

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148469.7 - phase: 0
(540 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
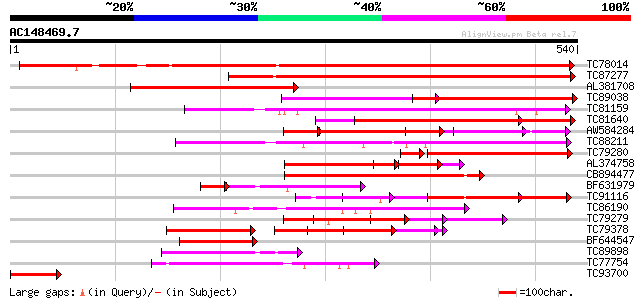
Score E
Sequences producing significant alignments: (bits) Value
TC78014 similar to GP|17529090|gb|AAL38755.1 unknown protein {Ar... 457 e-129
TC87277 similar to PIR|T45588|T45588 arm repeat containing prote... 355 2e-98
AL381708 similar to PIR|E96734|E96 unknown protein F23N20.1 [imp... 323 8e-89
TC89038 similar to PIR|D86364|D86364 hypothetical protein AAB721... 259 2e-69
TC81159 weakly similar to PIR|T45588|T45588 arm repeat containin... 187 1e-47
TC81640 similar to GP|15292863|gb|AAK92802.1 unknown protein {Ar... 172 3e-43
AW584284 similar to PIR|T45588|T45 arm repeat containing protein... 143 2e-38
TC88211 similar to PIR|E96534|E96534 Hypothetical protein [impor... 155 4e-38
TC79280 similar to GP|21592960|gb|AAM64910.1 unknown {Arabidopsi... 121 3e-30
AL374758 weakly similar to PIR|T45588|T45 arm repeat containing ... 114 5e-29
CB894477 weakly similar to GP|10178181|db gene_id:MQN23.13~pir||... 123 2e-28
BF631979 similar to PIR|A86416|A86 hypothetical protein AAG12624... 88 2e-25
TC91116 similar to GP|20161458|dbj|BAB90382. hypothetical protei... 113 2e-25
TC86190 weakly similar to PIR|T08454|T08454 hypothetical protein... 109 2e-24
TC79279 similar to GP|16226454|gb|AAL16172.1 AT3g01400/T13O15_4 ... 109 2e-24
TC79378 similar to PIR|T00518|T00518 hypothetical protein At2g23... 106 3e-23
BF644547 similar to GP|14149112|dbj bg55 {Bruguiera gymnorrhiza}... 102 4e-22
TC89898 similar to GP|19715649|gb|AAL91644.1 AT3g07360/F21O3_7 {... 101 6e-22
TC77754 similar to GP|19911585|dbj|BAB86896. syringolide-induced... 101 8e-22
TC93700 98 9e-21
>TC78014 similar to GP|17529090|gb|AAL38755.1 unknown protein {Arabidopsis
thaliana}, partial (74%)
Length = 2305
Score = 457 bits (1175), Expect = e-129
Identities = 257/537 (47%), Positives = 351/537 (64%), Gaps = 8/537 (1%)
Frame = +3
Query: 10 QFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGFMISKMP------- 62
+FQ++T K+E LS + Y+ L+ISEEV+EQ++LV Q +RA + F ++
Sbjct: 552 KFQQITEKIEAALSDISYNKLEISEEVQEQIELVHAQFKRAKAQTEFADLQLDLDIAVAQ 731
Query: 63 -SFDSSQPLAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPFGAGSRLER 121
D + + +S+ L L + + SEL + ++ G+ + F S L R
Sbjct: 732 KDKDPDPAILKRLSEKLH-----LRTMNDLKKESSELHELVITSNGELGDSFETVSSLLR 896
Query: 122 TRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELMRDPVIVA 181
++ + + S N+P K + VIP+DF CPISLELM+DPVIV+
Sbjct: 897 --------KLKDCVLAENPEVDTYDSENVPI--KHRSPVIPDDFRCPISLELMKDPVIVS 1046
Query: 182 TGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLTN 241
TGQTYERS IQ+W+D G+ TCPKTQQ L H LTPNYVL+SL+ WC + +E P +
Sbjct: 1047 TGQTYERSCIQKWLDAGHRTCPKTQQTLLHTALTPNYVLKSLIGLWCDSNGVELPKKQGS 1226
Query: 242 GKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAG 301
+ KKS S D D AI+ L+ KL+ +E+ RAA E+R L+KR+ DNR+ IAEAG
Sbjct: 1227 CRTKKSGTSLSDC--DKTAIKALLDKLTSNDIEQQRAAAGELRLLAKRNADNRVCIAEAG 1400
Query: 302 AIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEAREN 361
AIP+LV LL+S D TQE+A T++LNLSI E+NKG I+ AGAIP IV VL+ G+MEAREN
Sbjct: 1401 AIPLLVDLLSSTDPRTQEHADTALLNLSINESNKGTIVNAGAIPDIVDVLKNGSMEAREN 1580
Query: 362 AAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAI 421
AAATLFSLS+ DENK+ IGA+GAI AL+ LL G+PRGKKDAATA+FNLCIYQGNK RA+
Sbjct: 1581 AAATLFSLSVLDENKVAIGAAGAIPALIKLLCEGTPRGKKDAATAIFNLCIYQGNKARAV 1760
Query: 422 RAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRN 481
+AGI+ L+ + D+ MVDEAL I+++LA H E + +I +A IP+L++++RTG PRN
Sbjct: 1761 KAGIVAPLIRFMKDAGGGMVDEALAILTILAGHHEGRTAIGQAEPIPILVEVIRTGSPRN 1940
Query: 482 KENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRKLQ 538
+ENAAA+L +C D L GA L L+ GT+RAKRKA S+LE L++++
Sbjct: 1941 RENAAAVLWLVCTGDLLQLKLAKEHGAEEALQGLSENGTDRAKRKAGSILELLQRIE 2111
>TC87277 similar to PIR|T45588|T45588 arm repeat containing protein homolog
- Arabidopsis thaliana, partial (48%)
Length = 1393
Score = 355 bits (911), Expect = 2e-98
Identities = 184/331 (55%), Positives = 241/331 (72%)
Frame = +1
Query: 209 LQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKL 268
L LTPNYVLRSL+ QWC + IE P + + KS + + + IE+L++KL
Sbjct: 1 LTSTVLTPNYVLRSLIEQWCEANGIEPPKRPSTSEPSKSASAC--TPAERSKIESLIQKL 174
Query: 269 SCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNL 328
+ E+ R+A EIR L+KR+ DNR+ IAEAGAIP+LV LL+ D TQE+AVT++LNL
Sbjct: 175 TSGGPEDQRSAAGEIRLLAKRNADNRVAIAEAGAIPLLVGLLSVPDSRTQEHAVTALLNL 354
Query: 329 SIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLADENKIIIGASGAISAL 388
SIYE+NKG I+ +GA+P IV VL+ G+MEARENAAATLFSLS+ DENK+ IG+SGAIS L
Sbjct: 355 SIYESNKGSIVSSGAVPGIVHVLKRGSMEARENAAATLFSLSVIDENKVTIGSSGAISPL 534
Query: 389 VDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIM 448
V LL G+ RGKKDAATALFNLCIYQGNKG+A+RAG+I L+ +LT+ S MVDEAL I+
Sbjct: 535 VTLLSEGTQRGKKDAATALFNLCIYQGNKGKAVRAGVIPTLMRLLTEPSGGMVDEALAIL 714
Query: 449 SVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGA 508
++LASH E K +I A +PVL+ + G PRNKENAAA+L+ LC D ++ LG
Sbjct: 715 AILASHPEGKAAIGAAEAVPVLVKFIGNGSPRNKENAAAVLVHLCSGDRQYIAQAQELGV 894
Query: 509 VIPLSELARTGTERAKRKATSLLEHLRKLQQ 539
+ PL ELA+ GT+R KRKAT L+E + ++++
Sbjct: 895 MAPLLELAQNGTDRGKRKATQLIERMSRIRE 987
>AL381708 similar to PIR|E96734|E96 unknown protein F23N20.1 [imported] -
Arabidopsis thaliana, partial (18%)
Length = 482
Score = 323 bits (829), Expect = 8e-89
Identities = 160/160 (100%), Positives = 160/160 (100%)
Frame = +3
Query: 116 GSRLERTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELMR 175
GSRLERTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELMR
Sbjct: 3 GSRLERTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELMR 182
Query: 176 DPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQ 235
DPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQ
Sbjct: 183 DPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQ 362
Query: 236 PTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEE 275
PTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEE
Sbjct: 363 PTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEE 482
>TC89038 similar to PIR|D86364|D86364 hypothetical protein AAB72157.1
[imported] - Arabidopsis thaliana, partial (19%)
Length = 718
Score = 259 bits (661), Expect = 2e-69
Identities = 142/159 (89%), Positives = 148/159 (92%), Gaps = 2/159 (1%)
Frame = +3
Query: 384 AISALVDLLQNGSPRGKKDAATALFNLCIY-QGNKGRAIRAGIITALLNMLTD-SSKSMV 441
AISALVDLLQNGSPRGKKDAATAL+ +Y +GNKGRAIRAGIITA + + +KSMV
Sbjct: 3 AISALVDLLQNGSPRGKKDAATALYQFSVYIKGNKGRAIRAGIITAPVEHASQIQAKSMV 182
Query: 442 DEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLS 501
DEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLS
Sbjct: 183 DEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLS 362
Query: 502 CISRLGAVIPLSELARTGTERAKRKATSLLEHLRKLQQL 540
CISRLGAVIPLSELARTGTERAKRKATSLLEHLRKLQQL
Sbjct: 363 CISRLGAVIPLSELARTGTERAKRKATSLLEHLRKLQQL 479
Score = 50.8 bits (120), Expect = 1e-06
Identities = 44/153 (28%), Positives = 69/153 (44%), Gaps = 2/153 (1%)
Frame = +3
Query: 260 AIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQ- 318
AI LV L S + A + S N+ AG I V + +
Sbjct: 3 AISALVDLLQNGSPRGKKDAATALYQFSVYIKGNKGRAIRAGIITAPVEHASQIQAKSMV 182
Query: 319 ENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLAD-ENKI 377
+ A+T + L+ ++ K I+ A IP ++ +LR G +ENAAA L +L D +N
Sbjct: 183 DEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLS 362
Query: 378 IIGASGAISALVDLLQNGSPRGKKDAATALFNL 410
I GA+ L +L + G+ R K+ A + L +L
Sbjct: 363 CISRLGAVIPLSELARTGTERAKRKATSLLEHL 461
>TC81159 weakly similar to PIR|T45588|T45588 arm repeat containing protein
homolog - Arabidopsis thaliana, partial (12%)
Length = 1476
Score = 187 bits (474), Expect = 1e-47
Identities = 132/388 (34%), Positives = 204/388 (52%), Gaps = 20/388 (5%)
Frame = +1
Query: 167 CPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQ 226
CPISLELM DPV + TG TY+RS I +W GN+TCPKT + L + L PN VLR L+ Q
Sbjct: 4 CPISLELMSDPVTIETGHTYDRSSILKWFRSGNSTCPKTGKSLGSIELVPNLVLRRLIQQ 183
Query: 227 WCIEHNIEQPTGLTNGKIKKSDGSFRDVT-----GDIAA--IETLVRKLSCRS-----VE 274
+C + I S RD+T G +AA TL+ CRS VE
Sbjct: 184 YCNVNGI---------PFADSSRRSRDITRTVEPGSVAAEGAMTLLAGFLCRSLDNGNVE 336
Query: 275 ESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENN 334
+ A E+R L+K S +R E+G +P+L+ LL S D QENA+ ++LNLS Y +
Sbjct: 337 QKNHAAFEVRVLTKTSIFSRSCFVESGLVPLLLLLLASSDSSAQENAIAALLNLSKYIKS 516
Query: 335 KGLIMLAGAIPSIVQVLRAG-TMEARENAAATLFSLSLADENKIIIGAS-GAISALVDLL 392
+ ++ + IV VL G +EA+++AAA LF L+ E+ +IG AI +L+ L+
Sbjct: 517 RSEMVENWGLEMIVGVLNKGINIEAKQHAAAVLFYLASNPEHANLIGEEPEAIPSLISLI 696
Query: 393 QNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSK-SMVDEALTIMSVL 451
++ + R K+ A+F L N R + A I L+N+L S K +V ++L I++ L
Sbjct: 697 KDDNKRSVKNGLVAIFGLLKNHENHKRILAAQAIPLLVNILKASEKEDLVTDSLAILATL 876
Query: 452 ASHQEAKVSIVKASTIPVLIDLLRTGLPRN---KENAAAILLALCKRDTDNL--SCISRL 506
A + I++ + V ++++ + + KE+ ++LL+L +N+ +
Sbjct: 877 AEKSDGTSEILRFGALHVAVEVMSSSSTTSRLGKEHCVSLLLSLSINGGENVIAHLVKSS 1056
Query: 507 GAVIPLSELARTGTERAKRKATSLLEHL 534
+ L GT RA +KA+SL+ L
Sbjct: 1057SLMESLYSQLSEGTSRASKKASSLIRVL 1140
>TC81640 similar to GP|15292863|gb|AAK92802.1 unknown protein {Arabidopsis
thaliana}, partial (40%)
Length = 810
Score = 172 bits (436), Expect = 3e-43
Identities = 90/211 (42%), Positives = 133/211 (62%)
Frame = +2
Query: 329 SIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLADENKIIIGASGAISAL 388
+++E+NK LI AGA+ S++ VL+ GT +++NAA L SL+L +ENK IGASGAI L
Sbjct: 11 ALHEDNKKLIFNAGAVKSLIYVLKTGTETSKQNAACALLSLALVEENKSSIGASGAIPPL 190
Query: 389 VDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIM 448
V LL NGS RGKKDA T L+ LC + NK RA+ AG++ L+ ++ + M+++A+ ++
Sbjct: 191 VSLLLNGSNRGKKDALTTLYKLCSVKQNKERAVSAGVVKPLVELVAEQGNGMMEKAMVVL 370
Query: 449 SVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGA 508
+ LA E K +IV+ I L++ + G + KE A LL LC N + R G
Sbjct: 371 NSLAGFDEGKEAIVEEGGIAALVEAIEDGSVKGKEFAVLTLLQLCAESVTNRGLLVREGG 550
Query: 509 VIPLSELARTGTERAKRKATSLLEHLRKLQQ 539
+ PL L++ GT RAK KA +LL +LR+ +Q
Sbjct: 551 IPPLVALSQNGTPRAKHKAETLLRYLRESRQ 643
Score = 90.9 bits (224), Expect = 1e-18
Identities = 59/199 (29%), Positives = 101/199 (50%), Gaps = 1/199 (0%)
Frame = +2
Query: 292 DNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVL 351
DN+ LI AGA+ L+ +L + +++NA ++L+L++ E NK I +GAIP +V +L
Sbjct: 23 DNKKLIFNAGAVKSLIYVLKTGTETSKQNAACALLSLALVEENKSSIGASGAIPPLVSLL 202
Query: 352 RAGTMEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLC 411
G+ +++A TL+ L +NK ++G + LV+L+ + A L +L
Sbjct: 203 LNGSNRGKKDALTTLYKLCSVKQNKERAVSAGVVKPLVELVAEQGNGMMEKAMVVLNSLA 382
Query: 412 IYQGNKGRAIRAGIITALLNMLTDSS-KSMVDEALTIMSVLASHQEAKVSIVKASTIPVL 470
+ K + G I AL+ + D S K LT++ + A + +V+ IP L
Sbjct: 383 GFDEGKEAIVEEGGIAALVEAIEDGSVKGKEFAVLTLLQLCAESVTNRGLLVREGGIPPL 562
Query: 471 IDLLRTGLPRNKENAAAIL 489
+ L + G PR K A +L
Sbjct: 563 VALSQNGTPRAKHKAETLL 619
>AW584284 similar to PIR|T45588|T45 arm repeat containing protein homolog -
Arabidopsis thaliana, partial (24%)
Length = 542
Score = 143 bits (361), Expect(2) = 2e-38
Identities = 75/119 (63%), Positives = 95/119 (79%)
Frame = +2
Query: 296 LIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGT 355
L+ EAGA+P+LV LL++ T E+AVT++LNLSIYE+ KG I+ +GA+P I VL G+
Sbjct: 185 LLPEAGAVPLLVGLLSAPYSRTLEHAVTALLNLSIYESIKGSIVSSGAVPRIGHVLNRGS 364
Query: 356 MEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQ 414
+EAR+NAA+ LFSLSL DENK+ IG+ GAIS LV LL G+ RG+KDAATALFNLCIYQ
Sbjct: 365 VEARDNAASALFSLSLIDENKVTIGS*GAISPLVTLLSEGTQRGEKDAATALFNLCIYQ 541
Score = 58.5 bits (140), Expect = 6e-09
Identities = 38/116 (32%), Positives = 58/116 (49%)
Frame = +2
Query: 378 IIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSS 437
++ +GA+ LV LL R + A TAL NL IY+ KG + +G + + ++L S
Sbjct: 185 LLPEAGAVPLLVGLLSAPYSRTLEHAVTALLNLSIYESIKGSIVSSGAVPRIGHVLNRGS 364
Query: 438 KSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAAILLALC 493
D A + + L+ E KV+I I L+ LL G R +++AA L LC
Sbjct: 365 VEARDNAASALFSLSLIDENKVTIGS*GAISPLVTLLSEGTQRGEKDAATALFNLC 532
Score = 50.8 bits (120), Expect = 1e-06
Identities = 33/112 (29%), Positives = 59/112 (52%)
Frame = +2
Query: 423 AGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNK 482
AG + L+ +L+ ++ A+T + L+ ++ K SIV + +P + +L G +
Sbjct: 197 AGAVPLLVGLLSAPYSRTLEHAVTALLNLSIYESIKGSIVSSGAVPRIGHVLNRGSVEAR 376
Query: 483 ENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHL 534
+NAA+ L +L D +N I GA+ PL L GT+R ++ A + L +L
Sbjct: 377 DNAASALFSLSLID-ENKVTIGS*GAISPLVTLLSEGTQRGEKDAATALFNL 529
Score = 35.0 bits (79), Expect = 0.073
Identities = 26/85 (30%), Positives = 44/85 (51%), Gaps = 3/85 (3%)
Frame = +2
Query: 251 FRDVTGDIA---AIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLV 307
+ + G I A+ + L+ SVE A + + SLS +N++ I GAI LV
Sbjct: 290 YESIKGSIVSSGAVPRIGHVLNRGSVEARDNAASALFSLSLID-ENKVTIGS*GAISPLV 466
Query: 308 SLLTSEDVMTQENAVTSILNLSIYE 332
+LL+ +++A T++ NL IY+
Sbjct: 467 TLLSEGTQRGEKDAATALFNLCIYQ 541
Score = 34.3 bits (77), Expect(2) = 2e-38
Identities = 16/38 (42%), Positives = 27/38 (70%)
Frame = +1
Query: 261 IETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIA 298
IE+L++KL+ E+ R++ EIR L++R+ NR+ IA
Sbjct: 79 IESLIQKLTYGGPEDQRSSAGEIRLLAQRNAYNRVSIA 192
>TC88211 similar to PIR|E96534|E96534 Hypothetical protein [imported] -
Arabidopsis thaliana, partial (67%)
Length = 1478
Score = 155 bits (392), Expect = 4e-38
Identities = 117/390 (30%), Positives = 186/390 (47%), Gaps = 13/390 (3%)
Frame = +3
Query: 159 IVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNY 218
+ IP F CPISLELMRDPV V+TGQTY+R+ I+ W++ GNTTCP T+ L T PN+
Sbjct: 150 VQIPYHFRCPISLELMRDPVTVSTGQTYDRNSIESWVNTGNTTCPVTRTNLTDFTFIPNH 329
Query: 219 VLRSLVSQWCIEHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRA 278
LR L+ WC+ + + K D A + +L+ ++S S
Sbjct: 330 TLRRLIQDWCVSNRAFGVQRIPTPK----------QPADAALVRSLLNQISSHSAPTQLR 479
Query: 279 --AVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNK- 335
++ +RSL++ S NR LI+ +++ +L + + +N +++ L ++
Sbjct: 480 LNSLRRLRSLTRDSEYNRSLISSLNVRNIILPILFNNGLDELKNESLALIVLFPLSESEC 659
Query: 336 -GLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLADENK-----IIIGASGAISALV 389
L + I + +L + + R N+AA L + +A + + G +V
Sbjct: 660 TSLASDSDKINYLTSLLSHDSFDVRVNSAA-LIEIIVAGTHSPEIRLQVSNVDGIYDGVV 836
Query: 390 DLLQN--GSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTI 447
++L+N PR K ALF LC+ + + RA+ AG L++ L D K + AL
Sbjct: 837 EILKNPISYPRALKIGIKALFALCLVKQTRHRAVSAGAPVVLIDRLADFEKCDAERALAT 1016
Query: 448 MSVLASHQEAKVSIV-KASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISRL 506
+ +L S A T+P+L+ ++ R E AA L+ALC
Sbjct: 1017VELLCRVPAGCASFAGHALTVPMLVKIILKISDRATEYAAGALMALCSESERCQRETVAA 1196
Query: 507 GAVIPLSELARTG-TERAKRKATSLLEHLR 535
G + L L ++ TERAKRKA LL+ LR
Sbjct: 1197GVLTQLLLLVQSDCTERAKRKAQLLLKLLR 1286
>TC79280 similar to GP|21592960|gb|AAM64910.1 unknown {Arabidopsis
thaliana}, partial (46%)
Length = 809
Score = 121 bits (304), Expect(2) = 3e-30
Identities = 62/138 (44%), Positives = 94/138 (67%)
Frame = +2
Query: 399 GKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAK 458
GKKDA+TAL+ LC + NK RA++AGI+ L+ ++ D +MVD++ ++SVL S EAK
Sbjct: 83 GKKDASTALYTLCSVKENKMRAVKAGIMKVLVELMADFESNMVDKSAYVLSVLVSVPEAK 262
Query: 459 VSIVKASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELART 518
V++V+ +PVL++++ G R KE AA ILL +C+ S ++R GA+ PL L ++
Sbjct: 263 VALVEEGGVPVLVEIVEVGSQRQKEIAAVILLQICEDSVAVRSMVAREGAIPPLVALTQS 442
Query: 519 GTERAKRKATSLLEHLRK 536
GT RAK+KA L+E LR+
Sbjct: 443 GTNRAKQKAEKLIELLRQ 496
Score = 28.9 bits (63), Expect(2) = 3e-30
Identities = 14/23 (60%), Positives = 17/23 (73%)
Frame = +1
Query: 373 DENKIIIGASGAISALVDLLQNG 395
+ENK IG SGAI LV+LL +G
Sbjct: 4 EENKAAIGRSGAIPLLVNLLGSG 72
>AL374758 weakly similar to PIR|T45588|T45 arm repeat containing protein
homolog - Arabidopsis thaliana, partial (15%)
Length = 538
Score = 114 bits (284), Expect(2) = 5e-29
Identities = 65/111 (58%), Positives = 81/111 (72%), Gaps = 1/111 (0%)
Frame = +1
Query: 262 ETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENA 321
E LV KL+ S + R + EIR L+K DNR +IAE GAIP LV+LL S+D QE+
Sbjct: 49 EFLVGKLATGSTDIQRQSAYEIRLLAKTGMDNRRIIAEVGAIPFLVTLLVSKDSRIQEHV 228
Query: 322 VTSILNLSIYENNKGLIMLAGAIPSIVQVLRAG-TMEARENAAATLFSLSL 371
VT++ NLSIY+NNK LIM AGAI +IV+VL G TMEARENAAA ++SLS+
Sbjct: 229 VTALFNLSIYDNNKILIMAAGAIDNIVEVLEFGKTMEARENAAAAIYSLSM 381
Score = 42.7 bits (99), Expect = 3e-04
Identities = 30/88 (34%), Positives = 46/88 (52%), Gaps = 1/88 (1%)
Frame = +1
Query: 347 IVQVLRAGTMEARENAAATLFSLSLAD-ENKIIIGASGAISALVDLLQNGSPRGKKDAAT 405
+V L G+ + + +A + L+ +N+ II GAI LV LL + R ++ T
Sbjct: 55 LVGKLATGSTDIQRQSAYEIRLLAKTGMDNRRIIAEVGAIPFLVTLLVSKDSRIQEHVVT 234
Query: 406 ALFNLCIYQGNKGRAIRAGIITALLNML 433
ALFNL IY NK + AG I ++ +L
Sbjct: 235 ALFNLSIYDNNKILIMAAGAIDNIVEVL 318
Score = 32.3 bits (72), Expect(2) = 5e-29
Identities = 22/44 (50%), Positives = 27/44 (61%), Gaps = 2/44 (4%)
Frame = +2
Query: 371 LADENKIIIGASG-AISALVDLLQNGSPRGKKDAATA-LFNLCI 412
L D K+ IGAS AI ALV LL+ G+ GK+D LFNL +
Sbjct: 383 LYDXVKVQIGASSRAIPALVGLLKEGTIIGKRDVCNMHLFNLAV 514
>CB894477 weakly similar to GP|10178181|db
gene_id:MQN23.13~pir||T07721~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (34%)
Length = 812
Score = 123 bits (308), Expect = 2e-28
Identities = 66/191 (34%), Positives = 118/191 (61%)
Frame = +2
Query: 262 ETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENA 321
ETL++KL + V E + +RS+++ + R+ + + + SL+ S V+ Q NA
Sbjct: 239 ETLLKKLKSKEVFEQEQGLVSLRSITRNREEARVSLCTPRILSSIRSLIDSRYVVVQVNA 418
Query: 322 VTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLADENKIIIGA 381
V S++NLS+ ++NK I+ +G +P ++ VL+ G E++E+AA LFSL+L D+NK+ IG
Sbjct: 419 VASLVNLSLEKSNKMRIVRSGFVPFLIDVLKGGFSESQEHAAGALFSLALDDDNKMAIGV 598
Query: 382 SGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMV 441
GA+ L+ L++ S R + D+A AL++L + Q N+ + ++ G++ LL+M+ + +M
Sbjct: 599 LGALQPLMHALRSESERTRHDSALALYHLTLVQSNRVKMVKLGVVPTLLSMV--MTGTMA 772
Query: 442 DEALTIMSVLA 452
L I+ LA
Sbjct: 773 SRVLLILCNLA 805
>BF631979 similar to PIR|A86416|A86 hypothetical protein AAG12624.1
[imported] - Arabidopsis thaliana, partial (20%)
Length = 559
Score = 87.8 bits (216), Expect(2) = 2e-25
Identities = 55/141 (39%), Positives = 79/141 (56%), Gaps = 8/141 (5%)
Frame = +3
Query: 207 QKLQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLTNGKIKKSDG-SFRDVTGDIAAIET-- 263
++ H L PN LR+L+ QWC H I L ++ ++ G +F AA+E
Sbjct: 78 KRFAHTRLVPNRALRNLIVQWCSAHGIP----LEPPEVMEAMGEAFASACPTKAALEANR 245
Query: 264 -----LVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQ 318
L+++L+ S A EIR L+K +NR +AEAGAIP L LL+S + + Q
Sbjct: 246 ATANLLIQQLANGSQSGKTVAAREIRLLAKTGRENRAFLAEAGAIPYLRDLLSSPNSVAQ 425
Query: 319 ENAVTSILNLSIYENNKGLIM 339
EN+VT++LNLSIY+ NK IM
Sbjct: 426 ENSVTALLNLSIYDKNKSRIM 488
Score = 46.6 bits (109), Expect(2) = 2e-25
Identities = 20/28 (71%), Positives = 23/28 (81%)
Frame = +2
Query: 182 TGQTYERSYIQRWIDCGNTTCPKTQQKL 209
TGQTY+RS I RW+D G+TTCPKT Q L
Sbjct: 2 TGQTYDRSSISRWMDEGHTTCPKTGQTL 85
Score = 40.8 bits (94), Expect = 0.001
Identities = 25/74 (33%), Positives = 40/74 (53%), Gaps = 1/74 (1%)
Frame = +3
Query: 347 IVQVLRAGTMEARENAAATLFSLSLAD-ENKIIIGASGAISALVDLLQNGSPRGKKDAAT 405
++Q L G+ + AA + L+ EN+ + +GAI L DLL + + ++++ T
Sbjct: 261 LIQQLANGSQSGKTVAAREIRLLAKTGRENRAFLAEAGAIPYLRDLLSSPNSVAQENSVT 440
Query: 406 ALFNLCIYQGNKGR 419
AL NL IY NK R
Sbjct: 441 ALLNLSIYDKNKSR 482
>TC91116 similar to GP|20161458|dbj|BAB90382. hypothetical protein~similar
to Arabidopsis thaliana chromosome 2 T20D16.23, partial
(18%)
Length = 753
Score = 113 bits (282), Expect = 2e-25
Identities = 62/137 (45%), Positives = 90/137 (65%)
Frame = +1
Query: 399 GKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAK 458
GKKDAATALFNL I+ NK R ++AG + L+ ++ D + MVD+A+ +++ LA+ E +
Sbjct: 16 GKKDAATALFNLSIFHENKNRIVQAGAVKHLVELM-DPAAGMVDKAVAVLANLATIPEGR 192
Query: 459 VSIVKASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELART 518
++I + IPVL++++ G R KENAAA LL LC LS + + GAV PL L+++
Sbjct: 193 IAIGQEGGIPVLVEVVELGSARGKENAAAALLHLCLHSNRFLSMVLQHGAVPPLVALSQS 372
Query: 519 GTERAKRKATSLLEHLR 535
GT RAK KA +LL R
Sbjct: 373 GTPRAKEKAQALLNQFR 423
Score = 73.6 bits (179), Expect = 2e-13
Identities = 51/174 (29%), Positives = 85/174 (48%), Gaps = 2/174 (1%)
Frame = +1
Query: 318 QENAVTSILNLSIYENNKGLIMLAGAIPSIVQVL--RAGTMEARENAAATLFSLSLADEN 375
+++A T++ NLSI+ NK I+ AGA+ +V+++ AG ++ A A L +L+ E
Sbjct: 19 KKDAATALFNLSIFHENKNRIVQAGAVKHLVELMDPAAGMVD---KAVAVLANLATIPEG 189
Query: 376 KIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTD 435
+I IG G I LV++++ GS RGK++AA AL +LC++
Sbjct: 190 RIAIGQEGGIPVLVEVVELGSARGKENAAAALLHLCLHSNR------------------- 312
Query: 436 SSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAAIL 489
+S++ H +P L+ L ++G PR KE A A+L
Sbjct: 313 -----------FLSMVLQH----------GAVPPLVALSQSGTPRAKEKAQALL 411
Score = 51.2 bits (121), Expect = 1e-06
Identities = 29/95 (30%), Positives = 55/95 (57%), Gaps = 1/95 (1%)
Frame = +1
Query: 273 VEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYE 332
V+++ A +A + ++ + RI I + G IPVLV ++ +ENA ++L+L ++
Sbjct: 139 VDKAVAVLANLATIP----EGRIAIGQEGGIPVLVEVVELGSARGKENAAAALLHLCLHS 306
Query: 333 NN-KGLIMLAGAIPSIVQVLRAGTMEARENAAATL 366
N +++ GA+P +V + ++GT A+E A A L
Sbjct: 307 NRFLSMVLQHGAVPPLVALSQSGTPRAKEKAQALL 411
>TC86190 weakly similar to PIR|T08454|T08454 hypothetical protein F22O6.170
- Arabidopsis thaliana, partial (60%)
Length = 1159
Score = 109 bits (273), Expect = 2e-24
Identities = 84/304 (27%), Positives = 149/304 (48%), Gaps = 22/304 (7%)
Frame = +1
Query: 157 DAIVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWI-DCGNTTCPKTQQKLQHLT-- 213
D + +P F+CPISLE+M+DPV V+TG TY+R I++W+ N TCP T+Q L T
Sbjct: 118 DEVEVPSFFVCPISLEIMKDPVTVSTGITYDRESIEKWLFSKKNMTCPVTKQLLSDYTDL 297
Query: 214 LTPNYVLRSLVSQWCIEHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSV 273
TPN+ LR L+ WC + G+ K + +T L+++ S S+
Sbjct: 298 TTPNHTLRRLIQAWC---TLNASQGIERIPTPKPPINKTQIT-------KLIKEASHSSL 447
Query: 274 EESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDV----------MTQENAVT 323
+ +RS++ S N+ I +AGA+ + S++ + +++ N V
Sbjct: 448 TVQLKCLKRLRSMASGSETNKRCIEDAGAVEFIASIVINNSCNIDFVNNSGSLSETNTVD 627
Query: 324 SILNL--SIYENNKGLIMLAG-----AIPSIVQVLRAGTMEARENAAATLFSLS-LADEN 375
L++ +++ + GL L I S+ +V++ G E+R A L S+S +A+
Sbjct: 628 EALSILHNLHVSEAGLKKLLAFRNGEFIESLTKVMQKGFFESRAYAVFLLKSMSEVAEPV 807
Query: 376 KIIIGASGAISALVDLLQNG-SPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLT 434
+++ LV +L++ S + K A L LC + N+ + + AG + L+ +L
Sbjct: 808 QLLHQKIELFVELVQVLKDQISQKVSKAALQTLIALCPWGRNRIKGVEAGTVPVLIELLL 987
Query: 435 DSSK 438
++ K
Sbjct: 988 ENCK 999
>TC79279 similar to GP|16226454|gb|AAL16172.1 AT3g01400/T13O15_4
{Arabidopsis thaliana}, partial (45%)
Length = 663
Score = 109 bits (273), Expect = 2e-24
Identities = 61/120 (50%), Positives = 79/120 (65%)
Frame = +1
Query: 261 IETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQEN 320
I LV L S+EE + A EIR L+K +NRI IA+AGAI L+SL+TS+D+ QE
Sbjct: 259 IRQLVSDLHSDSIEEQKQAAMEIRLLAKNKPENRIKIAKAGAIKPLISLVTSQDLQLQEY 438
Query: 321 AVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLADENKIIIG 380
VT+ILNLS+ + NK LI +GAI +V+ L +GT A+ENAA L LS +ENK IG
Sbjct: 439 GVTAILNLSLCDENKELIASSGAIKPLVRALNSGTSTAKENAACALLRLSQVEENKAAIG 618
Score = 63.5 bits (153), Expect = 2e-10
Identities = 46/134 (34%), Positives = 71/134 (52%), Gaps = 6/134 (4%)
Frame = +1
Query: 290 STDNRILIAEAGA-----IPVLVSLLTSEDVMTQENAVTSILNLSIYE-NNKGLIMLAGA 343
ST R+ +A A I LVS L S+ + Q+ A I L+ + N+ I AGA
Sbjct: 205 STSRRLFLACASENSDDLIRQLVSDLHSDSIEEQKQAAMEIRLLAKNKPENRIKIAKAGA 384
Query: 344 IPSIVQVLRAGTMEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDA 403
I ++ ++ + ++ +E + +LSL DENK +I +SGAI LV L +G+ K++A
Sbjct: 385 IKPLISLVTSQDLQLQEYGVTAILNLSLCDENKELIASSGAIKPLVRALNSGTSTAKENA 564
Query: 404 ATALFNLCIYQGNK 417
A AL L + NK
Sbjct: 565 ACALLRLSQVEENK 606
Score = 43.1 bits (100), Expect = 3e-04
Identities = 35/133 (26%), Positives = 65/133 (48%), Gaps = 2/133 (1%)
Frame = +1
Query: 344 IPSIVQVLRAGTMEARENAAATLFSLSL-ADENKIIIGASGAISALVDLLQNGSPRGKKD 402
I +V L + ++E ++ AA + L+ EN+I I +GAI L+ L+ + + ++
Sbjct: 259 IRQLVSDLHSDSIEEQKQAAMEIRLLAKNKPENRIKIAKAGAIKPLISLVTSQDLQLQEY 438
Query: 403 AATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIV 462
TA+ NL + NK +G I L+ L + + + A + L+ +E K +I
Sbjct: 439 GVTAILNLSLCDENKELIASSGAIKPLVRALNSGTSTAKENAACALLRLSQVEENKAAIG 618
Query: 463 K-ASTIPVLIDLL 474
K +L++LL
Sbjct: 619 KIRCNFRLLVNLL 657
>TC79378 similar to PIR|T00518|T00518 hypothetical protein At2g23140
[imported] - Arabidopsis thaliana, partial (36%)
Length = 2133
Score = 106 bits (264), Expect = 3e-23
Identities = 44/85 (51%), Positives = 61/85 (71%)
Frame = +1
Query: 150 LPEVKKPDAIVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKL 209
L + + + +P DF CP+SLELM DPVIVA+GQTYER++I+ WID G T CPKT Q L
Sbjct: 799 LKQSQSNSPVPVPPDFCCPLSLELMTDPVIVASGQTYERAFIKNWIDLGLTVCPKTHQTL 978
Query: 210 QHLTLTPNYVLRSLVSQWCIEHNIE 234
H L PNY +++L++ WC +N++
Sbjct: 979 AHTNLIPNYTVKALIANWCESNNVK 1053
Score = 90.5 bits (223), Expect = 1e-18
Identities = 55/117 (47%), Positives = 74/117 (63%), Gaps = 1/117 (0%)
Frame = +1
Query: 253 DVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTS 312
D++G A + +LV L ++ R A +EIR L+K + DNRI IA GAI +LV LL S
Sbjct: 1768 DLSGIEAQVRSLVEGLRSIDLDIQREATSEIRLLAKHNMDNRIAIANCGAINILVDLLQS 1947
Query: 313 EDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSI-VQVLRAGTMEARENAAATLFS 368
D QENAVT++LNLSI +NNK I ++ VL G+ E++EN+AATLFS
Sbjct: 1948 ADSRVQENAVTALLNLSINDNNKTAIAKCRVQLNL*YHVLEHGSPESQENSAATLFS 2118
Score = 51.2 bits (121), Expect = 1e-06
Identities = 34/100 (34%), Positives = 55/100 (55%), Gaps = 1/100 (1%)
Frame = +1
Query: 319 ENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLAD-ENKI 377
E V I++ S E L + + S+V+ LR+ ++ + A + + L+ + +N+I
Sbjct: 1717 ERLVPRIISSSAIEPRVDLSGIEAQVRSLVEGLRSIDLDIQREATSEIRLLAKHNMDNRI 1896
Query: 378 IIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNK 417
I GAI+ LVDLLQ+ R +++A TAL NL I NK
Sbjct: 1897 AIANCGAINILVDLLQSADSRVQENAVTALLNLSINDNNK 2016
Score = 47.0 bits (110), Expect = 2e-05
Identities = 37/129 (28%), Positives = 67/129 (51%), Gaps = 3/129 (2%)
Frame = +1
Query: 284 RSLSKRSTDNRILIAEAGA-IPVLVSLLTSEDVMTQENAVTSILNLSIYE-NNKGLIMLA 341
R +S + + R+ ++ A + LV L S D+ Q A + I L+ + +N+ I
Sbjct: 1732 RIISSSAIEPRVDLSGIEAQVRSLVEGLRSIDLDIQREATSEIRLLAKHNMDNRIAIANC 1911
Query: 342 GAIPSIVQVLRAGTMEARENAAATLFSLSLADENKIIIGASGA-ISALVDLLQNGSPRGK 400
GAI +V +L++ +ENA L +LS+ D NK I ++ +L++GSP +
Sbjct: 1912 GAINILVDLLQSADSRVQENAVTALLNLSINDNNKTAIAKCRVQLNL*YHVLEHGSPESQ 2091
Query: 401 KDAATALFN 409
+++A LF+
Sbjct: 2092 ENSAATLFS 2118
Score = 33.9 bits (76), Expect = 0.16
Identities = 29/113 (25%), Positives = 53/113 (46%), Gaps = 3/113 (2%)
Frame = +1
Query: 426 ITALLNMLTDSSKSMVDEALTIMSVLASHQ-EAKVSIVKASTIPVLIDLLRTGLPRNKEN 484
+ +L+ L + EA + + +LA H + +++I I +L+DLL++ R +EN
Sbjct: 1792 VRSLVEGLRSIDLDIQREATSEIRLLAKHNMDNRIAIANCGAINILVDLLQSADSRVQEN 1971
Query: 485 AAAILLALCKRDTDNLSCISRLGAVIPL--SELARTGTERAKRKATSLLEHLR 535
A LL L D +N + I++ + L L E + A +L +R
Sbjct: 1972 AVTALLNLSIND-NNKTAIAKCRVQLNL*YHVLEHGSPESQENSAATLFSFVR 2127
>BF644547 similar to GP|14149112|dbj bg55 {Bruguiera gymnorrhiza}, partial
(8%)
Length = 681
Score = 102 bits (254), Expect = 4e-22
Identities = 41/75 (54%), Positives = 58/75 (76%)
Frame = +1
Query: 162 PEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLR 221
PE++ CPISL LM DPV++A+G+TYER +IQ+W D GN CPKT++KL HL +TPN L+
Sbjct: 376 PEEYACPISLRLMYDPVVIASGETYERMWIQKWFDEGNVICPKTKKKLLHLAMTPNVALK 555
Query: 222 SLVSQWCIEHNIEQP 236
L+S+WC +++ P
Sbjct: 556 ELISKWCKTNDVSIP 600
>TC89898 similar to GP|19715649|gb|AAL91644.1 AT3g07360/F21O3_7 {Arabidopsis
thaliana}, partial (15%)
Length = 816
Score = 101 bits (252), Expect = 6e-22
Identities = 53/135 (39%), Positives = 76/135 (56%)
Frame = +1
Query: 145 SGSGNLPEVKKPDAIVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPK 204
S S N K + P++F CP+S E MRDPVIVA+GQTY+R +IQRW + GN TCP+
Sbjct: 349 SQSSNSMSFKINKNVTFPDEFKCPLSKEFMRDPVIVASGQTYDRPFIQRWFNAGNRTCPQ 528
Query: 205 TQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETL 264
T Q L H LTPN+++R ++ +W +E L N SD ++ D + L
Sbjct: 529 THQVLSHTLLTPNHLIREMIEKWSKNQGLE----LCNAMSYISDEGAKE--ADQSHFLCL 690
Query: 265 VRKLSCRSVEESRAA 279
+ K+S ++ AA
Sbjct: 691 LEKMSSTLCDQKEAA 735
>TC77754 similar to GP|19911585|dbj|BAB86896. syringolide-induced protein
13-1-1 {Glycine max}, partial (85%)
Length = 1577
Score = 101 bits (251), Expect = 8e-22
Identities = 64/229 (27%), Positives = 115/229 (49%), Gaps = 12/229 (5%)
Frame = +1
Query: 136 KTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWI 195
+T+ ++ G+G+ + I IP +F CP+SL+LM+DPV ++TG TY+R I +WI
Sbjct: 127 RTSKAKHQLPGAGD--GELTVEEITIPTNFRCPVSLDLMKDPVTLSTGITYDRFSIDKWI 300
Query: 196 DCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLTNGKIKKSDGSFRDVT 255
+ GN TCP T QKL +TPN+ +R ++ WC+E++ + +I S +V
Sbjct: 301 EAGNKTCPVTNQKLSTFEITPNHTIRKMIQSWCVENSSYGIERIPTPRIPVSGYEVNEVC 480
Query: 256 GDIAAIETLVRKLSCRSVEESRAA--VAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTS- 312
+ + CR+++E + V +I+ + S N+ +I G VL ++ S
Sbjct: 481 TRLLS--------GCRNLDEKKCVEFVGKIKIWWRESERNKRVIIGNGVSSVLATVFDSF 636
Query: 313 ------EDVMTQENA---VTSILNLSIYENNKGLIMLAGAIPSIVQVLR 352
E V+ E +T I+ S ++ + + + ++ +V V R
Sbjct: 637 SCVSFEEHVVVLEEVLEILTWIVKTSFGDSKTKMCLSSSSLNCLVWVFR 783
>TC93700
Length = 504
Score = 97.8 bits (242), Expect = 9e-21
Identities = 48/49 (97%), Positives = 49/49 (99%)
Frame = +2
Query: 1 DGAAKKIIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRR 49
DGAAKKIIFQFQRVTWKLEKLLSSLPYDDLDIS+EVKEQVDLVRNQLRR
Sbjct: 356 DGAAKKIIFQFQRVTWKLEKLLSSLPYDDLDISQEVKEQVDLVRNQLRR 502
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.131 0.359
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,544,849
Number of Sequences: 36976
Number of extensions: 163427
Number of successful extensions: 998
Number of sequences better than 10.0: 96
Number of HSP's better than 10.0 without gapping: 909
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 946
length of query: 540
length of database: 9,014,727
effective HSP length: 101
effective length of query: 439
effective length of database: 5,280,151
effective search space: 2317986289
effective search space used: 2317986289
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148469.7


