

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148485.1 - phase: 0
(55 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
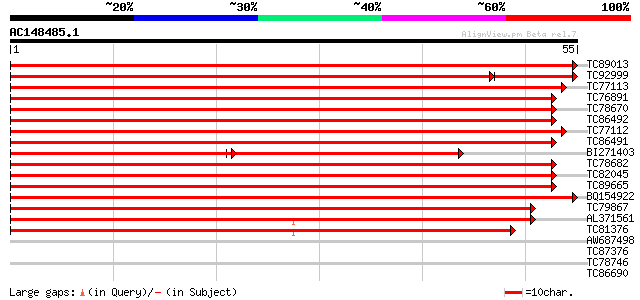
Sequences producing significant alignments: (bits) Value
TC89013 homologue to GP|11932089|emb|CAC19183. expansin {Cicer a... 108 6e-25
TC92999 similar to GP|11932089|emb|CAC19183. expansin {Cicer ari... 104 8e-24
TC77113 homologue to GP|21615409|emb|CAD33924. alpha-expansin 4 ... 94 1e-20
TC76891 homologue to GP|21615407|emb|CAD33923. alpha-expansin 3 ... 93 2e-20
TC78670 homologue to GP|11932092|emb|CAC19184. expansin {Cicer a... 92 4e-20
TC86492 homologue to GP|11907554|dbj|BAB19676. expansin {Prunus ... 91 7e-20
TC77112 homologue to GP|21615409|emb|CAD33924. alpha-expansin 4 ... 89 5e-19
TC86491 homologue to GP|11907554|dbj|BAB19676. expansin {Prunus ... 89 5e-19
BI271403 similar to SP|Q38864|EXP5 Alpha-expansin 5 precursor (A... 52 5e-17
TC78682 similar to GP|13898655|gb|AAK48848.1 expansin {Prunus ce... 79 4e-16
TC82045 similar to GP|13898655|gb|AAK48848.1 expansin {Prunus ce... 77 1e-15
TC89665 similar to GP|13898655|gb|AAK48848.1 expansin {Prunus ce... 76 3e-15
BQ154922 similar to GP|17645715|em unnamed protein product {Lyco... 75 4e-15
TC79867 similar to PIR|T10083|T10083 expansin S2 precursor - cuc... 69 5e-13
AL371561 similar to PIR|F86259|F862 protein T12C24.10 [imported]... 50 2e-07
TC81376 similar to SP|Q9LN94|EXP7_ARATH Alpha-expansin 7 precurs... 49 4e-07
AW687498 similar to PIR|T50657|T50 beta-expansin [imported] - Ar... 34 0.013
TC87376 similar to GP|21592834|gb|AAM64784.1 putative beta-expan... 27 1.2
TC78746 similar to GP|22137234|gb|AAM91462.1 AT5g39360/MUL8_40 {... 26 3.6
TC86690 similar to GP|6041851|gb|AAF02160.1| unknown protein {Ar... 26 3.6
>TC89013 homologue to GP|11932089|emb|CAC19183. expansin {Cicer arietinum},
partial (94%)
Length = 1318
Score = 108 bits (269), Expect = 6e-25
Identities = 48/55 (87%), Positives = 49/55 (88%)
Frame = +3
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQTYTGRQFLY 55
MSRNWGQNWQ NSYLNGQSLSFVVTT NGHS+VSFN AP WSFGQTYTGRQF Y
Sbjct: 567 MSRNWGQNWQRNSYLNGQSLSFVVTTSNGHSVVSFNAAPAGWSFGQTYTGRQFNY 731
>TC92999 similar to GP|11932089|emb|CAC19183. expansin {Cicer arietinum},
partial (96%)
Length = 767
Score = 104 bits (259), Expect(2) = 8e-24
Identities = 47/47 (100%), Positives = 47/47 (100%)
Frame = +2
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQT 47
MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQT
Sbjct: 581 MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQT 721
Score = 20.8 bits (42), Expect(2) = 8e-24
Identities = 7/8 (87%), Positives = 8/8 (99%)
Frame = +1
Query: 48 YTGRQFLY 55
YTG+QFLY
Sbjct: 721 YTGKQFLY 744
>TC77113 homologue to GP|21615409|emb|CAD33924. alpha-expansin 4 {Cicer
arietinum}, partial (27%)
Length = 1070
Score = 94.0 bits (232), Expect = 1e-20
Identities = 42/54 (77%), Positives = 46/54 (84%)
Frame = +2
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQTYTGRQFL 54
MSRNWGQNWQSN+YLNGQSLSF VTT +G ++VS NVAP WSFGQTYTG Q L
Sbjct: 431 MSRNWGQNWQSNNYLNGQSLSFKVTTSDGRTVVSNNVAPAGWSFGQTYTGAQIL 592
>TC76891 homologue to GP|21615407|emb|CAD33923. alpha-expansin 3 {Cicer
arietinum}, complete
Length = 1432
Score = 92.8 bits (229), Expect = 2e-20
Identities = 40/53 (75%), Positives = 46/53 (86%)
Frame = +3
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQTYTGRQF 53
MSRNWGQNWQSN+YLNGQ+LSF VTT +G +++S NVAP WSFGQTYTG QF
Sbjct: 705 MSRNWGQNWQSNNYLNGQALSFKVTTSDGKTVISNNVAPSGWSFGQTYTGAQF 863
>TC78670 homologue to GP|11932092|emb|CAC19184. expansin {Cicer arietinum},
complete
Length = 1043
Score = 92.0 bits (227), Expect = 4e-20
Identities = 40/53 (75%), Positives = 45/53 (84%)
Frame = +1
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQTYTGRQF 53
MSRNWGQNWQSNSYLNGQSLSF VTT +G ++ SFNVAP +W FGQT+ G QF
Sbjct: 667 MSRNWGQNWQSNSYLNGQSLSFQVTTSDGRTMTSFNVAPANWQFGQTFQGAQF 825
>TC86492 homologue to GP|11907554|dbj|BAB19676. expansin {Prunus persica},
partial (95%)
Length = 1257
Score = 91.3 bits (225), Expect = 7e-20
Identities = 40/53 (75%), Positives = 45/53 (84%)
Frame = +3
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQTYTGRQF 53
MSRNWGQNWQSN+YLNGQSLSF VTT +G +I S NV P +W FGQT+TGRQF
Sbjct: 780 MSRNWGQNWQSNNYLNGQSLSFQVTTSDGRTITSNNVVPGNWQFGQTFTGRQF 938
>TC77112 homologue to GP|21615409|emb|CAD33924. alpha-expansin 4 {Cicer
arietinum}, complete
Length = 859
Score = 88.6 bits (218), Expect = 5e-19
Identities = 41/54 (75%), Positives = 45/54 (82%)
Frame = +1
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQTYTGRQFL 54
MSRNWGQNWQSN+YLNGQSLSF VTT +G ++VS NVAP SFGQTYTG Q L
Sbjct: 619 MSRNWGQNWQSNNYLNGQSLSFKVTTSDGRTVVSNNVAPAGRSFGQTYTGAQIL 780
>TC86491 homologue to GP|11907554|dbj|BAB19676. expansin {Prunus persica},
partial (95%)
Length = 1166
Score = 88.6 bits (218), Expect = 5e-19
Identities = 39/53 (73%), Positives = 44/53 (82%)
Frame = +2
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQTYTGRQF 53
MSRNWGQNWQSN+YLNGQSLSF VTT +G +I S NV P +W FGQT+TG QF
Sbjct: 671 MSRNWGQNWQSNNYLNGQSLSFQVTTSDGRTITSNNVVPGNWQFGQTFTGGQF 829
>BI271403 similar to SP|Q38864|EXP5 Alpha-expansin 5 precursor (At-EXP5)
(AtEx5) (Ath-ExpAlpha-1.4). [Mouse-ear cress], partial
(65%)
Length = 664
Score = 52.4 bits (124), Expect(2) = 5e-17
Identities = 22/22 (100%), Positives = 22/22 (100%)
Frame = +2
Query: 1 MSRNWGQNWQSNSYLNGQSLSF 22
MSRNWGQNWQSNSYLNGQSLSF
Sbjct: 530 MSRNWGQNWQSNSYLNGQSLSF 595
Score = 49.7 bits (117), Expect(2) = 5e-17
Identities = 21/23 (91%), Positives = 21/23 (91%)
Frame = +3
Query: 22 FVVTTGNGHSIVSFNVAPPSWSF 44
FVVTTGNGHSIVSFNVAPP W F
Sbjct: 594 FVVTTGNGHSIVSFNVAPPXWXF 662
>TC78682 similar to GP|13898655|gb|AAK48848.1 expansin {Prunus cerasus},
partial (91%)
Length = 1236
Score = 79.0 bits (193), Expect = 4e-16
Identities = 33/53 (62%), Positives = 43/53 (80%)
Frame = +1
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQTYTGRQF 53
+SRNWGQNWQSN+ L GQ+LSF V++ + S S+N+APPSW FGQT+TG+ F
Sbjct: 706 LSRNWGQNWQSNAVLVGQALSFRVSSSDRRSSTSWNIAPPSWQFGQTFTGKNF 864
>TC82045 similar to GP|13898655|gb|AAK48848.1 expansin {Prunus cerasus},
partial (91%)
Length = 814
Score = 77.4 bits (189), Expect = 1e-15
Identities = 33/53 (62%), Positives = 42/53 (78%)
Frame = +3
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQTYTGRQF 53
MSRNWGQNWQSNS L GQSLSF VT+ + + S+N+ P +W FGQT+TG++F
Sbjct: 624 MSRNWGQNWQSNSVLVGQSLSFRVTSSDKRTSTSWNIVPSNWQFGQTFTGKKF 782
>TC89665 similar to GP|13898655|gb|AAK48848.1 expansin {Prunus cerasus},
partial (92%)
Length = 1006
Score = 75.9 bits (185), Expect = 3e-15
Identities = 33/53 (62%), Positives = 41/53 (77%)
Frame = +3
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQTYTGRQF 53
MSRNWGQNWQSNS L GQ+LSF VT + + S+N+AP +W FGQT+TG+ F
Sbjct: 684 MSRNWGQNWQSNSVLVGQALSFRVTGSDRRTSTSWNIAPHNWQFGQTFTGKNF 842
>BQ154922 similar to GP|17645715|em unnamed protein product {Lycopersicon
esculentum}, partial (45%)
Length = 599
Score = 75.5 bits (184), Expect = 4e-15
Identities = 31/55 (56%), Positives = 40/55 (72%)
Frame = +2
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQTYTGRQFLY 55
M RNWGQNW N+ L Q LSF VT+ +G ++ ++NVAP WSFGQT+ G+QF Y
Sbjct: 188 MGRNWGQNWHINALLQNQPLSFEVTSSDGKTVTAYNVAPKDWSFGQTFEGKQFDY 352
>TC79867 similar to PIR|T10083|T10083 expansin S2 precursor - cucumber,
partial (87%)
Length = 1211
Score = 68.6 bits (166), Expect = 5e-13
Identities = 28/51 (54%), Positives = 39/51 (75%)
Frame = +2
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQTYTGR 51
MSRNWG NWQSN+YLNGQS+SF VTT +G + ++ P +W FGQ+++ +
Sbjct: 536 MSRNWGANWQSNAYLNGQSMSFKVTTTDGVTRTFQDIVPSNWGFGQSFSSK 688
>AL371561 similar to PIR|F86259|F862 protein T12C24.10 [imported] -
Arabidopsis thaliana, partial (32%)
Length = 500
Score = 50.1 bits (118), Expect = 2e-07
Identities = 23/52 (44%), Positives = 37/52 (70%), Gaps = 1/52 (1%)
Frame = +3
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTT-GNGHSIVSFNVAPPSWSFGQTYTGR 51
MS NWG ++Q+ + L GQ+LSF +T+ +I+++NVAP +W+ G TY+ R
Sbjct: 90 MSHNWGASYQAFATLGGQALSFRLTSYTTKETIIAWNVAPSNWNVGLTYSSR 245
>TC81376 similar to SP|Q9LN94|EXP7_ARATH Alpha-expansin 7 precursor
(At-EXP7) (AtEx7) (Ath-ExpAlpha-1.26). [Mouse-ear
cress], partial (80%)
Length = 997
Score = 48.9 bits (115), Expect = 4e-07
Identities = 22/50 (44%), Positives = 36/50 (72%), Gaps = 1/50 (2%)
Frame = +1
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTT-GNGHSIVSFNVAPPSWSFGQTYT 49
MS NWG ++Q+ + L+GQ+LSF +T+ +I+++NVAP +W G TY+
Sbjct: 631 MSHNWGASYQAFATLSGQTLSFKITSYTTKETIIAWNVAPSNWGAGLTYS 780
>AW687498 similar to PIR|T50657|T50 beta-expansin [imported] - Arabidopsis
thaliana (fragment), partial (42%)
Length = 415
Score = 33.9 bits (76), Expect = 0.013
Identities = 18/51 (35%), Positives = 24/51 (46%)
Frame = +3
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQTYTGR 51
M+ WG NW + S ++T G SI +V P +WS TYT R
Sbjct: 204 MNHLWGANWNIVTGPLRGPFSVKLSTSTGKSITVKDVIPSNWSPKSTYTSR 356
>TC87376 similar to GP|21592834|gb|AAM64784.1 putative
beta-expansin/allergen protein {Arabidopsis thaliana},
partial (81%)
Length = 976
Score = 27.3 bits (59), Expect = 1.2
Identities = 13/43 (30%), Positives = 20/43 (46%)
Frame = +2
Query: 1 MSRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWS 43
M+ WG NW + S ++T G S+ + +V P WS
Sbjct: 710 MNHLWGANWNIVTGPLRGPFSVKLSTSTGKSLTAKDVIPSHWS 838
>TC78746 similar to GP|22137234|gb|AAM91462.1 AT5g39360/MUL8_40 {Arabidopsis
thaliana}, partial (97%)
Length = 1222
Score = 25.8 bits (55), Expect = 3.6
Identities = 9/19 (47%), Positives = 13/19 (68%)
Frame = -2
Query: 30 HSIVSFNVAPPSWSFGQTY 48
HS+ + N+APP+WS Y
Sbjct: 1212 HSLKTTNIAPPNWSLRFRY 1156
>TC86690 similar to GP|6041851|gb|AAF02160.1| unknown protein {Arabidopsis
thaliana}, partial (63%)
Length = 1592
Score = 25.8 bits (55), Expect = 3.6
Identities = 15/49 (30%), Positives = 23/49 (46%)
Frame = +3
Query: 2 SRNWGQNWQSNSYLNGQSLSFVVTTGNGHSIVSFNVAPPSWSFGQTYTG 50
+R W QN+ +NSY G G S +++N S +G +Y G
Sbjct: 426 TRPWEQNYGTNSYGGG-------ALGGYGSTMNYNSGYGSGMYGSSYGG 551
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.130 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,222,873
Number of Sequences: 36976
Number of extensions: 22561
Number of successful extensions: 155
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 151
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 152
length of query: 55
length of database: 9,014,727
effective HSP length: 31
effective length of query: 24
effective length of database: 7,868,471
effective search space: 188843304
effective search space used: 188843304
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 51 (24.3 bits)
Medicago: description of AC148485.1


