

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148483.6 + phase: 0
(125 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
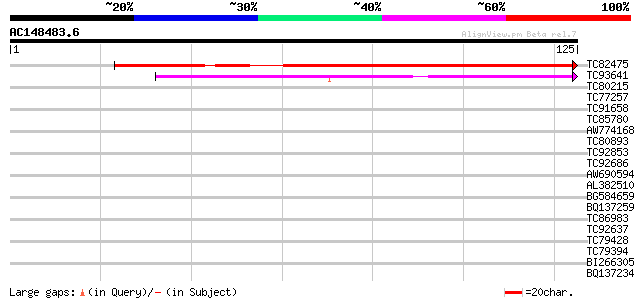
Sequences producing significant alignments: (bits) Value
TC82475 similar to GP|20146289|dbj|BAB89071. peptide deformylase... 99 6e-22
TC93641 similar to SP|Q9FV54|DEFC_LYCES Peptide deformylase chl... 39 4e-04
TC80215 similar to GP|7547106|gb|AAF63778.1| unknown protein {Ar... 31 0.11
TC77257 weakly similar to GP|14134199|gb|AAK54282.1 fatty acid 9... 30 0.25
TC91658 similar to GP|14329812|emb|CAC40753. putative nucleosome... 29 0.55
TC85780 similar to SP|P49608|ACOC_CUCMA Aconitate hydratase cyt... 28 0.71
AW774168 similar to GP|20197888|gb| Expressed protein {Arabidops... 27 1.6
TC80893 similar to GP|4557063|gb|AAD22502.1| expressed protein {... 27 1.6
TC92853 similar to GP|20268748|gb|AAM14077.1 unknown protein {Ar... 27 2.1
TC92686 weakly similar to PIR|T39560|T39560 dolichyl-phosphate-m... 27 2.1
AW690594 similar to GP|23498163|emb hypothetical protein {Plasmo... 27 2.7
AL382510 weakly similar to GP|13194672|gb| calmodulin-like prote... 27 2.7
BG584659 weakly similar to GP|12963415|gb| regulator of gene sil... 27 2.7
BQ137259 homologue to GP|15387793|emb hypothetical predicted pro... 26 3.5
TC86983 similar to GP|9971154|dbj|BAB12429.1 mkpA {Dictyostelium... 26 3.5
TC92637 similar to PIR|T39903|T39903 serine-rich protein - fissi... 26 3.5
TC79428 similar to PIR|T09661|T09661 ascorbate oxidase promoter-... 26 4.6
TC79394 weakly similar to PIR|T49019|T49019 probable RNA binding... 26 4.6
BI266305 similar to GP|10176943|dbj gene_id:K20J1.7~unknown prot... 26 4.6
BQ137234 similar to EGAD|35719|3709 hypothetical protein {Burkho... 25 6.0
>TC82475 similar to GP|20146289|dbj|BAB89071. peptide deformylase-like
protein {Oryza sativa (japonica cultivar-group)},
partial (52%)
Length = 654
Score = 98.6 bits (244), Expect = 6e-22
Identities = 58/102 (56%), Positives = 69/102 (66%)
Frame = +3
Query: 24 VSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIHEPAR 83
++LSSSSS N I+ + F FG T +K LP VKAGDPV+HEPA+
Sbjct: 159 LTLSSSSSQNATIRTRAGFF--FGRTKDDK-------KKKKMDLPDTVKAGDPVLHEPAQ 311
Query: 84 EVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
EVD SEI SDK+Q IIDDMI VMRKAPGVG+AAPQIG+ R+
Sbjct: 312 EVDPSEIMSDKVQKIIDDMIRVMRKAPGVGLAAPQIGVSSRI 437
>TC93641 similar to SP|Q9FV54|DEFC_LYCES Peptide deformylase chloroplast
precursor (EC 3.5.1.88) (PDF) (Polypeptide deformylase).
[Tomato], partial (59%)
Length = 644
Score = 39.3 bits (90), Expect = 4e-04
Identities = 25/94 (26%), Positives = 49/94 (51%), Gaps = 1/94 (1%)
Frame = +1
Query: 33 NNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPY-IVKAGDPVIHEPAREVDHSEIK 91
N L+ + S+ S S+ +E A L + P I K DP + + + + +
Sbjct: 31 NRFFYLTLSSSSRTKSAFSASQNEFASLGDLEFEAPLKIAKYPDPKLRKKNKRIGTFD-- 204
Query: 92 SDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
D ++ ++D+M VM + G+G++APQ+GI +++
Sbjct: 205 -DNLKKLVDEMFDVMYETDGIGLSAPQVGINVQL 303
>TC80215 similar to GP|7547106|gb|AAF63778.1| unknown protein {Arabidopsis
thaliana}, partial (50%)
Length = 1486
Score = 31.2 bits (69), Expect = 0.11
Identities = 19/70 (27%), Positives = 39/70 (55%)
Frame = +1
Query: 41 TKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIHEPAREVDHSEIKSDKIQNIID 100
TK++ + L+ + A LR+ V LP + KAG+ + A + +E+++D++ ++ID
Sbjct: 424 TKYTHSPAHLAVARRDHAALRRIVTGLPQLAKAGE--VKNEAESL-AAELRADEVSSVID 594
Query: 101 DMILVMRKAP 110
+ R+ P
Sbjct: 595 RRDVPGRETP 624
>TC77257 weakly similar to GP|14134199|gb|AAK54282.1 fatty acid
9-hydroperoxide lyase {Cucumis melo}, partial (81%)
Length = 1989
Score = 30.0 bits (66), Expect = 0.25
Identities = 18/46 (39%), Positives = 24/46 (52%), Gaps = 1/46 (2%)
Frame = +3
Query: 20 SNGVVSLSSSSSCNNKI-QLSSTKFSKFGSTLSSPSSETALLRKTV 64
S LSS+S+C+ + L + K S F + L SPSS T K V
Sbjct: 252 SKNTTPLSSNSTCHQVVSSLQTQKSSLFSTVLPSPSSSTTPKSKNV 389
>TC91658 similar to GP|14329812|emb|CAC40753. putative nucleosome assembly
protein 1 {Atropa belladonna}, partial (46%)
Length = 583
Score = 28.9 bits (63), Expect = 0.55
Identities = 25/72 (34%), Positives = 34/72 (46%)
Frame = -2
Query: 11 LRAFPMPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYI 70
L FP P S+ S SSSSS SS+ S S+ SS SS T++ K+++
Sbjct: 201 LLLFPFPSSSSSSSSSSSSSS-------SSSSSSSSSSSPSSSSSSTSISSKSLS----- 58
Query: 71 VKAGDPVIHEPA 82
PV H+ A
Sbjct: 57 --PASPVNHDTA 28
>TC85780 similar to SP|P49608|ACOC_CUCMA Aconitate hydratase cytoplasmic
(EC 4.2.1.3) (Citrate hydro-lyase) (Aconitase)., partial
(97%)
Length = 3124
Score = 28.5 bits (62), Expect = 0.71
Identities = 17/56 (30%), Positives = 28/56 (49%)
Frame = +3
Query: 45 KFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIHEPAREVDHSEIKSDKIQNIID 100
+FG S P AL ++ LPY ++ ++ R D ++KSD ++ IID
Sbjct: 138 EFGKYYSLP----ALNDSRIDALPYSIRI---LLESAIRNCDEFQVKSDDVEKIID 284
>AW774168 similar to GP|20197888|gb| Expressed protein {Arabidopsis
thaliana}, partial (26%)
Length = 625
Score = 27.3 bits (59), Expect = 1.6
Identities = 22/77 (28%), Positives = 34/77 (43%), Gaps = 2/77 (2%)
Frame = +2
Query: 14 FPMPVLSNGVVSLSSSSSCNNKIQL--SSTKFSKFGSTLSSPSSETALLRKTVNKLPYIV 71
FP P S + +S+ NNKIQ+ + T+ + +P S RK KLP
Sbjct: 17 FPFPFSSQPKILFPNSN*KNNKIQILQNLTEKQHLNTRFHAPPSWNNRSRKRTPKLPTRT 196
Query: 72 KAGDPVIHEPAREVDHS 88
+ P EP ++D +
Sbjct: 197ET-PPQQREPDPDLDRN 244
>TC80893 similar to GP|4557063|gb|AAD22502.1| expressed protein {Arabidopsis
thaliana}, partial (39%)
Length = 883
Score = 27.3 bits (59), Expect = 1.6
Identities = 17/34 (50%), Positives = 21/34 (61%)
Frame = -1
Query: 22 GVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSS 55
GV S SSSSS ++ SS+ S S+ SSPSS
Sbjct: 565 GVSSSSSSSSSSSSSSPSSSSSSSPSSSSSSPSS 464
>TC92853 similar to GP|20268748|gb|AAM14077.1 unknown protein {Arabidopsis
thaliana}, partial (21%)
Length = 761
Score = 26.9 bits (58), Expect = 2.1
Identities = 15/35 (42%), Positives = 21/35 (59%)
Frame = -2
Query: 23 VVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSET 57
V SLSSSS+ N+ + L S+ S+ SS SS +
Sbjct: 493 VQSLSSSSALNSSLSLPSSSLVSSSSSSSSSSSSS 389
>TC92686 weakly similar to PIR|T39560|T39560
dolichyl-phosphate-mannose--protein O-mannosyl
transferase - fission yeast, partial (9%)
Length = 768
Score = 26.9 bits (58), Expect = 2.1
Identities = 18/49 (36%), Positives = 24/49 (48%)
Frame = -2
Query: 15 PMPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKT 63
P+P L+ V+ S C N + SS K GS L+S S + LL T
Sbjct: 731 PLPRLAGVRVTASQLLLCQNMER*SSNHLRKKGSDLTSFSGFSRLLEST 585
>AW690594 similar to GP|23498163|emb hypothetical protein {Plasmodium
falciparum 3D7}, partial (10%)
Length = 633
Score = 26.6 bits (57), Expect = 2.7
Identities = 15/42 (35%), Positives = 23/42 (54%)
Frame = -3
Query: 25 SLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNK 66
S SSSSS ++ SS+ S+ +SPSS ++L T +
Sbjct: 157 SSSSSSSSSSSCSSSSSSSCSSSSSSTSPSSSSSLCSSTFRR 32
>AL382510 weakly similar to GP|13194672|gb| calmodulin-like protein
{Pennisetum ciliare}, partial (24%)
Length = 399
Score = 26.6 bits (57), Expect = 2.7
Identities = 16/54 (29%), Positives = 26/54 (47%)
Frame = -2
Query: 23 VVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDP 76
++S+ S +K + SST F+ S SPSS L + N L + + + P
Sbjct: 221 ILSIVFISQPLSKPKQSSTLFNSIASIYPSPSSSNILKASSNNSLFFDISSNSP 60
>BG584659 weakly similar to GP|12963415|gb| regulator of gene silencing
{Nicotiana tabacum}, partial (34%)
Length = 677
Score = 26.6 bits (57), Expect = 2.7
Identities = 13/41 (31%), Positives = 22/41 (52%)
Frame = -2
Query: 15 PMPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSS 55
P P SN +++S+S + N+ + + + FG SPSS
Sbjct: 328 PSPSESNASIAISTSFN*NSPLIFAIRLLNSFGEIFPSPSS 206
>BQ137259 homologue to GP|15387793|emb hypothetical predicted protein
LM15-1.83 unknown function {Leishmania major}, partial
(3%)
Length = 1050
Score = 26.2 bits (56), Expect = 3.5
Identities = 15/30 (50%), Positives = 17/30 (56%)
Frame = -3
Query: 3 AARRLVWRLRAFPMPVLSNGVVSLSSSSSC 32
AARRL WR A P SL+S+SSC
Sbjct: 670 AARRLAWRAAASP--------CSLASTSSC 605
>TC86983 similar to GP|9971154|dbj|BAB12429.1 mkpA {Dictyostelium
discoideum}, partial (0%)
Length = 1141
Score = 26.2 bits (56), Expect = 3.5
Identities = 12/19 (63%), Positives = 13/19 (68%)
Frame = +1
Query: 20 SNGVVSLSSSSSCNNKIQL 38
SN SLSSSSSC K+ L
Sbjct: 352 SNSTTSLSSSSSCTPKLSL 408
>TC92637 similar to PIR|T39903|T39903 serine-rich protein - fission yeast
(Schizosaccharomyces pombe), partial (7%)
Length = 671
Score = 26.2 bits (56), Expect = 3.5
Identities = 25/77 (32%), Positives = 32/77 (41%), Gaps = 9/77 (11%)
Frame = -1
Query: 2 EAARRLVWRLRAFPMPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFG---------STLSS 52
EA L RLR P+P S S SSSSS ++ S++ K G L S
Sbjct: 569 EAETFLYCRLRLLPLPPPS*SSSS*SSSSSSSSSSTSSASSSIKLGLLRFFPPPPPPLFS 390
Query: 53 PSSETALLRKTVNKLPY 69
SS L + N P+
Sbjct: 389 SSSSAFLFIEIFNLFPH 339
>TC79428 similar to PIR|T09661|T09661 ascorbate oxidase promoter-binding
protein AOBP - winter squash, partial (27%)
Length = 997
Score = 25.8 bits (55), Expect = 4.6
Identities = 15/29 (51%), Positives = 19/29 (64%), Gaps = 1/29 (3%)
Frame = -3
Query: 27 SSSSSCNNKIQLSS-TKFSKFGSTLSSPS 54
SSSSS ++ + LSS T + G T SSPS
Sbjct: 284 SSSSSFSDSLSLSSSTSIAAAGETFSSPS 198
>TC79394 weakly similar to PIR|T49019|T49019 probable RNA binding protein -
Arabidopsis thaliana, partial (74%)
Length = 1909
Score = 25.8 bits (55), Expect = 4.6
Identities = 16/39 (41%), Positives = 22/39 (56%)
Frame = -2
Query: 20 SNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETA 58
S+ LS+SSS + + SST S S+ SS SS T+
Sbjct: 417 SSACFFLSASSSSSKGLLSSSTSSSSSSSSSSSTSSSTS 301
>BI266305 similar to GP|10176943|dbj gene_id:K20J1.7~unknown protein
{Arabidopsis thaliana}, partial (11%)
Length = 485
Score = 25.8 bits (55), Expect = 4.6
Identities = 20/97 (20%), Positives = 46/97 (46%)
Frame = +2
Query: 13 AFPMPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVK 72
+FP+P+ + SLSSS L S+ S S++ P + T + +++P+++
Sbjct: 125 SFPLPITPSSSSSLSSSPPSFRSQTLPSSSSSSL-SSIIVPQTSTKHDHHSRSRIPFLLP 301
Query: 73 AGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKA 109
+ ++P+ + + +K N+ D++ K+
Sbjct: 302 KKNNK-NKPSSSI-NMNMKPSSSANVTSDIVFKRSKS 406
>BQ137234 similar to EGAD|35719|3709 hypothetical protein {Burkholderia
cepacia}, partial (6%)
Length = 1184
Score = 25.4 bits (54), Expect = 6.0
Identities = 13/33 (39%), Positives = 21/33 (63%)
Frame = -1
Query: 3 AARRLVWRLRAFPMPVLSNGVVSLSSSSSCNNK 35
AARRLV+ L F PVLS+ + ++ + S ++
Sbjct: 659 AARRLVFSLILFAAPVLSSPALLVTRAVSARSR 561
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.132 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,759,546
Number of Sequences: 36976
Number of extensions: 48670
Number of successful extensions: 433
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 419
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 432
length of query: 125
length of database: 9,014,727
effective HSP length: 84
effective length of query: 41
effective length of database: 5,908,743
effective search space: 242258463
effective search space used: 242258463
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC148483.6


