

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148405.4 - phase: 0
(609 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
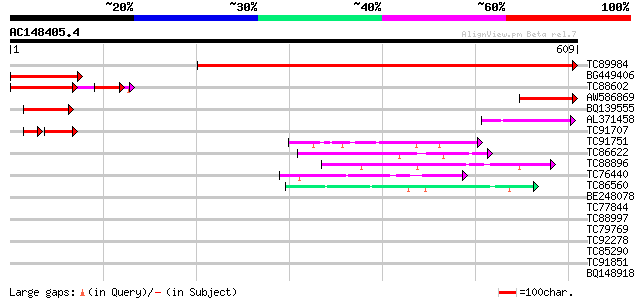
Sequences producing significant alignments: (bits) Value
TC89984 similar to GP|16604601|gb|AAL24093.1 unknown protein {Ar... 783 0.0
BG449406 similar to PIR|T04629|T04 hypothetical protein F10N7.30... 160 2e-39
TC88602 similar to PIR|T04629|T04629 hypothetical protein F10N7.... 158 5e-39
AW586869 125 4e-29
BQ139555 similar to GP|20198060|gb| unknown protein {Arabidopsis... 74 2e-13
AL371458 similar to GP|9294595|dbj| emb|CAA16573.1~gene_id:MVC8.... 68 1e-11
TC91707 weakly similar to GP|9294595|dbj|BAB02876.1 emb|CAA16573... 39 3e-07
TC91751 similar to GP|2739382|gb|AAC14505.1| unknown protein {Ar... 45 6e-05
TC86622 weakly similar to GP|20197623|gb|AAM15156.1 putative myo... 45 6e-05
TC88896 weakly similar to GP|6728968|gb|AAF26966.1| unknown prot... 45 8e-05
TC76440 weakly similar to SP|Q9LD55|IF3A_ARATH Eukaryotic transl... 43 3e-04
TC86560 similar to GP|8843737|dbj|BAA97285.1 myosin heavy chain-... 42 7e-04
BE248078 similar to GP|14334774|gb| unknown protein {Arabidopsis... 38 0.010
TC77844 similar to PIR|T49049|T49049 hypothetical protein T5P19.... 36 0.049
TC88997 weakly similar to GP|9719733|gb|AAF97835.1| EST gb|AI997... 35 0.084
TC79769 homologue to GP|7158882|gb|AAF37579.1| heat shock transc... 34 0.19
TC92278 similar to SP|O43093|KINH_SYNRA Kinesin heavy chain (Syn... 33 0.24
TC85290 similar to GP|7767653|gb|AAF69150.1| F27F5.2 {Arabidopsi... 33 0.32
TC91851 weakly similar to PIR|F96673|F96673 hypothetical protein... 33 0.32
BQ148918 weakly similar to PIR|T05232|T05 hypothetical protein F... 32 0.54
>TC89984 similar to GP|16604601|gb|AAL24093.1 unknown protein {Arabidopsis
thaliana}, partial (32%)
Length = 1303
Score = 783 bits (2023), Expect = 0.0
Identities = 408/408 (100%), Positives = 408/408 (100%)
Frame = +3
Query: 202 NSYDSSLLGNNTMELFSQPGHAKVLGHIRKLSNESVGSDGSSIRGSDISNFGIPNSSGDG 261
NSYDSSLLGNNTMELFSQPGHAKVLGHIRKLSNESVGSDGSSIRGSDISNFGIPNSSGDG
Sbjct: 3 NSYDSSLLGNNTMELFSQPGHAKVLGHIRKLSNESVGSDGSSIRGSDISNFGIPNSSGDG 182
Query: 262 SVDLPGSASVSRETDIVGHTKLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMED 321
SVDLPGSASVSRETDIVGHTKLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMED
Sbjct: 183 SVDLPGSASVSRETDIVGHTKLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMED 362
Query: 322 LIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRR 381
LIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRR
Sbjct: 363 LIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRR 542
Query: 382 KSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKA 441
KSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKA
Sbjct: 543 KSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKA 722
Query: 442 DIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGL 501
DIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGL
Sbjct: 723 DIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGL 902
Query: 502 LLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAAS 561
LLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAAS
Sbjct: 903 LLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAAS 1082
Query: 562 NVDKTNDIKDGVICGDEMRKIISDLFLDNVRLRKQTNRVTRHMLSNWI 609
NVDKTNDIKDGVICGDEMRKIISDLFLDNVRLRKQTNRVTRHMLSNWI
Sbjct: 1083NVDKTNDIKDGVICGDEMRKIISDLFLDNVRLRKQTNRVTRHMLSNWI 1226
>BG449406 similar to PIR|T04629|T04 hypothetical protein F10N7.30 -
Arabidopsis thaliana, partial (19%)
Length = 684
Score = 160 bits (404), Expect = 2e-39
Identities = 68/78 (87%), Positives = 74/78 (94%)
Frame = +2
Query: 1 MQRRSPPKHRHDGTSPLPLGMDWSPAPRKWNGRDTEWPHNHRTGWSYCVIIPSWVFVPKS 60
MQRRSPPKHRHDGTSPLPLGMDWSPAPRKWNGRDTEWPHNHRTGWSYCVIIPSWVFVPKS
Sbjct: 194 MQRRSPPKHRHDGTSPLPLGMDWSPAPRKWNGRDTEWPHNHRTGWSYCVIIPSWVFVPKS 373
Query: 61 KNSDPIVMSAFQDASQQN 78
KNSDPIV S++ ++++
Sbjct: 374 KNSDPIVASSWYTITRRD 427
>TC88602 similar to PIR|T04629|T04629 hypothetical protein F10N7.30 -
Arabidopsis thaliana, partial (25%)
Length = 1021
Score = 158 bits (400), Expect = 5e-39
Identities = 68/72 (94%), Positives = 68/72 (94%)
Frame = +1
Query: 1 MQRRSPPKHRHDGTSPLPLGMDWSPAPRKWNGRDTEWPHNHRTGWSYCVIIPSWVFVPKS 60
MQRRSPPKHRHDGTSPLPLGMDWSPAPRKWNGRDTEWPHNHRTGWSYCVIIPSWVFVPKS
Sbjct: 328 MQRRSPPKHRHDGTSPLPLGMDWSPAPRKWNGRDTEWPHNHRTGWSYCVIIPSWVFVPKS 507
Query: 61 KNSDPIVMSAFQ 72
KNSDPIV Q
Sbjct: 508 KNSDPIVFYRLQ 543
Score = 65.1 bits (157), Expect = 8e-11
Identities = 32/32 (100%), Positives = 32/32 (100%)
Frame = +2
Query: 92 VQSSLHSGLSLTAGSSSVASDYGSDTPYDASE 123
VQSSLHSGLSLTAGSSSVASDYGSDTPYDASE
Sbjct: 890 VQSSLHSGLSLTAGSSSVASDYGSDTPYDASE 985
Score = 49.3 bits (116), Expect = 4e-06
Identities = 34/69 (49%), Positives = 38/69 (54%), Gaps = 3/69 (4%)
Frame = +1
Query: 69 SAFQDASQQNSVKDPDSNNRDYSVQSSLHSGLSLTAGSSSVASDYGSDTPYDASEVGT-- 126
S+FQDASQQNSVKDPDSNNRDYS ++ S L Y +G
Sbjct: 820 SSFQDASQQNSVKDPDSNNRDYS--GAIISPFGLIINCWKFICCIRLW**YTI*CIGK*E 993
Query: 127 -PRIGQDDN 134
PRIGQDDN
Sbjct: 994 LPRIGQDDN 1020
>AW586869
Length = 422
Score = 125 bits (315), Expect = 4e-29
Identities = 61/62 (98%), Positives = 62/62 (99%)
Frame = -1
Query: 548 EVENLEKDAGSAASNVDKTNDIKDGVICGDEMRKIISDLFLDNVRLRKQTNRVTRHMLSN 607
+VENLEKDAGSAASNVDKTNDIKDGVICGDEMRKIISDLFLDNVRLRKQTNRVTRHMLSN
Sbjct: 341 QVENLEKDAGSAASNVDKTNDIKDGVICGDEMRKIISDLFLDNVRLRKQTNRVTRHMLSN 162
Query: 608 WI 609
WI
Sbjct: 161 WI 156
>BQ139555 similar to GP|20198060|gb| unknown protein {Arabidopsis thaliana},
partial (3%)
Length = 336
Score = 73.6 bits (179), Expect = 2e-13
Identities = 30/55 (54%), Positives = 38/55 (68%), Gaps = 1/55 (1%)
Frame = +2
Query: 15 SPLPLGMDWSPAPRKWNGRDTEWPHNHRTGWSYCVIIPSWVFVPKS-KNSDPIVM 68
SPLP GMDWSP P WNG T WPH + W++ IPSW+ VP+S +SDP+V+
Sbjct: 101 SPLPFGMDWSPPPPNWNGPTTLWPHTSSSSWTFSSTIPSWLLVPQSTPSSDPVVV 265
>AL371458 similar to GP|9294595|dbj| emb|CAA16573.1~gene_id:MVC8.4~strong
similarity to unknown protein {Arabidopsis thaliana},
partial (7%)
Length = 510
Score = 67.8 bits (164), Expect = 1e-11
Identities = 38/101 (37%), Positives = 59/101 (57%)
Frame = +1
Query: 507 QECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKT 566
QECSV E+ED+ + +S S DA + L TSD++I +LLAE + L +D + +
Sbjct: 1 QECSVNFLVEEEDKLNVDTSPS-DALDLLATSDNRIGLLLAEAQLLAQDVENVVDAAEGY 177
Query: 567 NDIKDGVICGDEMRKIISDLFLDNVRLRKQTNRVTRHMLSN 607
D +E+RK+++++F+DN LRKQ N V R L +
Sbjct: 178 RTTDDTGTTNEELRKMLANIFVDNASLRKQVNSVIRCALKS 300
>TC91707 weakly similar to GP|9294595|dbj|BAB02876.1
emb|CAA16573.1~gene_id:MVC8.4~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (15%)
Length = 684
Score = 39.3 bits (90), Expect(2) = 3e-07
Identities = 15/21 (71%), Positives = 15/21 (71%)
Frame = +2
Query: 15 SPLPLGMDWSPAPRKWNGRDT 35
SPLP GMDWSP P WNG T
Sbjct: 197 SPLPFGMDWSPPPPNWNGPTT 259
Score = 33.5 bits (75), Expect(2) = 3e-07
Identities = 14/36 (38%), Positives = 22/36 (60%), Gaps = 1/36 (2%)
Frame = +3
Query: 38 PHNHRTGWSYCVIIPSWVFVPKS-KNSDPIVMSAFQ 72
P+ + W++ IPSW+ VP+S +SDP+V Q
Sbjct: 270 PNTSSSSWTFSSTIPSWLLVPQSTPSSDPVVFYKIQ 377
>TC91751 similar to GP|2739382|gb|AAC14505.1| unknown protein {Arabidopsis
thaliana}, partial (38%)
Length = 1200
Score = 45.4 bits (106), Expect = 6e-05
Identities = 64/225 (28%), Positives = 105/225 (46%), Gaps = 16/225 (7%)
Frame = +2
Query: 300 KLNRILSTMQRRLVTAKTDMEDLIV---RLNQEIAAKDFLTTKVKDLEVELETSKQKNKE 356
K +RI + +R LV + D D + R E A + TK + L+ EL+++K+ +E
Sbjct: 242 KAHRIQTVERRNLVEQELDKADEEIPEYRKQAETAEQ----TKNQVLK-ELDSTKRLIEE 406
Query: 357 ---NLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDV 413
NL++A E+ Q + D E + + EME + ES +V E +
Sbjct: 407 LKLNLERAQTEEQ----QARQDSELAKLRVEEMEQGIADES--SVAAKAQLEVAKARYTA 568
Query: 414 LLQDLDAAREQLEILSKQYGEL-----EAKSKADIKVLV-KEV-KSLRSSQTEL---KKE 463
+ DL A +E+L+ L K+Y L EA KA+ V KEV KS+ EL K+
Sbjct: 569 AITDLAAVKEELDALRKEYASLVTDRDEAIKKAEEAVTASKEVEKSVEDLTIELIATKES 748
Query: 464 LSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQE 508
L + EAE+ + +Q + W K+L++ L+++ E
Sbjct: 749 LETAHAAHLEAEEQRIGTVMARDQDSLNWEKELKQAEEELQRINE 883
>TC86622 weakly similar to GP|20197623|gb|AAM15156.1 putative myosin heavy
chain {Arabidopsis thaliana}, partial (17%)
Length = 2546
Score = 45.4 bits (106), Expect = 6e-05
Identities = 49/218 (22%), Positives = 98/218 (44%), Gaps = 9/218 (4%)
Frame = +3
Query: 310 RRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERERF 369
+ ++ +K EDL+ + N+E+ + V D L + Q K+ L+ IL E+
Sbjct: 1269 KEILASKNAAEDLVTQHNEEVQTLKSQISSVIDDRNLLNETNQNLKKELESIILDLEEKL 1448
Query: 370 TQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQD---LDAAREQLE 426
+ Q + + L+ + +++++ +S + + + L ++ + AA Q E
Sbjct: 1449 KEHQKNEDSLKSEVETLKIEIAEKSALQSRLHEIEAQLAKAESRLHEEVGSVQAAASQRE 1628
Query: 427 I-LSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKEL-----SESIKEKYEAEKLLLH 480
+ LS ++ + E K + E+ L EL+KEL + + ++ E++KL L
Sbjct: 1629 VDLSSKFEDYEQK--------ISEINVLNGKVAELEKELHLAQDTIANQKGEESQKLELE 1784
Query: 481 EREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDE 518
K+ E+ +K + LL KQ+ E +L DE
Sbjct: 1785 AALKNSVEELETKK--NEISLLQKQVIEFEQKLQQADE 1892
>TC88896 weakly similar to GP|6728968|gb|AAF26966.1| unknown protein
{Arabidopsis thaliana}, partial (27%)
Length = 1097
Score = 45.1 bits (105), Expect = 8e-05
Identities = 59/271 (21%), Positives = 123/271 (44%), Gaps = 20/271 (7%)
Frame = +2
Query: 336 LTTKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDM---------EELRRKSLEM 386
L T+++ L+ ELE +K +++ ++ LIE+ M +E R+K E+
Sbjct: 8 LQTEIEALKHELEKAKGYDEKLAEKETLIEQLNVESEAAKMAESYARSVLDECRKKVEEL 187
Query: 387 EMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELE---AKSKADI 443
EMK++ + L+ E+ +Q + + L A ++ L ++ G LE + + D+
Sbjct: 188 EMKVEEANQLERSASLSLETATKQLEGKNELLHDAESEISSLKEKLGMLEMTVGRQRGDL 367
Query: 444 KVLVKEVKSLRSSQTELKKELS--ESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGL 501
+ + + + + E+ K++ ES E EK EK + + + LE+
Sbjct: 368 EDAERCLLAAKEENIEMSKKIESLESEIETVSKEKAQALNNEKLSASSV--QTLLEEKNK 541
Query: 502 LLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKT-SDDQIDILL---AEVENLEKDAG 557
L+ +L+ C ++E++T L S A +++ + D + LL AE E+ E
Sbjct: 542 LINELEICR-----DEEEKTKLAMDSLASALHEVSAEARDTKEKLLANQAEHESYETQIE 706
Query: 558 SAASNVDKTNDIKDGVI--CGDEMRKIISDL 586
S+++ + + + ++ E+ + SDL
Sbjct: 707 DLKSDLEASKEKYESMLNDAHHEIEDLKSDL 799
>TC76440 weakly similar to SP|Q9LD55|IF3A_ARATH Eukaryotic translation
initiation factor 3 subunit 10 (eIF-3 theta), partial
(41%)
Length = 1953
Score = 43.1 bits (100), Expect = 3e-04
Identities = 48/214 (22%), Positives = 104/214 (48%), Gaps = 12/214 (5%)
Frame = +1
Query: 290 QLVLPLDQR-NKLNRILSTMQ-------RRLVTAKTDMEDLIVRLNQEIAAKDFL--TTK 339
Q++ P D++ +KL +L T+ +RL+ K+ +E +++ K+ + +
Sbjct: 319 QMICPPDRKQSKLGALLPTLSEVVAKEHKRLLARKSIIEKRKEEQERQLLEKEREEESKR 498
Query: 340 VKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNV- 398
++ L+++ E +++ ++Q I+R + + + D EE LE E +LK + V
Sbjct: 499 LRLLKIDEEAEQRRLATEIEQR-KIQRLQREKEERDREEAEALRLEAEKRLKRKGKKPVI 675
Query: 399 -GQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQ 457
G +++ES++Q L ++ RE+ E+ K ++ L K + L ++
Sbjct: 676 EGGQISRESLMQ-----LTLVEQVRERQEMEKK------------LQKLAKTMDHLERAK 804
Query: 458 TELKKELSESIKEKYEAEKLLLHEREKSEQAEIA 491
E L ++ ++ E+ +LHERE+ Q E++
Sbjct: 805 REEAAPLIDAAYQQRLVEERVLHEREQQLQVELS 906
>TC86560 similar to GP|8843737|dbj|BAA97285.1 myosin heavy chain-like
{Arabidopsis thaliana}, partial (28%)
Length = 2450
Score = 42.0 bits (97), Expect = 7e-04
Identities = 65/316 (20%), Positives = 123/316 (38%), Gaps = 44/316 (13%)
Frame = +2
Query: 297 QRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKN-K 355
++ KL + S Q+ LV + D + E + K L K K+ E+ S +
Sbjct: 956 EQTKLAYVESQQQQALVLTEKDALRQSYKATLEQSKKKLLALK-KEFNPEITKSLEAQLA 1132
Query: 356 ENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLL 415
E + + + E + D++ ++ + E++ K V ++ T S+V+ V L
Sbjct: 1133 ETMNEIAALHTELENKRSSDLDSVKTVTSELD-GAKESLQKVVDEENTLRSLVETLKVEL 1309
Query: 416 QDLDAAREQLEI---------------LSKQYGELEAKSKADIKV--------------- 445
+++ +L+ L K ELEA S + KV
Sbjct: 1310 ENVKKEHSELKEKESELESTVGNLHVKLRKSKSELEACSADESKVRGASEEMILTLSRLT 1489
Query: 446 -----LVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCG 500
+EV+ +++ ELKKE + EAEK L E++E A+ A +E+
Sbjct: 1490 SETEEARREVEDMKNKTDELKKEAEATKLALEEAEKKLKEATEEAEAAKAAEASAIEQIT 1669
Query: 501 LLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKL--------KTSDDQIDILLAEVENL 552
+L ++ E + + ST+ F L K +D ++D A+VE +
Sbjct: 1670 VLTERTSARRASTSSE----SGAAMTISTEEFESLKRKVEESDKLADMKVDAAKAQVEAV 1837
Query: 553 EKDAGSAASNVDKTND 568
+ ++ T++
Sbjct: 1838 KASENEVLKKLEATSE 1885
>BE248078 similar to GP|14334774|gb| unknown protein {Arabidopsis thaliana},
partial (24%)
Length = 392
Score = 38.1 bits (87), Expect = 0.010
Identities = 32/112 (28%), Positives = 56/112 (49%), Gaps = 2/112 (1%)
Frame = +3
Query: 389 KLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVK 448
KLKSE + + ES V + + + DL++ E ++ ++++ + K++ +KV +
Sbjct: 45 KLKSELEAQNSEKVNWESRVAKLEKKIHDLNSKLEDVQKINEEQKKQIRKTERALKVAEE 224
Query: 449 EVKSLRSSQTELKKELSESIKEKY--EAEKLLLHEREKSEQAEIAWRKQLEK 498
E+ + T KELSES+ E + E K LL E+ +K LEK
Sbjct: 225 EMLKAKLEATTKAKELSESVAESHWNEHGKPLL---------EVISQKALEK 353
>TC77844 similar to PIR|T49049|T49049 hypothetical protein T5P19.130 -
Arabidopsis thaliana, partial (70%)
Length = 1969
Score = 35.8 bits (81), Expect = 0.049
Identities = 39/153 (25%), Positives = 69/153 (44%), Gaps = 5/153 (3%)
Frame = +2
Query: 342 DLEVELETSKQKNKENLQQAILIERERFTQMQ-----WDMEELRRKSLEMEMKLKSESGG 396
D +V ET++ +L++ I E ++ D+ K+L KL +E+
Sbjct: 395 DHQVFQETAEDNQGPSLKEVIEQEASNLSEQNNRISVRDLASKFDKNLSAAAKLSNEAK- 571
Query: 397 NVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSS 456
+E + VLL+ L A E L+ G ++K D+ + V++L
Sbjct: 572 ------LREVPSLEGHVLLKKLRDALEYLK------GRFTGRNKEDVANAISMVEALAVK 715
Query: 457 QTELKKELSESIKEKYEAEKLLLHEREKSEQAE 489
T+ + EL I+EK+E +KLL ++ SE A+
Sbjct: 716 LTQNEGEL---IQEKFEVKKLLNFLKQASEDAK 805
>TC88997 weakly similar to GP|9719733|gb|AAF97835.1| EST gb|AI997943 comes
from this gene. {Arabidopsis thaliana}, partial (29%)
Length = 1323
Score = 35.0 bits (79), Expect = 0.084
Identities = 46/210 (21%), Positives = 98/210 (45%), Gaps = 15/210 (7%)
Frame = +2
Query: 370 TQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTK------ESIVQQNDVLLQDLDAARE 423
T +QW++E ++ E+ K+ S+ +V ++L + ++ +N + + ++ RE
Sbjct: 8 TAVQWNLEVEVKQVAELRQKMASKE--SVHEELRRSLRNPNQTGASRNQLASKGVEFERE 181
Query: 424 QLE----ILSKQYGELEAKSK---ADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEK 476
LE ++ + +L+ K++ ADI++ KE++ + ELK+ L + + +
Sbjct: 182 ILEAEHSFINDKVAQLQEKARKLEADIEMTRKEIEEPTEVEVELKRRLHQMTDHLIQKQA 361
Query: 477 LLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKL- 535
+ E SE+A + +R +E LL + S +SSS+D + L
Sbjct: 362 KV--ESLSSEKASLIFR--IEAVSRLLDENMSVS-------GSTAMNPASSSSDLESGLW 508
Query: 536 KTSDDQI-DILLAEVENLEKDAGSAASNVD 564
+ S+ + +L A + + +K GS +D
Sbjct: 509 ELSNSKFKPMLKARIHSGKKQLGSLLQQID 598
>TC79769 homologue to GP|7158882|gb|AAF37579.1| heat shock transcription
factor {Medicago sativa}, complete
Length = 1979
Score = 33.9 bits (76), Expect = 0.19
Identities = 36/176 (20%), Positives = 72/176 (40%)
Frame = +1
Query: 404 KESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKE 463
++S+V + + L QD REQL + + +Y + + +++ L Q ++
Sbjct: 1009 RQSMVDEIEKLKQD----REQLLMETNRYQHDWETYEIQMHCSKDQLEKLEHKQQKMLSS 1176
Query: 464 LSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFL 523
+SE++++ A LL + + R ++ E SV LP E+ +
Sbjct: 1177 VSEALQKPMIAVNLLPLAEAMERKRRLPARSGCFNNEASVEDAMETSVALPRENAE---- 1344
Query: 524 QSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKTNDIKDGVICGDEM 579
+ST N + DQ++ +A E L + G + D+ + C D +
Sbjct: 1345 --DNSTLTLNTERL--DQLEASVAFWETLAHEVGGNFVHTHSNMDLDESTCCADSL 1500
>TC92278 similar to SP|O43093|KINH_SYNRA Kinesin heavy chain (Synkin).
{Syncephalastrum racemosum}, partial (10%)
Length = 709
Score = 33.5 bits (75), Expect = 0.24
Identities = 42/197 (21%), Positives = 87/197 (43%), Gaps = 1/197 (0%)
Frame = +3
Query: 346 ELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKE 405
EL T +++ E+ + + + ++ ++ E L RK E+E++L + +L E
Sbjct: 138 ELSTLRRELSES-KSLVTQHEQTINELHYENEHLTRKRDELEIRLTT-------LELEYE 293
Query: 406 SIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELS 465
++ + + ++ + + E +++ G+LEA+ A +V KE+ ELK+EL
Sbjct: 294 ELLDKT-IAEEEANNNIDITETIAELRGKLEAQFVAKKEVQQKEI-------DELKQELE 449
Query: 466 ESIKEKYEAEKLLLHEREKSEQAEIAWR-KQLEKCGLLLKQLQECSVELPYEDEDRTFLQ 524
+ +E ++ L ++ +E + A K+++ G Q E E +T Q
Sbjct: 450 KKNEELHKLNSALADLKKVNEDLQTAINDKEIKSDG----QKNVADKEKDMERIXKTMAQ 617
Query: 525 SSSSTDAFNKLKTSDDQ 541
+ D K D Q
Sbjct: 618 QLADFDVMKKALMRDLQ 668
>TC85290 similar to GP|7767653|gb|AAF69150.1| F27F5.2 {Arabidopsis
thaliana}, partial (32%)
Length = 1558
Score = 33.1 bits (74), Expect = 0.32
Identities = 29/106 (27%), Positives = 56/106 (52%), Gaps = 3/106 (2%)
Frame = +1
Query: 296 DQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNK 355
++R K + +R+ K + ++ I ++NQ + K LT V +E K++ K
Sbjct: 784 EKRKKNVKRRRKKRRKENVRKKNQKNDIRKMNQIVIIKT*LTAMV----IERRRKKKRIK 951
Query: 356 -ENLQQAILIERERFTQMQWDMEELRRKSLE--MEMKLKSESGGNV 398
EN+++ I R M W ++++RRKSLE +M++ + S G++
Sbjct: 952 RENIEEGI-----RAVWMMWILKKMRRKSLENREDMEVTARSLGSM 1074
Score = 28.9 bits (63), Expect = 6.0
Identities = 39/176 (22%), Positives = 76/176 (43%), Gaps = 1/176 (0%)
Frame = +3
Query: 313 VTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVEL-ETSKQKNKENLQQAILIERERFTQ 371
V + ED +++E K +K L EL E +K+K ++ ++ + + FT
Sbjct: 402 VATTSVFEDFKSAVSEEATCKTISEINLKLLFEELLERAKEKEEKEAKKRQRLADD-FTN 578
Query: 372 MQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQ 431
+ + ++++ S E K E T+E I N+ +++ E + L ++
Sbjct: 579 LLYTLKDIITSSTWEECKALFED--------TQEYISIGNESYSKEI--FEEYITYLKEK 728
Query: 432 YGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQ 487
E E K + + ++ E ++E E KEK + EK E+EKS++
Sbjct: 729 AKEKERKREEE------------KAKKEKEREEKEKRKEKEKKEKEREREKEKSKE 860
>TC91851 weakly similar to PIR|F96673|F96673 hypothetical protein F13O11.30
[imported] - Arabidopsis thaliana, partial (12%)
Length = 981
Score = 33.1 bits (74), Expect = 0.32
Identities = 36/145 (24%), Positives = 63/145 (42%), Gaps = 1/145 (0%)
Frame = +3
Query: 410 QNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIK 469
Q +V +DL A+EQ+ K+ + + K +V + + L+ + K+ ES
Sbjct: 375 QLNVAQEDLKKAKEQILQAEKEKVKAIDELKEAQRVAEEANEKLQEALVAQKQAKEESEI 554
Query: 470 EKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSST 529
EK+ A +L E + E W+K+LE L S+ E+ +R + +
Sbjct: 555 EKFRAVELEQAGIETVNKKEEEWQKELESV-RNQHALDVASLASTTEELERVKQELTMMC 731
Query: 530 DAFNK-LKTSDDQIDILLAEVENLE 553
DA N+ L +DD + E +E
Sbjct: 732 DAKNQALNHADDAAKVAEVHAEKVE 806
>BQ148918 weakly similar to PIR|T05232|T05 hypothetical protein F18A5.20 -
Arabidopsis thaliana, partial (17%)
Length = 639
Score = 32.3 bits (72), Expect = 0.54
Identities = 36/140 (25%), Positives = 64/140 (45%), Gaps = 8/140 (5%)
Frame = +1
Query: 263 VDLPGSASVSRETDIVGHTKLKSSGDAQLVL---PLDQRNK----LNRILSTMQRRLVTA 315
VD PG+ S T + ++ + D + + PL+ N L L + TA
Sbjct: 82 VDSPGTGIYSSTTQ--ENIEIAGNEDNHVTMYENPLEGENAAYAALYLELEKERAAAATA 255
Query: 316 KTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERERFTQ-MQW 374
+ +I RL +E ++ + + + L +E S + + N+ Q ILI RER ++
Sbjct: 256 ADEAMAMISRLQEEKSSMEMEMRQYERL-IEERVSYDEEEMNIMQEILIRRERENLFLEK 432
Query: 375 DMEELRRKSLEMEMKLKSES 394
++E R+ SL +L S+S
Sbjct: 433 ELETFRQMSLTESCELHSKS 492
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.310 0.129 0.356
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,472,169
Number of Sequences: 36976
Number of extensions: 207286
Number of successful extensions: 939
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 930
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 936
length of query: 609
length of database: 9,014,727
effective HSP length: 102
effective length of query: 507
effective length of database: 5,243,175
effective search space: 2658289725
effective search space used: 2658289725
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148405.4


