

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148483.13 + phase: 0
(63 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
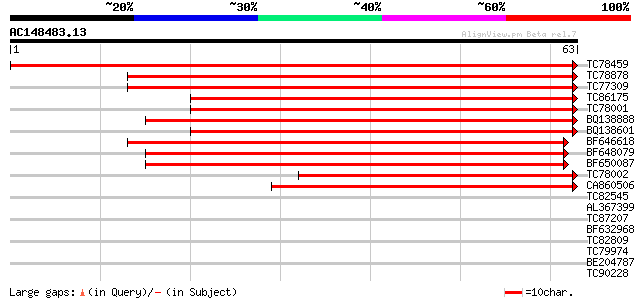
TC78459 similar to GP|12643061|gb|AAK00450.1 unknown protein {Or... 134 5e-33
TC78878 similar to GP|4519792|dbj|BAA75744.1 Asp1 {Arabidopsis t... 78 6e-16
TC77309 similar to GP|4519792|dbj|BAA75744.1 Asp1 {Arabidopsis t... 77 1e-15
TC86175 similar to GP|23617209|dbj|BAC20880. contains ESTs C2856... 70 2e-13
TC78001 similar to GP|21618169|gb|AAM67219.1 ARF GAP-like zinc f... 69 4e-13
BQ138888 weakly similar to PIR|S47006|S470 zinc finger protein G... 69 5e-13
BQ138601 similar to GP|23617209|db contains ESTs C28562(C61610) ... 65 7e-12
BF646618 similar to GP|21594052|gb putative GTPase activating pr... 62 4e-11
BF648079 similar to GP|6648206|gb|A putative GTPase activating p... 61 1e-10
BF650087 similar to GP|6648206|gb|A putative GTPase activating p... 61 1e-10
TC78002 homologue to GP|23617209|dbj|BAC20880. contains ESTs C28... 53 3e-08
CA860506 similar to GP|12697977|dbj KIAA1716 protein {Homo sapie... 50 2e-07
TC82545 similar to GP|17064858|gb|AAL32583.1 putative protein {A... 37 0.001
AL367399 similar to GP|17064858|gb| putative protein {Arabidopsi... 37 0.002
TC87207 similar to GP|9802557|gb|AAF99759.1| F22O13.16 {Arabidop... 36 0.003
BF632968 weakly similar to GP|9858772|gb| BAC19.4 {Lycopersicon ... 27 1.6
TC82809 26 2.7
TC79974 similar to PIR|T48577|T48577 hypothetical protein T31B5.... 26 3.5
BE204787 25 5.9
TC90228 similar to PIR|T47675|T47675 glycoprotein-like - Arabido... 25 7.7
>TC78459 similar to GP|12643061|gb|AAK00450.1 unknown protein {Oryza
sativa}, partial (67%)
Length = 1619
Score = 134 bits (338), Expect = 5e-33
Identities = 63/63 (100%), Positives = 63/63 (100%)
Frame = +2
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS
Sbjct: 194 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 373
Query: 61 FVR 63
FVR
Sbjct: 374 FVR 382
>TC78878 similar to GP|4519792|dbj|BAA75744.1 Asp1 {Arabidopsis thaliana},
partial (53%)
Length = 1839
Score = 78.2 bits (191), Expect = 6e-16
Identities = 31/50 (62%), Positives = 41/50 (82%)
Frame = +2
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
R+L+++ NK C DC+ KNP WASV+YG+F+C++CS HR LGVHISFVR
Sbjct: 113 RELQSEPSNKICVDCSQKNPQWASVSYGVFMCLECSGKHRGLGVHISFVR 262
>TC77309 similar to GP|4519792|dbj|BAA75744.1 Asp1 {Arabidopsis thaliana},
partial (77%)
Length = 2042
Score = 77.0 bits (188), Expect = 1e-15
Identities = 32/50 (64%), Positives = 40/50 (80%)
Frame = +2
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
R L+++ NK C DC+ KNP WASV+YGIF+C++CS HR LGVHISFVR
Sbjct: 176 RDLQSQPGNKICVDCSQKNPQWASVSYGIFMCLECSGKHRGLGVHISFVR 325
>TC86175 similar to GP|23617209|dbj|BAC20880. contains ESTs C28562(C61610)
C25917(C11090)~similar to ARF GAP-like zinc
finger-containing protein, partial (47%)
Length = 1922
Score = 69.7 bits (169), Expect = 2e-13
Identities = 29/43 (67%), Positives = 33/43 (76%)
Frame = +2
Query: 21 ENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
EN+ C DC AK P WASV GIF+C+ CS +HRSLGVHIS VR
Sbjct: 218 ENRECADCKAKGPRWASVNLGIFICMSCSGIHRSLGVHISKVR 346
>TC78001 similar to GP|21618169|gb|AAM67219.1 ARF GAP-like zinc
finger-containing protein ZIGA3 {Arabidopsis thaliana},
partial (46%)
Length = 1265
Score = 68.9 bits (167), Expect = 4e-13
Identities = 29/43 (67%), Positives = 33/43 (76%)
Frame = +1
Query: 21 ENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
EN+ C DC AK P WASV GIF+C+ CS +HRSLGVHIS VR
Sbjct: 301 ENRECADCKAKAPRWASVNLGIFICMQCSGIHRSLGVHISKVR 429
>BQ138888 weakly similar to PIR|S47006|S470 zinc finger protein GCS1 - yeast
(Saccharomyces cerevisiae), partial (25%)
Length = 665
Score = 68.6 bits (166), Expect = 5e-13
Identities = 26/48 (54%), Positives = 35/48 (72%)
Frame = +3
Query: 16 LKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
++ ++N C DCNA +P WAS GIF+C+ CS +HR LGVHISF+R
Sbjct: 126 IQKTNDNNKCVDCNAPSPQWASPKLGIFMCLSCSGIHRGLGVHISFIR 269
>BQ138601 similar to GP|23617209|db contains ESTs C28562(C61610)
C25917(C11090)~similar to ARF GAP-like zinc
finger-containing protein, partial (22%)
Length = 651
Score = 64.7 bits (156), Expect = 7e-12
Identities = 27/43 (62%), Positives = 32/43 (73%)
Frame = +2
Query: 21 ENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
+N+ C DC K P WASV GIF+C+ CS +HRSLGVHIS VR
Sbjct: 215 DNRECADCWTKAPRWASVNLGIFICMQCSGIHRSLGVHISKVR 343
>BF646618 similar to GP|21594052|gb putative GTPase activating protein
{Arabidopsis thaliana}, partial (38%)
Length = 652
Score = 62.0 bits (149), Expect = 4e-11
Identities = 25/49 (51%), Positives = 33/49 (67%)
Frame = +1
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFV 62
+ L + +N+ C DCNA +P WAS G+F+C+ C VHRSLG HIS V
Sbjct: 259 KDLLHQKDNRVCSDCNAPDPKWASANIGVFICLKCCGVHRSLGTHISKV 405
>BF648079 similar to GP|6648206|gb|A putative GTPase activating protein
{Arabidopsis thaliana}, partial (19%)
Length = 582
Score = 60.8 bits (146), Expect = 1e-10
Identities = 25/47 (53%), Positives = 32/47 (67%)
Frame = +2
Query: 16 LKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFV 62
L ++ NK C DC + P W S + G+F+CI CS +HRSLGVHIS V
Sbjct: 332 LMRQAGNKFCADCGSSEPKWVSSSLGVFICIKCSGIHRSLGVHISKV 472
>BF650087 similar to GP|6648206|gb|A putative GTPase activating protein
{Arabidopsis thaliana}, partial (21%)
Length = 589
Score = 60.8 bits (146), Expect = 1e-10
Identities = 25/47 (53%), Positives = 32/47 (67%)
Frame = +3
Query: 16 LKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFV 62
L ++ NK C DC + P W S + G+F+CI CS +HRSLGVHIS V
Sbjct: 372 LMRQAGNKFCADCGSSEPKWVSSSLGVFICIKCSGIHRSLGVHISKV 512
>TC78002 homologue to GP|23617209|dbj|BAC20880. contains ESTs C28562(C61610)
C25917(C11090)~similar to ARF GAP-like zinc
finger-containing protein, partial (26%)
Length = 737
Score = 52.8 bits (125), Expect = 3e-08
Identities = 22/31 (70%), Positives = 25/31 (79%)
Frame = +3
Query: 33 PTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
P WASV GIF+C+ CS +HRSLGVHIS VR
Sbjct: 468 PRWASVNLGIFICMQCSGIHRSLGVHISKVR 560
>CA860506 similar to GP|12697977|dbj KIAA1716 protein {Homo sapiens},
partial (7%)
Length = 362
Score = 50.1 bits (118), Expect = 2e-07
Identities = 19/34 (55%), Positives = 26/34 (75%)
Frame = +3
Query: 30 AKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
+K+P WAS + LCI+CS +HRSLGVH+S V+
Sbjct: 6 SKDPQWASTNFSTLLCIECSGIHRSLGVHVSKVK 107
>TC82545 similar to GP|17064858|gb|AAL32583.1 putative protein {Arabidopsis
thaliana}, partial (22%)
Length = 688
Score = 37.4 bits (85), Expect = 0.001
Identities = 13/40 (32%), Positives = 21/40 (52%)
Frame = +2
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHR 53
R L N+ C +CN+ P + + F+C +CS +HR
Sbjct: 200 RTLLKLEPNRRCINCNSLGPQYVCTNFWTFVCTNCSGIHR 319
>AL367399 similar to GP|17064858|gb| putative protein {Arabidopsis thaliana},
partial (15%)
Length = 461
Score = 37.0 bits (84), Expect = 0.002
Identities = 13/40 (32%), Positives = 21/40 (52%)
Frame = +1
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHR 53
R L + N+ C +CN+ P + + F+C CS +HR
Sbjct: 217 RGLLKLTPNRRCINCNSMGPQYVCTNFWTFVCTKCSGIHR 336
>TC87207 similar to GP|9802557|gb|AAF99759.1| F22O13.16 {Arabidopsis
thaliana}, partial (25%)
Length = 617
Score = 36.2 bits (82), Expect = 0.003
Identities = 13/40 (32%), Positives = 21/40 (52%)
Frame = +2
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAV 51
+ R L N+ C +CN+ P + T+ F+CI CS +
Sbjct: 362 IIRGLMKLPPNRRCINCNSLGPQYVCTTFWTFVCITCSGI 481
>BF632968 weakly similar to GP|9858772|gb| BAC19.4 {Lycopersicon esculentum},
partial (56%)
Length = 644
Score = 26.9 bits (58), Expect = 1.6
Identities = 13/26 (50%), Positives = 17/26 (65%), Gaps = 2/26 (7%)
Frame = -2
Query: 14 RKLKTKSE--NKSCFDCNAKNPTWAS 37
+KL TK+E SC+DC +PT AS
Sbjct: 385 QKLPTKTEVIQWSCYDCECTHPTCAS 308
>TC82809
Length = 691
Score = 26.2 bits (56), Expect = 2.7
Identities = 15/39 (38%), Positives = 22/39 (55%), Gaps = 3/39 (7%)
Frame = +3
Query: 20 SENKSCFDCNA---KNPTWASVTYGIFLCIDCSAVHRSL 55
S+ K F N+ KNP + S+T+ + I C A +RSL
Sbjct: 27 SKTKPLFHFNSIPKKNPNFLSLTFDLVQFIICDANNRSL 143
>TC79974 similar to PIR|T48577|T48577 hypothetical protein T31B5.120 -
Arabidopsis thaliana, partial (61%)
Length = 1835
Score = 25.8 bits (55), Expect = 3.5
Identities = 12/43 (27%), Positives = 21/43 (47%), Gaps = 9/43 (20%)
Frame = +1
Query: 16 LKTKSENKSCFDCNAK---------NPTWASVTYGIFLCIDCS 49
+K K++ K C + + P WAS+ G+ +CI+ S
Sbjct: 1666 IKVKNQLKCCEELSGMINVLIVGHPEPDWASLNLGVLVCIEXS 1794
>BE204787
Length = 579
Score = 25.0 bits (53), Expect = 5.9
Identities = 9/23 (39%), Positives = 15/23 (65%)
Frame = -1
Query: 9 KNAVFRKLKTKSENKSCFDCNAK 31
K+ + ++KT++E SC DC K
Sbjct: 96 KDLNYMEMKTRTET*SCLDCRPK 28
>TC90228 similar to PIR|T47675|T47675 glycoprotein-like - Arabidopsis
thaliana, complete
Length = 1312
Score = 24.6 bits (52), Expect = 7.7
Identities = 13/40 (32%), Positives = 19/40 (47%)
Frame = +1
Query: 18 TKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGV 57
T+ + +C+ NPTW S+ Y C C +R L V
Sbjct: 796 TEESSWTCWRRWISNPTWLSLQY----CYLCKLYNRDLPV 903
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.132 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,424,673
Number of Sequences: 36976
Number of extensions: 31032
Number of successful extensions: 255
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 254
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 255
length of query: 63
length of database: 9,014,727
effective HSP length: 39
effective length of query: 24
effective length of database: 7,572,663
effective search space: 181743912
effective search space used: 181743912
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 51 (24.3 bits)
Medicago: description of AC148483.13


