Arabidopsis thaliana, partial (14%)
Length = 533
Score = 113 bits (282), Expect = 1e-25
Identities = 58/133 (43%), Positives = 82/133 (61%), Gaps = 4/133 (3%)
Frame = +1
Query: 178 SLESMVDFTISKLGNNVE---AVSTEVDKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENY 234
SLES+ DF G N + +T V K+S LQ Y I+K V ++ N + CH Y
Sbjct: 7 SLESLFDFANYMFGPNSKFKVLTTTHVTKSSIPLQNYTISK-VNEISVPN-AVGCHPMPY 180
Query: 235 PYAVFYCH-KTDSTEVYSVPLEGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVP 293
P+AVFYCH + T +Y + +EG +G +V+ A+CH DTS+W+ H+AF VL V+PG P
Sbjct: 181 PFAVFYCHSQKGDTSLYEIVVEGENGGIVQAAAICHMDTSKWDADHVAFRVLNVKPGNSP 360
Query: 294 ICHILPQDHVVWV 306
+CH P D++VWV
Sbjct: 361 VCHFSPPDNLVWV 399
>BE318954 similar to PIR|T07587|T07 probable polygalacturonase (EC 3.2.1.15)
1 - tomato, partial (23%)
Length = 556
Score = 105 bits (263), Expect = 2e-23
Identities = 53/152 (34%), Positives = 83/152 (53%), Gaps = 5/152 (3%)
Frame = +2
Query: 159 TISECEANGIKGEEKLCVTSLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQYVIA---K 215
++ ECE +GE K CV S+E M+DF S LG NV +TE N NG ++ ++
Sbjct: 8 SLGECERVPARGETKKCVGSIEDMIDFAASVLGRNVVVRTTE---NVNGSKKDLMVGHVS 178
Query: 216 GVKKLGEKNKTIVCHKENYPYAVFYCHKTDSTEVYSVPL--EGVDGNMVKTIAVCHTDTS 273
G+ G+ +T+ CH+ + Y ++YCH VY L + + + +AVCH DTS
Sbjct: 179 GINNGGKMTRTVSCHQSLFQYLLYYCHSVPKVRVYQAELLDPKIKDKINQGVAVCHLDTS 358
Query: 274 EWNPKHLAFYVLKVQPGTVPICHILPQDHVVW 305
+W+P H AF L PG + +CH + ++ + W
Sbjct: 359 DWSPTHGAFVSLGSGPGQIEVCHWIFENDMSW 454
>AW329328 similar to PIR|D96529|D965 probable RAB7 GTP-binding Protein
[imported] - Arabidopsis thaliana, partial (6%)
Length = 591
Score = 100 bits (250), Expect = 5e-22
Identities = 47/107 (43%), Positives = 65/107 (59%), Gaps = 1/107 (0%)
Frame = +1
Query: 203 KNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPYAVFYCHKTDS-TEVYSVPLEGVDGNM 261
K+S Q Y I + + ++ K + CH YPYAVFYCH +S VY V L G +G+
Sbjct: 49 KSSVTFQNYTIVEIMMEI-RAPKMVACHTVPYPYAVFYCHSQESENRVYKVLLGGENGDK 225
Query: 262 VKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVWVSK 308
V+ + VCH DTS+W P H++F VL V PG+ +CH P D+ +WV K
Sbjct: 226 VEAMVVCHLDTSQWAPSHVSFQVLGVTPGSSSVCHFFPADNYIWVPK 366
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,379,887
Number of Sequences: 36976
Number of extensions: 144442
Number of successful extensions: 714
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 632
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 658
length of query: 308
length of database: 9,014,727
effective HSP length: 96
effective length of query: 212
effective length of database: 5,465,031
effective search space: 1158586572
effective search space used: 1158586572
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148532.11


