

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
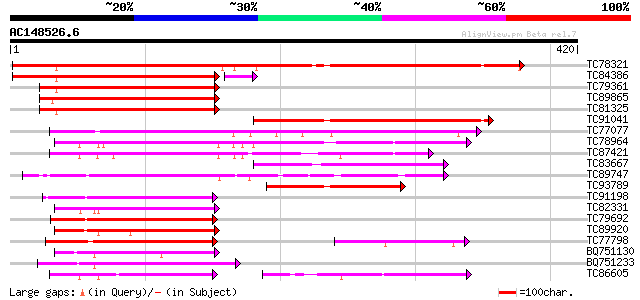
Query= AC148526.6 + phase: 0 /pseudo
(420 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC78321 similar to GP|21689841|gb|AAM67564.1 unknown protein {Ar... 357 6e-99
TC84386 similar to GP|21689841|gb|AAM67564.1 unknown protein {Ar... 192 2e-50
TC79361 similar to GP|17380890|gb|AAL36257.1 putative membrane t... 179 1e-45
TC89865 similar to GP|17380890|gb|AAL36257.1 putative membrane t... 176 1e-44
TC81325 similar to GP|17380890|gb|AAL36257.1 putative membrane t... 169 2e-42
TC91041 similar to GP|21689841|gb|AAM67564.1 unknown protein {Ar... 165 3e-41
TC77077 similar to GP|9293909|dbj|BAB01812.1 sugar transporter p... 125 2e-29
TC78964 weakly similar to PIR|T00450|T00450 probable monosacchar... 124 5e-29
TC87421 sugar transporter 107 1e-23
TC83667 similar to GP|17380890|gb|AAL36257.1 putative membrane t... 104 7e-23
TC89747 similar to PIR|F96696|F96696 protein F1N21.12 [imported]... 100 1e-21
TC93789 similar to GP|21689841|gb|AAM67564.1 unknown protein {Ar... 95 4e-20
TC91198 similar to GP|16648753|gb|AAL25568.1 AT5g16150/T21H19_70... 95 4e-20
TC82331 weakly similar to EGAD|120604|128852 hypothetical protei... 93 2e-19
TC79692 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_14... 92 5e-19
TC89920 similar to GP|12039393|gb|AAG46179.1 putative sugar tran... 88 7e-18
TC77798 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_14... 80 1e-15
BQ751130 weakly similar to GP|15289796|dbj putative monosacchari... 77 2e-14
BQ751233 similar to GP|18307448|em probable low-affinity hexose ... 76 2e-14
TC86605 similar to PIR|T01853|T01853 probable hexose transport p... 75 5e-14
>TC78321 similar to GP|21689841|gb|AAM67564.1 unknown protein {Arabidopsis
thaliana}, partial (80%)
Length = 2093
Score = 357 bits (915), Expect = 6e-99
Identities = 206/415 (49%), Positives = 270/415 (64%), Gaps = 36/415 (8%)
Frame = +2
Query: 3 EGGHPLADKTEFTECWRRTAESPYLMRLALS---GGLLFGYDTGVISGASLYIRDEFEQV 59
EGG P AD + F EC + ++PY++RLA S GGLLFGYDTGVISGA LYIRDEF V
Sbjct: 149 EGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDEFPAV 328
Query: 60 DKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWV 119
+KK WLQE IVS A AGAIIGAA GG++ND+ GRK +I++AD +F+ G++++AAAP P
Sbjct: 329 EKKTWLQEAIVSTAIAGAIIGAAIGGWINDRFGRKVSIIVADTLFLLGSIILAAAPNPAT 508
Query: 120 IIIGRLLVGLGVGAASMTAPLYISEASPAKIRGALVC--------------RLHQWFTHN 165
+I+GR+ VGLGVG ASM +PLYISEASP ++RGALV ++ FT
Sbjct: 509 LIVGRVFVGLGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTKA 688
Query: 166 ----RWSISLLPHQFGIYQV------------YRQSKEEEAKQILSKIYRPGEVEEEMKA 209
RW + + I V YR+ KEEEAK IL KIY + + E++A
Sbjct: 689 PGTWRWMLGVAAAPAVIQIVLMLSLPESPRWLYRKGKEEEAKVILKKIYEVEDYDNEIQA 868
Query: 210 MHESIEAEKAEDGLIGHSLAQKLKGAWSNDVVRRGLYAGITVQVVQQIVGINTIMYYSPT 269
+ ES+E E E I S+ Q +K VRRGLYAG+ + QQ GINT+MYYSP+
Sbjct: 869 LKESVEMELKETEKI--SIMQLVK----TTSVRRGLYAGVGLAFFQQFTGINTVMYYSPS 1030
Query: 270 IVQFAGIASNSTAFALSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSV 329
IVQ AG AS TA LSL+TSGLNA G+I+S+ ID+ GR+KL LISL G+ +SL L+V
Sbjct: 1031IVQLAGFASKRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLTGVVLSLTLLTV 1210
Query: 330 TFNQAAHHAPSLSIQDSLSFGGNSTCKAY-TTAPNHLSWNCMQCLHED--CAFCA 381
TF ++ HAP +S+ +S + N+TC + T A N+ +WNCM+C+ + C FCA
Sbjct: 1211TFRESEIHAPMVSLVESSHY--NNTCPEFKTAATNNHNWNCMKCIRAESTCGFCA 1369
>TC84386 similar to GP|21689841|gb|AAM67564.1 unknown protein {Arabidopsis
thaliana}, partial (30%)
Length = 664
Score = 192 bits (489), Expect(2) = 2e-50
Identities = 96/156 (61%), Positives = 123/156 (78%), Gaps = 3/156 (1%)
Frame = +2
Query: 3 EGGHPLADKTEFTECWRRTAESPYLMRLALS---GGLLFGYDTGVISGASLYIRDEFEQV 59
EGG P AD + F EC + ++PY++RLA S GG LFGYDTGVISGA LYIRD+F+ V
Sbjct: 113 EGGVPEADVSAFRECLSLSWKNPYVLRLAFSAGIGGFLFGYDTGVISGALLYIRDDFKAV 292
Query: 60 DKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWV 119
D++ WLQE IVS A AGAIIGA+ GG++ND+ GRKK I++AD +F G+++MAAA P +
Sbjct: 293 DRQTWLQEAIVSTALAGAIIGASVGGWINDRFGRKKAIILADALFFIGSVIMAAAINPAI 472
Query: 120 IIIGRLLVGLGVGAASMTAPLYISEASPAKIRGALV 155
+I+GR+ VGLGVG ASM +PLYISEASP ++RGALV
Sbjct: 473 LIVGRVFVGLGVGMASMASPLYISEASPTRVRGALV 580
Score = 24.6 bits (52), Expect(2) = 2e-50
Identities = 8/24 (33%), Positives = 14/24 (58%)
Frame = +1
Query: 160 QWFTHNRWSISLLPHQFGIYQVYR 183
QW +H+ W+ L +Q G ++ R
Sbjct: 586 QWISHHWWTXPFLCYQLGFHKCTR 657
>TC79361 similar to GP|17380890|gb|AAL36257.1 putative membrane transporter
protein {Arabidopsis thaliana}, partial (55%)
Length = 949
Score = 179 bits (455), Expect = 1e-45
Identities = 90/136 (66%), Positives = 114/136 (83%), Gaps = 3/136 (2%)
Frame = +1
Query: 23 ESPYLMRLALS---GGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAII 79
++PY++ LA GGLLFGYDTGVISGA LYI+D+F QV +LQETIVSMA AGAI+
Sbjct: 157 KNPYILGLAAVAGIGGLLFGYDTGVISGALLYIKDDFPQVRNSNFLQETIVSMAIAGAIV 336
Query: 80 GAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAP 139
GAAFGG++ND GRKK L+ADV+F+ GA++MAAAP P+V+I GRLLVGLGVG AS+TAP
Sbjct: 337 GAAFGGWLNDAYGRKKATLLADVIFILGAILMAAAPDPYVLIAGRLLVGLGVGIASVTAP 516
Query: 140 LYISEASPAKIRGALV 155
+YI+E +P++IRG+LV
Sbjct: 517 VYIAEVAPSEIRGSLV 564
>TC89865 similar to GP|17380890|gb|AAL36257.1 putative membrane transporter
protein {Arabidopsis thaliana}, partial (33%)
Length = 617
Score = 176 bits (447), Expect = 1e-44
Identities = 88/136 (64%), Positives = 114/136 (83%), Gaps = 3/136 (2%)
Frame = +1
Query: 23 ESPYLMRL---ALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAII 79
++PY++ L A GGLLFGYDTGVISGA LYI+D+F+ V +LQETIVSMA AGAI+
Sbjct: 79 KNPYILCLTSVASIGGLLFGYDTGVISGALLYIKDDFQAVRYSHFLQETIVSMAVAGAIV 258
Query: 80 GAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAP 139
GAA GG+MND+ GRKK ++ADV+F+ GA+VMAAAP P+++I+GR+LVGLGVG AS+TAP
Sbjct: 259 GAAVGGWMNDRYGRKKATIIADVIFILGAIVMAAAPDPYILILGRVLVGLGVGIASVTAP 438
Query: 140 LYISEASPAKIRGALV 155
+YI+E SP++IRG LV
Sbjct: 439 VYIAELSPSEIRGGLV 486
>TC81325 similar to GP|17380890|gb|AAL36257.1 putative membrane transporter
protein {Arabidopsis thaliana}, partial (32%)
Length = 636
Score = 169 bits (428), Expect = 2e-42
Identities = 85/136 (62%), Positives = 111/136 (81%), Gaps = 3/136 (2%)
Frame = +3
Query: 23 ESPYLMRLALS---GGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAII 79
++PY++ + + GGLLFGYDTGVISGA LYI+D+F+ V +LQETIVSMA GAII
Sbjct: 156 KNPYILGVTAAAGIGGLLFGYDTGVISGALLYIKDDFDDVRNSSFLQETIVSMALVGAII 335
Query: 80 GAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAP 139
GAA GG++ND GRKK L ADVVF G++VMA+AP +V+I+GRLLVG+GVG AS+TAP
Sbjct: 336 GAATGGWINDAFGRKKATLSADVVFTLGSVVMASAPDAYVLILGRLLVGIGVGVASVTAP 515
Query: 140 LYISEASPAKIRGALV 155
+YI+E+SP++IRG+LV
Sbjct: 516 VYIAESSPSEIRGSLV 563
>TC91041 similar to GP|21689841|gb|AAM67564.1 unknown protein {Arabidopsis
thaliana}, partial (31%)
Length = 564
Score = 165 bits (418), Expect = 3e-41
Identities = 89/179 (49%), Positives = 123/179 (67%), Gaps = 1/179 (0%)
Frame = +3
Query: 181 VYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAE-KAEDGLIGHSLAQKLKGAWSND 239
++R+ KEEEAK +L +IY P E E E+ + ES+E E K + S+ + LK
Sbjct: 45 LFRKGKEEEAKALLRRIYSPEEAEAEINTLKESVELEIKESESSDKASIIKMLK----TK 212
Query: 240 VVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIV 299
VRRGLYAG+ +Q+ QQ VGINT+MYYSP I+Q AG ASN TA LSLVTSGLNA G+I+
Sbjct: 213 TVRRGLYAGMGLQIFQQFVGINTVMYYSPAIIQLAGFASNQTALLLSLVTSGLNAFGSIL 392
Query: 300 SMVLIDRFGRRKLMLISLIGIFVSLVTLSVTFNQAAHHAPSLSIQDSLSFGGNSTCKAY 358
S+ ID+ GR+KL+L SL G+ +SLV L+V F++++ H+P +S ++ F N+TC Y
Sbjct: 393 SIYFIDKTGRKKLLLFSLSGVVLSLVVLTVVFHESSTHSPMISAIETSHF--NNTCTDY 563
>TC77077 similar to GP|9293909|dbj|BAB01812.1 sugar transporter protein
{Arabidopsis thaliana}, partial (85%)
Length = 1752
Score = 125 bits (315), Expect = 2e-29
Identities = 98/366 (26%), Positives = 173/366 (46%), Gaps = 46/366 (12%)
Frame = +1
Query: 30 LALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMND 89
LA +L GYD GV+SGA++YI+ + + D K E ++ + + ++IG+ G +D
Sbjct: 127 LASMTSILLGYDIGVMSGAAIYIKRDLKVSDGKI---EVLLGIINIYSLIGSCLAGRTSD 297
Query: 90 KMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAK 149
+GR+ TI+ A +F GAL+M +P ++ GR + G+G+G A M AP+Y +E SPA
Sbjct: 298 WIGRRYTIVFAGAIFFVGALLMGFSPNYNFLMFGRFVAGVGIGYALMIAPVYTAEVSPAS 477
Query: 150 IRGALVCRLHQWFTH-------NRWSISLLPHQFG-------------IYQVYRQSKEEE 189
RG L + + ++ S L + G I V + E
Sbjct: 478 SRGFLTSFPEVFINSGILLGYISNYAFSKLSLKLGWRMMLGVGAIPSVILAVGVLAMPES 657
Query: 190 AKQILSK---------IYRPGEVEEEMKAMHESIE-----AEKAEDGLIGHSLAQKLKGA 235
+ ++ + + + + +EE + I+ E D ++ + +G
Sbjct: 658 PRWLVMRGRLGDAIKVLNKTSDSKEEAQLRLAEIKQAAGIPEDCNDDVVEVKVKNTGEGV 837
Query: 236 WS------NDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVT 289
W VR + A + + QQ G++ ++ YSPTI + AGI ++ ++
Sbjct: 838 WKELFLYPTPAVRHIVIAALGIHFFQQASGVDAVVLYSPTIFKKAGINGDTHLLIATIAV 1017
Query: 290 SGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSVTF------NQAAHHAPSLSI 343
+ + +V+ ++DR+GRR L+L S+ G+ VSL+TL+V+ N + A LSI
Sbjct: 1018GFVKTLFILVATFMLDRYGRRPLLLTSVGGMVVSLLTLAVSLTIIDHSNTKLNWAIGLSI 1197
Query: 344 QDSLSF 349
LS+
Sbjct: 1198ATVLSY 1215
>TC78964 weakly similar to PIR|T00450|T00450 probable monosaccharide
transport protein T14N5.7 - Arabidopsis thaliana,
partial (98%)
Length = 1771
Score = 124 bits (312), Expect = 5e-29
Identities = 101/357 (28%), Positives = 163/357 (45%), Gaps = 48/357 (13%)
Frame = +1
Query: 34 GGLLFGYDTGVISGASL---YIRDEFEQVDKKPW--LQET------------IVSMASAG 76
GG LFGYD GV G + ++ F V +K L+ET S
Sbjct: 136 GGSLFGYDLGVSGGVTSMDDFLEKFFPDVYRKKHAHLKETDYCKYDNQVLTLFTSSLYFS 315
Query: 77 AIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASM 136
A++ F Y+ GRK TI++ + F+ GA++ AAA +IIGR+ +G G+G +
Sbjct: 316 ALVMTFFASYLTRNKGRKATIIVGALSFLIGAILNAAAQNIPTLIIGRVFLGGGIGFGNQ 495
Query: 137 TAPLYISEASPAKIRGA-------------LVCRLHQWFTH----NRWSISL-------- 171
PLY+SE +PA RGA L+ L +FT + W ISL
Sbjct: 496 AVPLYLSEMAPASSRGAVNQLFQFTTCAGILIANLVNYFTDKIHPHGWRISLGLAGIPAV 675
Query: 172 LPHQFGIY------QVYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGLIG 225
L GI+ + Q + +EA+++L K+ V+ E + + ++ E +A
Sbjct: 676 LMLLGGIFCAETPNSLVEQGRLDEARKVLEKVRGTKNVDAEFEDLKDASELAQAVKSPFK 855
Query: 226 HSLAQKLKGAWSNDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFAL 285
L +K + + + + QQ+ G N+I++Y+P I Q +G SN+ F+
Sbjct: 856 VLLKRKYR--------PQLIIGALGTPAFQQLTGNNSILFYAPVIFQSSGFGSNAALFS- 1008
Query: 286 SLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSVTFNQAAHHAPSLS 342
S +T+G V T++SM L+D+FG RK L + + ++ +V H LS
Sbjct: 1009SFITNGALLVATVISMFLVDKFGTRKFFLEAGFEMICCMIITAVVLAVEFGHGKELS 1179
>TC87421 sugar transporter
Length = 1753
Score = 107 bits (266), Expect = 1e-23
Identities = 87/339 (25%), Positives = 153/339 (44%), Gaps = 54/339 (15%)
Frame = +2
Query: 30 LALSGGLLFGYDTGVISGASL---YIRDEFEQVDKKP-----------WLQETIVSMASA 75
+A GGL+FGYD G+ G + +++ F V +K + +T+ S+
Sbjct: 113 VAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQYCQYDSQTLTMFTSS 292
Query: 76 ---GAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVG 132
A++ + + + GRK ++L ++F+ GAL+ A W++I+GR+L+G G+G
Sbjct: 293 LYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGRILLGFGIG 472
Query: 133 AASMTAPLYISEASPAKIRGA-------------LVCRLHQWFTHN-----RWSISL--- 171
A+ PLY+SE +P K RGA LV + +F W +SL
Sbjct: 473 FANQPVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWGWRLSLGGA 652
Query: 172 -------------LPHQFGIYQVYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEK 218
LP + + + AK L +I +V+EE + + EA
Sbjct: 653 MVPALIITIGSLVLPDTPN--SMIERGDRDGAKAQLKRIRGIEDVDEEFNDLVAASEA-- 820
Query: 219 AEDGLIGHSLAQKLKGAWSNDVVRR---GLYAGITVQVVQQIVGINTIMYYSPTIVQFAG 275
+ +++ W N + R+ L + + QQ GIN IM+Y+P + G
Sbjct: 821 ----------SMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLFNSIG 970
Query: 276 IASNSTAFALSLVTSGLNAVGTIVSMVLIDRFGRRKLML 314
+++ + +++T +N V T VS+ +D++GRR L L
Sbjct: 971 FKDDASLMS-AVITGVVNVVATCVSIYGVDKWGRRALFL 1084
>TC83667 similar to GP|17380890|gb|AAL36257.1 putative membrane transporter
protein {Arabidopsis thaliana}, partial (26%)
Length = 454
Score = 104 bits (259), Expect = 7e-23
Identities = 57/145 (39%), Positives = 84/145 (57%)
Frame = +3
Query: 181 VYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGLIGHSLAQKLKGAWSNDV 240
+Y ++E EA +L KIY +E+E+ + E ++ + + + + + +
Sbjct: 33 LYINNRENEAIIVLGKIYDFDRLEDEVALLTAQSEQDRQKRADV------RYRHVFKSKE 194
Query: 241 VRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVS 300
+R AG +Q QQ GINT+MYYSPTIVQ AG SN A LSL+ +GLNA GT++
Sbjct: 195 IRLAFLAGAGLQAFQQFTGINTVMYYSPTIVQMAGFHSNELALQLSLIVAGLNAAGTVLG 374
Query: 301 MVLIDRFGRRKLMLISLIGIFVSLV 325
+ LID GR+KL L SL G+ SL+
Sbjct: 375 IYLIDHAGRKKLALYSLGGVIASLI 449
>TC89747 similar to PIR|F96696|F96696 protein F1N21.12 [imported] -
Arabidopsis thaliana, partial (64%)
Length = 1372
Score = 100 bits (248), Expect = 1e-21
Identities = 85/347 (24%), Positives = 155/347 (44%), Gaps = 31/347 (8%)
Frame = +3
Query: 10 DKTEFTECWRRTAESPYLMRLALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETI 69
DK W+ + P+++ +A LFGY GV++ I + + + +
Sbjct: 339 DKETTNPSWKLSL--PHVL-VATITSFLFGYHLGVVNEPLESISVDLG-FNGNTLAEGLV 506
Query: 70 VSMASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGL 129
VS+ GA+ G G++ D +GR++ + + + GA + AA + +++GRL VG
Sbjct: 507 VSICLGGALFGCLLSGWIADAVGRRRAFQLCALPMIIGAAMSAATNNLFGMLVGRLFVGT 686
Query: 130 GVGAASMTAPLYISEASPAKIRGAL---------------------VCRLHQWFTHNRWS 168
G+G A LY++E SPA +RG V + W+ W
Sbjct: 687 GLGLGPPVAALYVTEVSPAFVRGTYGALIQIATCFGILGSLFIGIPVKEISGWWRVCFW- 863
Query: 169 ISLLPHQF----------GIYQVYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEK 218
+S +P + +Y+Q + EA+ ++ V E AM + + ++
Sbjct: 864 VSTIPAAILALAMDFCAESPHWLYKQGRTAEAEAEFERLLG---VSEAKFAMSQLSKVDR 1034
Query: 219 AEDGLIGHSLAQKLKGAWSNDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIAS 278
ED ++ L G S V + G T+ +QQ+ GIN + Y+S T+ + AG+ S
Sbjct: 1035GED-TDTVKFSELLHGHHSKVV-----FIGSTLFALQQLSGINAVFYFSSTVFKSAGVPS 1196
Query: 279 NSTAFALSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLV 325
+ + + N G+I+S L+D+ GR+ L+ S G+ +S++
Sbjct: 1197DFANVCIGVA----NLTGSIISTGLMDKLGRKVLLFWSFFGMAISMI 1325
>TC93789 similar to GP|21689841|gb|AAM67564.1 unknown protein {Arabidopsis
thaliana}, partial (17%)
Length = 668
Score = 95.1 bits (235), Expect = 4e-20
Identities = 55/104 (52%), Positives = 69/104 (65%), Gaps = 1/104 (0%)
Frame = +3
Query: 191 KQILSKIYRPGEVEEEMKAMHESIEAE-KAEDGLIGHSLAQKLKGAWSNDVVRRGLYAGI 249
K IL KIY EV+ E++A+ ES+ E K + S+ LK VRRGLYAG+
Sbjct: 366 KAILRKIYPAEEVDAEIQALQESVAMELKEAESSEKISMMTLLK----TTSVRRGLYAGM 533
Query: 250 TVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLN 293
+Q+ QQ VGINT+MY+SPTIVQ AG ASN TA LSL+T+GLN
Sbjct: 534 GLQIFQQFVGINTVMYFSPTIVQLAGFASNQTAMLLSLITAGLN 665
>TC91198 similar to GP|16648753|gb|AAL25568.1 AT5g16150/T21H19_70
{Arabidopsis thaliana}, partial (31%)
Length = 673
Score = 95.1 bits (235), Expect = 4e-20
Identities = 54/130 (41%), Positives = 76/130 (57%)
Frame = +2
Query: 25 PYLMRLALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFG 84
PY+ +A G LFGY GV++GA Y+ + ++ + LQ IVS AGA +G+ G
Sbjct: 227 PYV-GVACLGAFLFGYHLGVVNGALEYLAKDL-RIAQNTVLQGWIVSTLLAGATVGSFTG 400
Query: 85 GYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISE 144
G + DK GR +T + + G + A A + +I+GR L G+G+G AS PLYISE
Sbjct: 401 GALADKFGRTRTFQLDAIPLAIGGFLCATAQSVQTMIVGRSLAGIGIGIASAIVPLYISE 580
Query: 145 ASPAKIRGAL 154
SP +IRGAL
Sbjct: 581 ISPTEIRGAL 610
>TC82331 weakly similar to EGAD|120604|128852 hypothetical protein M01F1.5
{Caenorhabditis elegans}, partial (10%)
Length = 848
Score = 92.8 bits (229), Expect = 2e-19
Identities = 57/133 (42%), Positives = 76/133 (56%), Gaps = 11/133 (8%)
Frame = +3
Query: 34 GGLLFGYDTGVISGASLY--IRDEFEQVDK-----KPW---LQETIVSMASAGAIIGAAF 83
GG LFGYDTG IS L+ R F + K W +Q +VS+ S G +IGA
Sbjct: 270 GGFLFGYDTGQISSMLLFTDFRQRFATGPEDANGVKTWVSIIQSLMVSLMSIGTLIGALS 449
Query: 84 GGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWV-IIIGRLLVGLGVGAASMTAPLYI 142
G Y D GR+K++ VVFV G +V A WV +++GR + GLGVG S+ P++
Sbjct: 450 GAYTADWWGRRKSLSFGVVVFVIGNIVQITAMDTWVHMMMGRFIAGLGVGNLSVGVPMFQ 629
Query: 143 SEASPAKIRGALV 155
SE SP +IRGA+V
Sbjct: 630 SECSPKEIRGAVV 668
>TC79692 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_140
{Arabidopsis thaliana}, partial (49%)
Length = 1353
Score = 91.7 bits (226), Expect = 5e-19
Identities = 47/124 (37%), Positives = 78/124 (62%)
Frame = +2
Query: 31 ALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMNDK 90
A G LL G+D I+G+ LYI+ EF Q+ +P ++ IV+M+ GA + G ++D
Sbjct: 182 AAIGNLLQGWDNATIAGSILYIKREF-QLQSEPTVEGLIVAMSLIGATVVTTCSGALSDL 358
Query: 91 MGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAKI 150
GR+ ++++ +++ +LVM +P ++++ RLL GLG+G A PLYISE +P +I
Sbjct: 359 FGRRPMLIISSLLYFLSSLVMFWSPNVYILLFARLLDGLGIGLAVTLVPLYISEIAPPEI 538
Query: 151 RGAL 154
RG+L
Sbjct: 539 RGSL 550
>TC89920 similar to GP|12039393|gb|AAG46179.1 putative sugar transporter
protein {Oryza sativa}, partial (33%)
Length = 654
Score = 87.8 bits (216), Expect = 7e-18
Identities = 48/128 (37%), Positives = 78/128 (60%), Gaps = 6/128 (4%)
Frame = +2
Query: 34 GGLLFGYDTGVISGASLYIRDEFEQVDKKPW--LQETIVSMASAGAIIGAAFGGYMN--- 88
GGLLFGYD G S A++ I+ + W L + + ++G++ GA G +
Sbjct: 176 GGLLFGYDIGATSSATISIQSS--SLSGITWYDLDAVEIGLLTSGSLYGALIGSVLAFNI 349
Query: 89 -DKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASP 147
D +GR++ +L+A ++++ GAL+ A AP +++IGRL+ G+G+G A AP+YI+E +P
Sbjct: 350 ADFLGRRRELLVAALMYLVGALITAFAPNFPLLVIGRLVFGIGIGLAMHAAPMYIAETAP 529
Query: 148 AKIRGALV 155
IRG LV
Sbjct: 530 TPIRGQLV 553
>TC77798 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_140
{Arabidopsis thaliana}, partial (88%)
Length = 2467
Score = 80.1 bits (196), Expect = 1e-15
Identities = 44/129 (34%), Positives = 78/129 (60%), Gaps = 1/129 (0%)
Frame = +3
Query: 27 LMRLALS-GGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGG 85
L+ +A S G L G+D I+G+ LYI+ + + ++ +V+M+ GA + G
Sbjct: 126 LVAIAASIGNFLQGWDNATIAGSILYIKKDLAL---QTTMEGLVVAMSLIGATVITTCSG 296
Query: 86 YMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEA 145
++D +GR+ ++++ V++ G+LVM +P +V+ + RLL G G+G A P+YISE
Sbjct: 297 PISDWLGRRPMMIISSVLYFLGSLVMLWSPNVYVLCLARLLDGFGIGLAVTLVPVYISET 476
Query: 146 SPAKIRGAL 154
+P+ IRG+L
Sbjct: 477 APSDIRGSL 503
Score = 64.7 bits (156), Expect = 6e-11
Identities = 41/113 (36%), Positives = 65/113 (57%), Gaps = 13/113 (11%)
Frame = +3
Query: 241 VRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIA---------SNSTAFALSLVTSG 291
V+ L GI +Q++QQ GIN ++YY+P I++ AG+A S S++F +S VT+
Sbjct: 1623 VKHALIVGIGIQLLQQFSGINGVLYYTPQILEEAGVAVLLADLGLSSTSSSFLISAVTTL 1802
Query: 292 LNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTL----SVTFNQAAHHAPS 340
L ++M L+D GRR+L+L+++ + VSLV L + F H A S
Sbjct: 1803 LMLPSIGLAMRLMDVTGRRQLLLVTIPVLIVSLVILVLGSVIDFGSVVHAAIS 1961
>BQ751130 weakly similar to GP|15289796|dbj putative monosaccharide
transporter 3 {Oryza sativa (japonica cultivar-group)},
partial (7%)
Length = 795
Score = 76.6 bits (187), Expect = 2e-14
Identities = 50/131 (38%), Positives = 70/131 (53%), Gaps = 9/131 (6%)
Frame = +3
Query: 34 GGLLFGYDTGVISGASLYIRDEFEQVDK------KPWLQETIVSMASAGAIIGAAFGGYM 87
GG+ FGYD+G I+G L E V+ Q IVS+ S G GA G +
Sbjct: 69 GGIFFGYDSGYINGV-LGANQFIEHVEGPGAEAISESHQSLIVSILSCGTFFGALIAGDV 245
Query: 88 NDKMGRKKTILMADVVFVAGALVM--AAAPAPW-VIIIGRLLVGLGVGAASMTAPLYISE 144
+D +GRK T+++ +++ G L+ AP I+ GRL+ G+GVG S LY+SE
Sbjct: 246 SDWIGRKWTVIIGCGIYMIGVLIQMFTGMGAPLGAIVAGRLIAGIGVGFESAIVILYMSE 425
Query: 145 ASPAKIRGALV 155
P K+RGALV
Sbjct: 426 ICPKKVRGALV 458
>BQ751233 similar to GP|18307448|em probable low-affinity hexose transporter
HXT3 {Neurospora crassa}, partial (30%)
Length = 659
Score = 76.3 bits (186), Expect = 2e-14
Identities = 50/161 (31%), Positives = 82/161 (50%), Gaps = 10/161 (6%)
Frame = +3
Query: 21 TAESPYLMRLALSGGLLFGYDTGVISGASLYIRDEFEQVDK---------KPWLQETIVS 71
T + +L +A GG +FGY +G ISG L ++D ++ + Q TIV
Sbjct: 144 TFAAAFLGLVASIGGFMFGYVSGQISGFFL-MQDYLDRFGEDLPDGTRGFSAARQGTIVG 320
Query: 72 MASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWV-IIIGRLLVGLG 130
+ G ++GA G + D +GR+ TI G ++ ++ + W IGRL+ G+G
Sbjct: 321 LLCIGCLVGALTAGKLADTLGRRLTISAFAFWSCVGIVIEISSQSAWYQFAIGRLVNGVG 500
Query: 131 VGAASMTAPLYISEASPAKIRGALVCRLHQWFTHNRWSISL 171
+GA S+ P+Y SE++PA IRG +V + T W+ +
Sbjct: 501 IGALSVVVPMYQSESTPAIIRGVIVSTYQLFITLGIWTAEM 623
>TC86605 similar to PIR|T01853|T01853 probable hexose transport protein
F9D12.17 - Arabidopsis thaliana, partial (91%)
Length = 3111
Score = 75.1 bits (183), Expect = 5e-14
Identities = 47/144 (32%), Positives = 78/144 (53%), Gaps = 19/144 (13%)
Frame = +2
Query: 30 LALSGGLLFGYDTGVISGASL---YIRDEFEQVDKKPW----------------LQETIV 70
+A +GGL+FGYD GV G + +++ F V ++ LQ
Sbjct: 1370 MAATGGLMFGYDVGVSGGVASMPPFLKKFFPTVLRQTTESDGSESNYCKYDNQGLQLFTS 1549
Query: 71 SMASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLG 130
S+ AG + F Y +GR+ T+L+A F+AG + A+A ++I+GR+L+G G
Sbjct: 1550 SLYLAGLTV-TFFASYTTRVLGRRLTMLIAGFFFIAGVSLNASAQNLLMLIVGRVLLGCG 1726
Query: 131 VGAASMTAPLYISEASPAKIRGAL 154
+G A+ P+++SE +P++IRGAL
Sbjct: 1727 IGFANQAVPVFLSEIAPSRIRGAL 1798
Score = 65.5 bits (158), Expect = 4e-11
Identities = 44/159 (27%), Positives = 80/159 (49%), Gaps = 4/159 (2%)
Frame = +3
Query: 188 EEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGLIGHSLAQKLKGAWSNDVVRRG--- 244
++ K +L KI +E E E +EA + +A+++K + N + R
Sbjct: 1995 DKGKAVLRKIRGTDNIEPEFL---ELVEASR---------VAKEVKHPFRNLLKRNNRPQ 2138
Query: 245 LYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMVLI 304
L I + + QQ GIN IM+Y+P + G N A +++T +N + TIVS+ +
Sbjct: 2139 LVISIALMIFQQFTGINAIMFYAPVLFNTLGF-KNDAALYSAVITGAINVISTIVSIYSV 2315
Query: 305 DRFGRRKLMLISLIGIFVSLVTLSVTFN-QAAHHAPSLS 342
D+ GRRKL+L + + + +S + +++ + H+ LS
Sbjct: 2316 DKLGRRKLLLEAGVQMLLSQMVIAIVLGIKVKDHSEELS 2432
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,588,109
Number of Sequences: 36976
Number of extensions: 180032
Number of successful extensions: 1074
Number of sequences better than 10.0: 88
Number of HSP's better than 10.0 without gapping: 1018
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1038
length of query: 420
length of database: 9,014,727
effective HSP length: 99
effective length of query: 321
effective length of database: 5,354,103
effective search space: 1718667063
effective search space used: 1718667063
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC148526.6


