

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148470.10 - phase: 0 /pseudo
(598 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
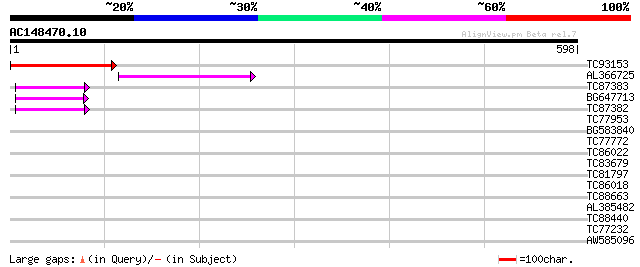
Score E
Sequences producing significant alignments: (bits) Value
TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Ci... 206 3e-53
AL366725 80 3e-15
TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA PO... 45 8e-05
BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent vir... 45 1e-04
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 43 4e-04
TC77953 similar to PIR|D84890|D84890 probable AT-hook DNA-bindin... 35 0.11
BG583840 GP|20146556|gb ATP synthase alpha subunit {Clethra barb... 32 0.69
TC77772 similar to GP|22136300|gb|AAM91228.1 unknown protein {Ar... 32 0.69
TC86022 homologue to GP|19423939|gb|AAL87312.1 putative RNA heli... 32 0.91
TC83679 weakly similar to PIR|G75403|G75403 DNA topoisomerase I ... 31 1.2
TC81797 similar to GP|17065224|gb|AAL32766.1 Unknown protein {Ar... 30 2.0
TC86018 similar to GP|19423939|gb|AAL87312.1 putative RNA helica... 30 3.4
TC88663 similar to GP|8778473|gb|AAF79481.1| F1L3.23 {Arabidopsi... 29 4.5
AL385482 similar to GP|5106924|gb|A putative cell wall protein {... 29 5.9
TC88440 weakly similar to GP|7293965|gb|AAF49324.1| Eip74EF gene... 29 5.9
TC77232 similar to SP|O04130|SERA_ARATH D-3-phosphoglycerate deh... 29 5.9
AW585096 similar to GP|11125147|emb MHC class I chain-related pr... 28 7.7
>TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Cicer
arietinum}, partial (8%)
Length = 516
Score = 206 bits (523), Expect = 3e-53
Identities = 94/112 (83%), Positives = 104/112 (91%)
Frame = +2
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLRNHPPTFKGRY PDGA KWLKE+E IFRVMQC E QKV+FGTHMLAEEA+DWW+SLLP
Sbjct: 116 FLRNHPPTFKGRYAPDGA*KWLKEIERIFRVMQCFETQKVQFGTHMLAEEADDWWISLLP 295
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEF 112
+LEQDDA +TWA+FR+EFL RY PEDVRG+KEIEFLELKQGDMSVTEYAA+F
Sbjct: 296 VLEQDDAVVTWAMFRKEFLGRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKF 451
>AL366725
Length = 485
Score = 79.7 bits (195), Expect = 3e-15
Identities = 66/145 (45%), Positives = 81/145 (55%)
Frame = +3
Query: 115 SSSLKMV*ELISSGPLVIKR**IFLIW*AVVGYMKRIPKLITR**VSAVARDSRVVRSRI 174
+SS +MV* S G LI * +RI +L+TR *+S R +RV S I
Sbjct: 48 ASSSRMV*GPTSRGQSDTNSSEFSLI**IPAESTRRILRLMTRL*MSGRPRANRVALSLI 227
Query: 175 VPLRTRVNRG*LMRGGLRGEMLLLRLFVSSVAKKATRVMYVTVMRRSISDVDRNDTH*LI 234
VPL R NR M G LR MLL RL VS++A+KATRVM+V R++S V R *LI
Sbjct: 228 VPLLIRANREWSMIGVLRRRMLLRRLSVSTMARKATRVMFVLKRSRNVSGVTRRVIL*LI 407
Query: 235 ARVGMLFAIIVMKRGILAHSARSLR 259
A +L A I KR IL +A SLR
Sbjct: 408 ASEMILCASIATKRVILVPNASSLR 482
>TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA POLYMERASE II
{Encephalitozoon cuniculi}, partial (0%)
Length = 1247
Score = 45.1 bits (105), Expect = 8e-05
Identities = 22/80 (27%), Positives = 41/80 (50%), Gaps = 2/80 (2%)
Frame = -2
Query: 7 PTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLPILEQDD 66
P F+G PD WL+ +E +F+ + E QKV+ L + A+ WW ++ +++
Sbjct: 724 PDFEGNLQPDDLLDWLQIMERLFKYKEVLEEQKVKIVAAKLKKLASIWWENVKRRRKREG 545
Query: 67 AA--MTWAVFRREFLNRYVP 84
+ TW R++ +Y+P
Sbjct: 544 KSKIKTWEKMRQKLTRKYLP 485
>BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent virus}, partial
(1%)
Length = 726
Score = 44.7 bits (104), Expect = 1e-04
Identities = 23/79 (29%), Positives = 38/79 (47%), Gaps = 2/79 (2%)
Frame = +2
Query: 7 PTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSL--LPILEQ 64
P F G+ PD WL+ +E +F+ + +E QKV+ L + A+ WW +L E
Sbjct: 347 PDF*GKLQPDEFVDWLQTIERVFKYKEVAEEQKVKIVAAKLKKHASIWWKNLKRKRNCEG 526
Query: 65 DDAAMTWAVFRREFLNRYV 83
TW R++ +Y+
Sbjct: 527 KSKIKTWDKMRQKLTRKYL 583
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 42.7 bits (99), Expect = 4e-04
Identities = 22/80 (27%), Positives = 38/80 (47%), Gaps = 2/80 (2%)
Frame = +2
Query: 7 PTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLPILEQDD 66
P F+G D WL+ +E +F + E QKV+ L + A WW +L +++
Sbjct: 629 PDFEGNLQLDDFLDWLQTIERVFEYKEVPEEQKVKIVAAKLKKHALIWWENLKRRRKREG 808
Query: 67 AA--MTWAVFRREFLNRYVP 84
+ TW R++ +Y+P
Sbjct: 809 KSKIKTWDKMRQKLTRKYLP 868
>TC77953 similar to PIR|D84890|D84890 probable AT-hook DNA-binding protein
[imported] - Arabidopsis thaliana, partial (57%)
Length = 1695
Score = 34.7 bits (78), Expect = 0.11
Identities = 22/73 (30%), Positives = 41/73 (56%), Gaps = 6/73 (8%)
Frame = +2
Query: 250 ILAHSARSLRRFGQVARYLH*LVRRL*I------RTDLSEVLVSLIVLL*LLL*ILVLLI 303
+L H R F + R+ H L+R L + R D +LV ++++L*LLL +L+L +
Sbjct: 941 LLLHFMEGSRFFRWLDRFFHHLLRPLLLV*PYI*RADKGRLLVEVLLVL*LLLDLLLLCL 1120
Query: 304 VSLLLIVHISWVW 316
+ L+ ++ ++W
Sbjct: 1121 LLLVTLLMRDFLW 1159
>BG583840 GP|20146556|gb ATP synthase alpha subunit {Clethra barbinervis},
partial (17%)
Length = 585
Score = 32.0 bits (71), Expect = 0.69
Identities = 16/35 (45%), Positives = 27/35 (76%)
Frame = +2
Query: 284 VLVSLIVLL*LLL*ILVLLIVSLLLIVHISWVWLY 318
+L+ L++LL LLL +L+LL++ LLL+ ++WLY
Sbjct: 5 LLLLLLLLLLLLLLLLLLLLLLLLLLYIYIYMWLY 109
Score = 29.6 bits (65), Expect = 3.4
Identities = 13/32 (40%), Positives = 27/32 (83%)
Frame = +2
Query: 288 LIVLL*LLL*ILVLLIVSLLLIVHISWVWLYL 319
L++LL LLL +L+LL++ LLL++ + ++++Y+
Sbjct: 5 LLLLLLLLLLLLLLLLLLLLLLLLLLYIYIYM 100
Score = 28.5 bits (62), Expect = 7.7
Identities = 14/35 (40%), Positives = 28/35 (80%)
Frame = +2
Query: 285 LVSLIVLL*LLL*ILVLLIVSLLLIVHISWVWLYL 319
L+ L++LL LLL +L+LL++ LLL++ +++++L
Sbjct: 2 LLLLLLLLLLLLLLLLLLLLLLLLLLLYIYIYMWL 106
>TC77772 similar to GP|22136300|gb|AAM91228.1 unknown protein {Arabidopsis
thaliana}, partial (62%)
Length = 2554
Score = 32.0 bits (71), Expect = 0.69
Identities = 21/59 (35%), Positives = 39/59 (65%), Gaps = 3/59 (5%)
Frame = +2
Query: 284 VLVSLIVLL*LLL*ILVLLIVSLLLIVHISWVWLYLI*MEK---WLLKLQLRVQ*LLLL 339
+L+ LI++L L+L*+++LL++ L I + W L +++ W L+L +R++ LLLL
Sbjct: 530 MLICLIIILGLIL*LVLLLLICL--IC*LRWNNLIFCRIQRVLIWFLRLCIRLERLLLL 700
>TC86022 homologue to GP|19423939|gb|AAL87312.1 putative RNA helicase
{Arabidopsis thaliana}, partial (89%)
Length = 1982
Score = 31.6 bits (70), Expect = 0.91
Identities = 24/53 (45%), Positives = 36/53 (67%), Gaps = 3/53 (5%)
Frame = -2
Query: 285 LVSLIVLL*LLL*ILVLLIVSLLLIVH---ISWVWLYLI*MEKWLLKLQLRVQ 334
L+ L++LL LLL +L++LIV L+LI+H + + L LI M LLK +RV+
Sbjct: 319 LLLLLLLLILLLILLLILIVLLMLILHHVPLHILLLLLILMMWSLLKPGVRVR 161
>TC83679 weakly similar to PIR|G75403|G75403 DNA topoisomerase I -
Deinococcus radiodurans (strain R1), partial (3%)
Length = 596
Score = 31.2 bits (69), Expect = 1.2
Identities = 20/54 (37%), Positives = 34/54 (62%)
Frame = -2
Query: 284 VLVSLIVLL*LLL*ILVLLIVSLLLIVHISWVWLYLI*MEKWLLKLQLRVQ*LL 337
+LV L + L L+L*+ +LL++ L L++H W L+L+ W L L + +Q +L
Sbjct: 301 LLVHLWLWLWLVL*LALLLVLLL*LVLHHQW--LFLLLSRPWRLLLDIFLQRVL 146
>TC81797 similar to GP|17065224|gb|AAL32766.1 Unknown protein {Arabidopsis
thaliana}, partial (17%)
Length = 758
Score = 30.4 bits (67), Expect = 2.0
Identities = 17/43 (39%), Positives = 29/43 (66%)
Frame = -2
Query: 289 IVLL*LLL*ILVLLIVSLLLIVHISWVWLYLI*MEKWLLKLQL 331
+VL+ +++ +L+LL++ LLL+ W+WL+L WLL L L
Sbjct: 493 LVLMAIMMFLLLLLLLVLLLL----WLWLWL-----WLLLLML 392
>TC86018 similar to GP|19423939|gb|AAL87312.1 putative RNA helicase
{Arabidopsis thaliana}, partial (12%)
Length = 885
Score = 29.6 bits (65), Expect = 3.4
Identities = 17/37 (45%), Positives = 29/37 (77%)
Frame = -1
Query: 284 VLVSLIVLL*LLL*ILVLLIVSLLLIVHISWVWLYLI 320
+L+ LI+LL LLL +L++LIV L+LI+ + V L+++
Sbjct: 693 LLLLLILLLILLLILLMMLIVLLMLILILHQVPLHIL 583
>TC88663 similar to GP|8778473|gb|AAF79481.1| F1L3.23 {Arabidopsis
thaliana}, partial (34%)
Length = 946
Score = 29.3 bits (64), Expect = 4.5
Identities = 22/56 (39%), Positives = 37/56 (65%)
Frame = +2
Query: 284 VLVSLIVLL*LLL*ILVLLIVSLLLIVHISWVWLYLI*MEKWLLKLQLRVQ*LLLL 339
+L ++++LL LLL +L+LL++ LLL++ L L+ LL L+LR++ LLL
Sbjct: 515 LLQAMLLLLRLLLLVLLLLLLVLLLLL------LLLLLRSMLLLHLKLRLRFGLLL 664
>AL385482 similar to GP|5106924|gb|A putative cell wall protein {Medicago
truncatula}, partial (42%)
Length = 402
Score = 28.9 bits (63), Expect = 5.9
Identities = 14/32 (43%), Positives = 26/32 (80%)
Frame = -2
Query: 284 VLVSLIVLL*LLL*ILVLLIVSLLLIVHISWV 315
+LV L+V++ LLL +L+L++V LLL+V + ++
Sbjct: 143 LLVLLLVVVLLLLVVLLLVVVLLLLVVELLYL 48
>TC88440 weakly similar to GP|7293965|gb|AAF49324.1| Eip74EF gene product
{Drosophila melanogaster}, partial (5%)
Length = 711
Score = 28.9 bits (63), Expect = 5.9
Identities = 16/41 (39%), Positives = 32/41 (78%)
Frame = -3
Query: 284 VLVSLIVLL*LLL*ILVLLIVSLLLIVHISWVWLYLI*MEK 324
+LV L++L+ LL ++VLL+V LLL++H+ + L+L+ +++
Sbjct: 313 LLVGLLLLVV*LL-LVVLLLVGLLLVLHLLQLVLHLLVLDE 194
>TC77232 similar to SP|O04130|SERA_ARATH D-3-phosphoglycerate dehydrogenase
chloroplast precursor (EC 1.1.1.95) (3-PGDH). [Mouse-ear
cress], partial (87%)
Length = 2474
Score = 28.9 bits (63), Expect = 5.9
Identities = 18/47 (38%), Positives = 33/47 (69%)
Frame = +3
Query: 285 LVSLIVLL*LLL*ILVLLIVSLLLIVHISWVWLYLI*MEKWLLKLQL 331
+ L++ L +LL*+L+LLI+ LLL + + ++ L+L+ K +L L+L
Sbjct: 597 IYKLLLNLVVLL*MLLLLILLLLLNMGLLFLLLWLVTFLKLMLPLKL 737
>AW585096 similar to GP|11125147|emb MHC class I chain-related protein A
{Homo sapiens}, partial (5%)
Length = 231
Score = 28.5 bits (62), Expect = 7.7
Identities = 14/34 (41%), Positives = 25/34 (73%)
Frame = +3
Query: 280 DLSEVLVSLIVLL*LLL*ILVLLIVSLLLIVHIS 313
++ +L++L++LL LLL L LLI+ L+L + I+
Sbjct: 111 EIMRLLLNLLLLLLLLLLFLTLLIILLILTIDIT 212
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.370 0.166 0.609
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,883,866
Number of Sequences: 36976
Number of extensions: 296650
Number of successful extensions: 4239
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 1319
Number of HSP's successfully gapped in prelim test: 185
Number of HSP's that attempted gapping in prelim test: 2710
Number of HSP's gapped (non-prelim): 1697
length of query: 598
length of database: 9,014,727
effective HSP length: 102
effective length of query: 496
effective length of database: 5,243,175
effective search space: 2600614800
effective search space used: 2600614800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148470.10


