

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148525.11 + phase: 0 /pseudo
(88 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
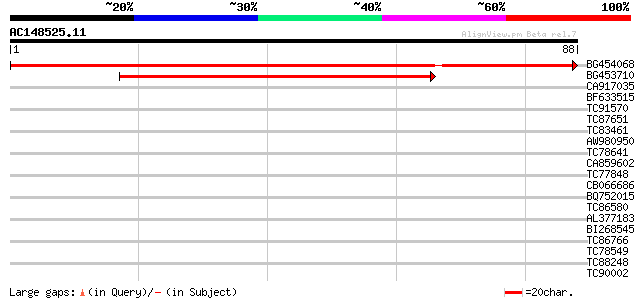
BG454068 weakly similar to GP|22202804|dbj hypothetical protein~... 75 3e-15
BG453710 weakly similar to GP|21537057|gb| unknown {Arabidopsis ... 56 2e-09
CA917035 similar to GP|7303305|gb|A CG6061-PA {Drosophila melano... 36 0.002
BF633515 35 0.004
TC91570 weakly similar to GP|7208782|emb|CAB76913.1 hypothetical... 28 0.80
TC87651 similar to GP|15292729|gb|AAK92733.1 putative polygalact... 26 3.0
TC83461 similar to PIR|T04778|T04778 hypothetical protein F10M10... 25 4.0
AW980950 25 5.2
TC78641 25 5.2
CA859602 25 6.8
TC77848 similar to PIR|F86296|F86296 hypothetical protein AAF185... 25 6.8
CB066686 similar to GP|14334416|gb| unknown protein {Arabidopsis... 24 8.9
BQ752015 homologue to GP|18376006|em probable ATP citrate lyase ... 24 8.9
TC86580 similar to GP|6513932|gb|AAF14836.1| unknown protein {Ar... 24 8.9
AL377183 GP|2598587|emb cycloartenol synthase {Medicago truncatu... 24 8.9
BI268545 24 8.9
TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E... 24 8.9
TC78549 similar to GP|15027268|emb|CAC44902. replication licensi... 24 8.9
TC88248 cycloartenol synthase 24 8.9
TC90002 weakly similar to GP|9757850|dbj|BAB08484.1 dimethylanil... 24 8.9
>BG454068 weakly similar to GP|22202804|dbj hypothetical protein~predicted by
GeneMark.hmm etc. {Oryza sativa (japonica
cultivar-group)}, partial (5%)
Length = 673
Score = 75.5 bits (184), Expect = 3e-15
Identities = 37/88 (42%), Positives = 54/88 (61%)
Frame = +3
Query: 1 MRVSGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIES 60
+R SGL L +Y +D GLL + RWH TSSFHLP+ EM + LDD+SCLLH+ +
Sbjct: 276 IRASGLYPLLETNYGQVDHGLLIAFSLRWHSETSSFHLPVGEMTIALDDISCLLHIPVGG 455
Query: 61 HMLYHNIYLTRSEAVDLMVELLGSDSGE 88
++L+H L+ + + +V LG + E
Sbjct: 456 NLLFHE-SLSIHQGTEYLVNYLGLEFEE 536
>BG453710 weakly similar to GP|21537057|gb| unknown {Arabidopsis thaliana},
partial (6%)
Length = 651
Score = 56.2 bits (134), Expect = 2e-09
Identities = 25/49 (51%), Positives = 33/49 (67%)
Frame = -1
Query: 18 DQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHMLYHN 66
D ++ + ERWH TSSFHLP+ EM +TLDD+ LLH+ I ML H+
Sbjct: 474 DPSMIRAMCERWHTETSSFHLPMGEMTITLDDVYNLLHIPIHGCMLDHD 328
>CA917035 similar to GP|7303305|gb|A CG6061-PA {Drosophila melanogaster},
partial (2%)
Length = 365
Score = 36.2 bits (82), Expect = 0.002
Identities = 16/34 (47%), Positives = 20/34 (58%)
Frame = +2
Query: 4 SGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFH 37
SGL L +Y +D LL ++RWH TSSFH
Sbjct: 263 SGLYLLLETNYGQVDHSLLIAFSDRWHSETSSFH 364
>BF633515
Length = 249
Score = 35.4 bits (80), Expect = 0.004
Identities = 17/42 (40%), Positives = 23/42 (54%)
Frame = +2
Query: 47 LDDLSCLLHLSIESHMLYHNIYLTRSEAVDLMVELLGSDSGE 88
LDD++CL HL IE ML H + + E L++ LG E
Sbjct: 2 LDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQE 127
>TC91570 weakly similar to GP|7208782|emb|CAB76913.1 hypothetical protein
{Cicer arietinum}, partial (17%)
Length = 1446
Score = 27.7 bits (60), Expect = 0.80
Identities = 17/55 (30%), Positives = 26/55 (46%)
Frame = -2
Query: 11 HVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHMLYH 65
H S L + LS++ WH SS + ++ + DLS L H + S + YH
Sbjct: 584 HYSLLALSVQDLSLL---WHRHLSSSLAYHYSLVFFVQDLSLL*HRHLSSSLAYH 429
>TC87651 similar to GP|15292729|gb|AAK92733.1 putative polygalacturonase PG1
{Arabidopsis thaliana}, partial (35%)
Length = 822
Score = 25.8 bits (55), Expect = 3.0
Identities = 24/99 (24%), Positives = 39/99 (39%), Gaps = 11/99 (11%)
Frame = +1
Query: 1 MRVSGLQWLQHVSYLVIDQGLLSVIAERWHEVTS---------SFHLP--IWEMIVTLDD 49
+++ W+ + YL L +I E W E T+ F LP + ++TL
Sbjct: 154 IQILNFSWISFMDYLFFGLPLYYLI*EMWMEGTTFTPSKRKYLPFFLPTLLLVFLLTLTQ 333
Query: 50 LSCLLHLSIESHMLYHNIYLTRSEAVDLMVELLGSDSGE 88
+ + L ML+H L + E V M L G+
Sbjct: 334 MILAIRLRTVFSMLHH---LEQLEMVQQMTHLHSRKHGK 441
>TC83461 similar to PIR|T04778|T04778 hypothetical protein F10M10.90 -
Arabidopsis thaliana, partial (25%)
Length = 663
Score = 25.4 bits (54), Expect = 4.0
Identities = 15/43 (34%), Positives = 24/43 (54%), Gaps = 3/43 (6%)
Frame = -2
Query: 32 VTSSFHLP-IWEMIVTLDDLSCLLHL--SIESHMLYHNIYLTR 71
VTS +P + + + T +LHL I +H L+H++ LTR
Sbjct: 653 VTSILKIPKLLQCLYTSPPFKPILHLFLKISNHRLHHHLILTR 525
>AW980950
Length = 548
Score = 25.0 bits (53), Expect = 5.2
Identities = 9/27 (33%), Positives = 18/27 (66%)
Frame = +1
Query: 21 LLSVIAERWHEVTSSFHLPIWEMIVTL 47
+++VI+ +W + SF LP+W + T+
Sbjct: 430 VIAVISNKWIYIYFSFPLPLWLLY*TI 510
>TC78641
Length = 550
Score = 25.0 bits (53), Expect = 5.2
Identities = 9/27 (33%), Positives = 18/27 (66%)
Frame = +1
Query: 21 LLSVIAERWHEVTSSFHLPIWEMIVTL 47
+++VI+ +W + SF LP+W + T+
Sbjct: 367 VIAVISNKWIYIYFSFPLPLWLLY*TI 447
>CA859602
Length = 587
Score = 24.6 bits (52), Expect = 6.8
Identities = 10/33 (30%), Positives = 22/33 (66%)
Frame = -1
Query: 36 FHLPIWEMIVTLDDLSCLLHLSIESHMLYHNIY 68
F LP++ ++ ++ L+ LLH+ + + + +NIY
Sbjct: 317 FILPLYNLLNDINVLNLLLHIFVYTLNI*YNIY 219
>TC77848 similar to PIR|F86296|F86296 hypothetical protein AAF18512.1
[imported] - Arabidopsis thaliana, complete
Length = 1592
Score = 24.6 bits (52), Expect = 6.8
Identities = 11/52 (21%), Positives = 24/52 (46%)
Frame = +2
Query: 14 YLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHMLYH 65
+L++ LL RW++ + W + + + L C + + S +L+H
Sbjct: 593 FLLVQVVLLLDFVHRWNDTWVGYDEQFWYIALFVVSLVCYVATFVFSGVLFH 748
>CB066686 similar to GP|14334416|gb| unknown protein {Arabidopsis thaliana},
partial (4%)
Length = 306
Score = 24.3 bits (51), Expect = 8.9
Identities = 13/36 (36%), Positives = 20/36 (55%)
Frame = +1
Query: 21 LLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHL 56
LLS++ E+ SF +WE+I+ L L LH+
Sbjct: 166 LLSLVREK----NFSFKFQVWEIIIELFLLRSYLHI 261
>BQ752015 homologue to GP|18376006|em probable ATP citrate lyase subunit 1
{Neurospora crassa}, partial (23%)
Length = 523
Score = 24.3 bits (51), Expect = 8.9
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = -2
Query: 4 SGLQWLQHVSYLVIDQGLLSVIAER 28
+GLQ + S LV++QG+L A+R
Sbjct: 213 AGLQTVDEASGLVVEQGVLGKRAQR 139
>TC86580 similar to GP|6513932|gb|AAF14836.1| unknown protein {Arabidopsis
thaliana}, partial (83%)
Length = 2108
Score = 24.3 bits (51), Expect = 8.9
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = +2
Query: 30 HEVTSSFHLPIWEMIVTLDDLSC 52
H+ S +HLP I++L LSC
Sbjct: 101 HKTDSKYHLPTSLFILSLSFLSC 169
>AL377183 GP|2598587|emb cycloartenol synthase {Medicago truncatula}, partial
(12%)
Length = 370
Score = 24.3 bits (51), Expect = 8.9
Identities = 12/30 (40%), Positives = 16/30 (53%)
Frame = +1
Query: 30 HEVTSSFHLPIWEMIVTLDDLSCLLHLSIE 59
H + +SF +WE IVT L L L +E
Sbjct: 88 HHIETSFPYGLWESIVTKSCLHKLQMLHLE 177
>BI268545
Length = 467
Score = 24.3 bits (51), Expect = 8.9
Identities = 13/39 (33%), Positives = 20/39 (50%)
Frame = -1
Query: 9 LQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTL 47
+ H SY D ++ +I + H + FHLPI+E L
Sbjct: 290 IDHRSY---DMFIIQIIEK*RHCLEKHFHLPIYESFAYL 183
>TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E8.80 -
Arabidopsis thaliana, partial (18%)
Length = 1260
Score = 24.3 bits (51), Expect = 8.9
Identities = 9/14 (64%), Positives = 12/14 (85%)
Frame = +3
Query: 53 LLHLSIESHMLYHN 66
L HLS+ESHM +H+
Sbjct: 537 LYHLSLESHMHHHH 578
>TC78549 similar to GP|15027268|emb|CAC44902. replication licensing factor
MCM7 homologue {Zea mays}, partial (73%)
Length = 1900
Score = 24.3 bits (51), Expect = 8.9
Identities = 6/15 (40%), Positives = 13/15 (86%)
Frame = -2
Query: 52 CLLHLSIESHMLYHN 66
CL+H+++ + +L+HN
Sbjct: 1434 CLIHITLSNSLLFHN 1390
>TC88248 cycloartenol synthase
Length = 1674
Score = 24.3 bits (51), Expect = 8.9
Identities = 12/30 (40%), Positives = 16/30 (53%)
Frame = +3
Query: 30 HEVTSSFHLPIWEMIVTLDDLSCLLHLSIE 59
H + +SF +WE IVT L L L +E
Sbjct: 1335 HHIETSFPYGLWESIVTKSCLHKLQMLHLE 1424
>TC90002 weakly similar to GP|9757850|dbj|BAB08484.1 dimethylaniline
monooxygenase (N-oxide-forming)-like protein
{Arabidopsis thaliana}, partial (26%)
Length = 778
Score = 24.3 bits (51), Expect = 8.9
Identities = 12/34 (35%), Positives = 17/34 (49%)
Frame = -2
Query: 6 LQWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLP 39
LQWL +L + G+ ++ E WH HLP
Sbjct: 195 LQWLLFHLFLGKECGICTLYNELWHLHR*RIHLP 94
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.137 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,356,949
Number of Sequences: 36976
Number of extensions: 48103
Number of successful extensions: 322
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 318
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 321
length of query: 88
length of database: 9,014,727
effective HSP length: 64
effective length of query: 24
effective length of database: 6,648,263
effective search space: 159558312
effective search space used: 159558312
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 51 (24.3 bits)
Medicago: description of AC148525.11


