

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148482.2 + phase: 0
(530 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
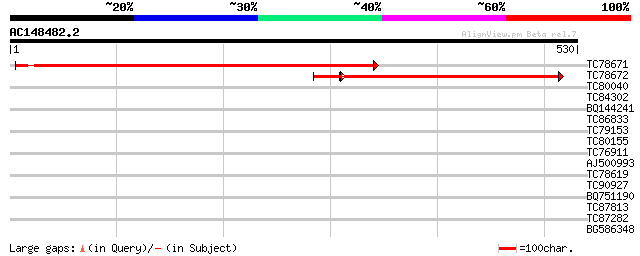
Score E
Sequences producing significant alignments: (bits) Value
TC78671 similar to GP|21536650|gb|AAM60982.1 sodium-dicarboxylat... 522 e-148
TC78672 similar to GP|21536650|gb|AAM60982.1 sodium-dicarboxylat... 352 e-105
TC80040 similar to GP|7621712|gb|AAF65155.1| 2C-methyl-D-erythri... 33 0.27
TC84302 similar to PIR|E96797|E96797 hypothetical protein F7O12.... 30 1.8
BQ144241 similar to PIR|S61505|S615 UDPglucose--starch glucosylt... 29 3.9
TC86833 weakly similar to PIR|H86427|H86427 unknown protein [imp... 29 3.9
TC79153 similar to SP|P28583|CDPK_SOYBN Calcium-dependent protei... 29 5.1
TC80155 similar to PIR|T07821|T07821 Ca2+/H+-exchanging protein ... 29 5.1
TC76911 similar to GP|13543234|gb|AAH05782.1 Unknown (protein fo... 29 5.1
AJ500993 similar to GP|5514645|emb cytochrome P450 {Cicer arieti... 28 6.7
TC78619 similar to GP|6552731|gb|AAF16530.1| T26F17.9 {Arabidops... 28 8.7
TC90927 similar to GP|9279712|dbj|BAB01269.1 cell division prote... 28 8.7
BQ751190 homologue to PIR|A72706|A727 hypothetical protein APE10... 28 8.7
TC87813 similar to SP|P49299|CYSZ_CUCMA Citrate synthase glyoxy... 28 8.7
TC87282 similar to GP|21553648|gb|AAM62741.1 Ser Thr specific pr... 28 8.7
BG586348 similar to GP|20465587|gb unknown protein {Arabidopsis ... 28 8.7
>TC78671 similar to GP|21536650|gb|AAM60982.1 sodium-dicarboxylate
cotransporter-like {Arabidopsis thaliana}, partial (50%)
Length = 1083
Score = 522 bits (1344), Expect = e-148
Identities = 253/340 (74%), Positives = 290/340 (84%), Gaps = 1/340 (0%)
Frame = +1
Query: 6 SLNATLLPLQEPIQETPNNNSITIFSILNLQNFYIILGPLLSIIICLLVKLDAPTTSTK- 64
+L A LLPL +N + S L NFY++LGPLLS+ ICL VKLD P +++
Sbjct: 79 NLKAPLLPLHRT-----QSNPFNLKSFFTLNNFYVLLGPLLSLFICLFVKLDTPNNNSRN 243
Query: 65 MLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIASADTVAHSYMDDVITLVLGSFILA 124
ML VIAWVF+WW+T+A+PLPVTSMCPLFLFPLFGIASADTVAHSYMDDVITLVLGSFILA
Sbjct: 244 MLAVIAWVFSWWVTSALPLPVTSMCPLFLFPLFGIASADTVAHSYMDDVITLVLGSFILA 423
Query: 125 LAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCATTFFVSMWLHNVATAVMMMPVATG 184
LAVERYNVHRRLAL VT VFCG+PLNP++LLLGLC T+FFVSMW+HNVA AVM+MPVATG
Sbjct: 424 LAVERYNVHRRLALTVTSVFCGEPLNPALLLLGLCGTSFFVSMWMHNVAAAVMLMPVATG 603
Query: 185 ILHRLPPAEEQPELMNKFSRAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKP 244
+L RLPP EQ EL+NKFSRAV+LTVVYA PIGG+STLTGTGVNLI+IGMWKSL P AKP
Sbjct: 604 LLQRLPPPSEQSELVNKFSRAVVLTVVYAVPIGGMSTLTGTGVNLILIGMWKSLVPGAKP 783
Query: 245 ISFNSWFFFGFPVAVLLLLCFWCILCLIYVRKGSSRALSDYLDRALLKRDLEALGPMSFA 304
ISFN+WFFF FPVA++ L+CFWCILCLIY+RKGS+ ALS YL + LKR+LEALGP+SFA
Sbjct: 784 ISFNTWFFFAFPVAIVFLVCFWCILCLIYLRKGSASALSPYLSKGHLKRELEALGPISFA 963
Query: 305 EKMVLCVFGLLIVLWMTRRISDDLPGWGVLFHGLVGDGSI 344
EKMVL VFGLLI+LWMTRRI+DD+PG G GLVGDGS+
Sbjct: 964 EKMVLSVFGLLIILWMTRRITDDIPGXGXXXXGLVGDGSV 1083
>TC78672 similar to GP|21536650|gb|AAM60982.1 sodium-dicarboxylate
cotransporter-like {Arabidopsis thaliana}, partial (43%)
Length = 1062
Score = 352 bits (903), Expect(2) = e-105
Identities = 171/209 (81%), Positives = 191/209 (90%)
Frame = +1
Query: 309 LCVFGLLIVLWMTRRISDDLPGWGVLFHGLVGDGSISVLAAVLLFIIPNMKQNGEKLMDW 368
LC+ G I+LWMTRRI+DD+PGWG FHGLVGDGS+SV+ AVLLFIIPN KQ GE+LMDW
Sbjct: 82 LCL-GCXIILWMTRRITDDIPGWGTFFHGLVGDGSVSVMVAVLLFIIPNRKQEGERLMDW 258
Query: 369 NECKKLPWNLILLLGAGFALADGVQSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIIT 428
NECKKLPWNLILLLGAGFALADGVQSSGLADV+SKAL+FL+D PYL I PAVSL+ SIIT
Sbjct: 259 NECKKLPWNLILLLGAGFALADGVQSSGLADVLSKALDFLEDAPYLQIAPAVSLISSIIT 438
Query: 429 EFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGH 488
EFITSNDATATL++PLLYHIA MHVHPLLLMIPG IATEFAF LPTSTPSNVVGFATGH
Sbjct: 439 EFITSNDATATLIIPLLYHIARTMHVHPLLLMIPGAIATEFAFLLPTSTPSNVVGFATGH 618
Query: 489 IEIKDMLKVGVPLKVAGVSVLALLMPTLG 517
IEI+D LKVG+PLKVAG++VL++ MP+LG
Sbjct: 619 IEIQDTLKVGLPLKVAGIAVLSVFMPSLG 705
Score = 48.1 bits (113), Expect(2) = e-105
Identities = 23/30 (76%), Positives = 26/30 (86%)
Frame = +3
Query: 285 YLDRALLKRDLEALGPMSFAEKMVLCVFGL 314
YL + LKR+LEALGP+SFAEKMVL VFGL
Sbjct: 6 YLSKGHLKRELEALGPISFAEKMVLSVFGL 95
>TC80040 similar to GP|7621712|gb|AAF65155.1| 2C-methyl-D-erythritol 2
4-cyclodiphosphate synthase {Catharanthus roseus},
partial (69%)
Length = 755
Score = 33.1 bits (74), Expect = 0.27
Identities = 28/99 (28%), Positives = 44/99 (44%), Gaps = 5/99 (5%)
Frame = +1
Query: 415 AIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIAT-----EF 469
A V A S T +S+ + PL +H +++ P L + P G AT E
Sbjct: 1 ARVTASSYSFPFTTSLKSSHPIPLSCSFPLKHH-TLHLKSSPSLFVSPSGAATPTTSIEI 177
Query: 470 AFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSV 508
++TPS V+ F GH L+ G PL + G+++
Sbjct: 178 DKSPISATPSRVLPFRVGHGFDLHRLEPGYPLIIGGINI 294
>TC84302 similar to PIR|E96797|E96797 hypothetical protein F7O12.4
[imported] - Arabidopsis thaliana, partial (7%)
Length = 832
Score = 30.4 bits (67), Expect = 1.8
Identities = 14/31 (45%), Positives = 19/31 (61%), Gaps = 3/31 (9%)
Frame = +2
Query: 246 SFNSWFFFGF---PVAVLLLLCFWCILCLIY 273
SF F+F F P+ + LLLCFW I+ +Y
Sbjct: 50 SFEEGFYFQFSNDPLNIFLLLCFWGIIFELY 142
>BQ144241 similar to PIR|S61505|S615 UDPglucose--starch glucosyltransferase
(EC 2.4.1.11) isoform II precursor - garden pea, partial
(4%)
Length = 753
Score = 29.3 bits (64), Expect = 3.9
Identities = 14/39 (35%), Positives = 22/39 (55%)
Frame = +3
Query: 104 TVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTL 142
T+ H Y+D VITLVL +E N H++ L++ +
Sbjct: 105 TIFHHYIDHVITLVLTLLCYFFTLEISNSHQKSTLSMLI 221
>TC86833 weakly similar to PIR|H86427|H86427 unknown protein [imported] -
Arabidopsis thaliana, partial (90%)
Length = 2640
Score = 29.3 bits (64), Expect = 3.9
Identities = 13/61 (21%), Positives = 29/61 (47%)
Frame = -3
Query: 17 PIQETPNNNSITIFSILNLQNFYIILGPLLSIIICLLVKLDAPTTSTKMLGVIAWVFTWW 76
P+ E N++ + +FS NL + +++ I +L + T +K ++ W +W
Sbjct: 1225 PV*EKDNSSILLLFSESNLLLGFQFRNQFINLFIAVLYAIHLFTNKSKAASLVGWSCAFW 1046
Query: 77 I 77
+
Sbjct: 1045 L 1043
>TC79153 similar to SP|P28583|CDPK_SOYBN Calcium-dependent protein kinase
SK5 (EC 2.7.1.-) (CDPK). [Soybean] {Glycine max},
partial (95%)
Length = 1751
Score = 28.9 bits (63), Expect = 5.1
Identities = 15/39 (38%), Positives = 22/39 (55%), Gaps = 8/39 (20%)
Frame = -1
Query: 254 GFPVAVLLLLCFWC--------ILCLIYVRKGSSRALSD 284
G P+ VL++LC+WC +LCL++ GS R D
Sbjct: 119 GEPMWVLVVLCWWCWISTFFNVVLCLVF---GSEREKED 12
>TC80155 similar to PIR|T07821|T07821 Ca2+/H+-exchanging protein - mung
bean, partial (35%)
Length = 850
Score = 28.9 bits (63), Expect = 5.1
Identities = 11/27 (40%), Positives = 16/27 (58%), Gaps = 2/27 (7%)
Frame = +3
Query: 245 ISFNSWFF--FGFPVAVLLLLCFWCIL 269
+ F W+F P + LLLC+WC+L
Sbjct: 384 LHFTGWYFSLLERPYSPTLLLCYWCLL 464
>TC76911 similar to GP|13543234|gb|AAH05782.1 Unknown (protein for
MGC:12025) {Mus musculus}, partial (57%)
Length = 1065
Score = 28.9 bits (63), Expect = 5.1
Identities = 21/75 (28%), Positives = 36/75 (48%), Gaps = 4/75 (5%)
Frame = -3
Query: 11 LLPLQEPIQETPNNNSITIFSILNLQNFYIILGPLLSIIICLLVKLDAPTTSTKMLGVI- 69
LLPL EP+ + T+F+ L++ F ++ L++++ V T+T + +
Sbjct: 691 LLPLLEPMLAS------TMFAKLSMLEFAMLSPLELAMLLEFSVAHVTHVTATMCIAALW 530
Query: 70 ---AWVFTWWITNAV 81
WV WWI AV
Sbjct: 529 VSALWVAAWWIMAAV 485
>AJ500993 similar to GP|5514645|emb cytochrome P450 {Cicer arietinum},
partial (27%)
Length = 608
Score = 28.5 bits (62), Expect = 6.7
Identities = 16/32 (50%), Positives = 20/32 (62%)
Frame = +2
Query: 23 NNNSITIFSILNLQNFYIILGPLLSIIICLLV 54
NNN ++I SILN + YI L+ I I LLV
Sbjct: 107 NNNIVSILSILNNLSKYIQYTMLVEIAIALLV 202
>TC78619 similar to GP|6552731|gb|AAF16530.1| T26F17.9 {Arabidopsis
thaliana}, complete
Length = 1428
Score = 28.1 bits (61), Expect = 8.7
Identities = 9/41 (21%), Positives = 22/41 (52%)
Frame = +3
Query: 349 AVLLFIIPNMKQNGEKLMDWNECKKLPWNLILLLGAGFALA 389
A ++ ++P M G +++W PW+ ++++ + LA
Sbjct: 717 ATMILVLPAMLLEGNGVLEWLNTHPYPWSALIIIFSSGVLA 839
>TC90927 similar to GP|9279712|dbj|BAB01269.1 cell division protein
FtsH-like {Arabidopsis thaliana}, partial (28%)
Length = 984
Score = 28.1 bits (61), Expect = 8.7
Identities = 7/26 (26%), Positives = 20/26 (76%)
Frame = +3
Query: 244 PISFNSWFFFGFPVAVLLLLCFWCIL 269
P +F ++ F+ + + ++++LCF+C++
Sbjct: 891 PANFGNFLFYAYLMYLIVVLCFFCLI 968
>BQ751190 homologue to PIR|A72706|A727 hypothetical protein APE1064 -
Aeropyrum pernix (strain K1), partial (4%)
Length = 808
Score = 28.1 bits (61), Expect = 8.7
Identities = 14/45 (31%), Positives = 21/45 (46%), Gaps = 2/45 (4%)
Frame = -2
Query: 44 PLLSIIICLLVKLDAPTTSTKML--GVIAWVFTWWITNAVPLPVT 86
P SI+ C LD+ + ++ L G I W +WW +P T
Sbjct: 540 PSTSILSCSTSPLDSSSAMSRFLLTGTIGWGASWWSGPGMPFART 406
>TC87813 similar to SP|P49299|CYSZ_CUCMA Citrate synthase glyoxysomal
precursor (EC 4.1.3.7) (GCS). [Pumpkin Winter squash],
partial (57%)
Length = 1235
Score = 28.1 bits (61), Expect = 8.7
Identities = 13/36 (36%), Positives = 18/36 (49%)
Frame = +1
Query: 180 PVATGILHRLPPAEEQPELMNKFSRAVILTVVYATP 215
P+ G LHR E P LMN R+++L + P
Sbjct: 301 PLPLGSLHRPMSKERSPLLMNVLGRSIVLKSLLMAP 408
>TC87282 similar to GP|21553648|gb|AAM62741.1 Ser Thr specific protein
kinase-like protein {Arabidopsis thaliana}, partial
(27%)
Length = 1338
Score = 28.1 bits (61), Expect = 8.7
Identities = 13/26 (50%), Positives = 17/26 (65%)
Frame = +3
Query: 247 FNSWFFFGFPVAVLLLLCFWCILCLI 272
FN++FFF P LLLL F C C++
Sbjct: 213 FNTFFFFLAPPPHLLLLRFSCASCVL 290
>BG586348 similar to GP|20465587|gb unknown protein {Arabidopsis thaliana},
partial (33%)
Length = 755
Score = 28.1 bits (61), Expect = 8.7
Identities = 16/39 (41%), Positives = 25/39 (64%)
Frame = -1
Query: 120 SFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGL 158
SFIL L+ +YN+ ++T V GD LNP++++L L
Sbjct: 689 SFILYLS--QYNLTCFWISSLTFVLIGDELNPAIVILAL 579
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.327 0.142 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,714,389
Number of Sequences: 36976
Number of extensions: 299145
Number of successful extensions: 2588
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 2544
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2584
length of query: 530
length of database: 9,014,727
effective HSP length: 101
effective length of query: 429
effective length of database: 5,280,151
effective search space: 2265184779
effective search space used: 2265184779
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148482.2


