

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148405.2 + phase: 0
(209 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
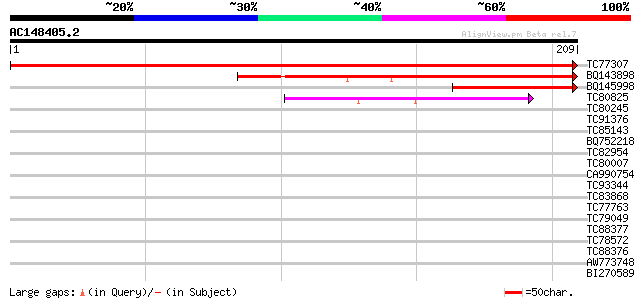
Sequences producing significant alignments: (bits) Value
TC77307 similar to PIR|T05402|T05402 H+-transporting ATP synthas... 391 e-110
BQ143898 similar to GP|14596171|gb| H+-transporting ATP synthase... 92 1e-19
BQ145998 similar to GP|14596171|gb| H+-transporting ATP synthase... 72 1e-13
TC80825 similar to PIR|F84614|F84614 probable kinesin heavy chai... 40 8e-04
TC80245 similar to PIR|T48581|T48581 hypothetical protein T31B5.... 38 0.003
TC91376 weakly similar to GP|17065054|gb|AAL32681.1 putative pro... 35 0.016
TC85143 similar to PIR|T11852|T11852 lipoxygenase (EC 1.13.11.12... 34 0.035
BQ752218 similar to GP|19170914|emb hypothetical protein {Enceph... 32 0.13
TC82954 weakly similar to GP|8809591|dbj|BAA97142.1 emb|CAB82814... 32 0.17
TC80007 31 0.30
CA990754 similar to GP|21397269|g Unknown protein {Oryza sativa ... 31 0.39
TC93344 weakly similar to GP|19310663|gb|AAL85062.1 unknown prot... 31 0.39
TC83868 30 0.66
TC77763 similar to GP|15215674|gb|AAK91382.1 AT4g27500/F27G19_10... 30 0.66
TC79049 weakly similar to PIR|T10798|T10798 pherophorin-S - Volv... 29 1.5
TC88377 similar to GP|8778315|gb|AAF79324.1| F14J16.29 {Arabidop... 29 1.5
TC78572 similar to GP|9801280|emb|CAC03581.1 putative 1-deoxy-D-... 29 1.5
TC88376 similar to GP|8778315|gb|AAF79324.1| F14J16.29 {Arabidop... 28 2.5
AW773748 similar to GP|10176892|dbj Na+-dependent inorganic phos... 28 3.3
BI270589 similar to PIR|C96829|C96 unknown protein F19K16.21 [im... 28 3.3
>TC77307 similar to PIR|T05402|T05402 H+-transporting ATP synthase (EC
3.6.1.34) chain 9 - Arabidopsis thaliana, partial (72%)
Length = 940
Score = 391 bits (1005), Expect = e-110
Identities = 209/209 (100%), Positives = 209/209 (100%)
Frame = +1
Query: 1 MASMIMASSKPIIRVPTNSLPSTPKLPSLQTITPNLKLQLSKLKSMTLAATSLSFVFAPP 60
MASMIMASSKPIIRVPTNSLPSTPKLPSLQTITPNLKLQLSKLKSMTLAATSLSFVFAPP
Sbjct: 178 MASMIMASSKPIIRVPTNSLPSTPKLPSLQTITPNLKLQLSKLKSMTLAATSLSFVFAPP 357
Query: 61 SLAFEKAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTS 120
SLAFEKAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTS
Sbjct: 358 SLAFEKAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTS 537
Query: 121 EEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEK 180
EEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEK
Sbjct: 538 EEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEK 717
Query: 181 QKEETVKALDSQIAALSQDIVNKVLPVSR 209
QKEETVKALDSQIAALSQDIVNKVLPVSR
Sbjct: 718 QKEETVKALDSQIAALSQDIVNKVLPVSR 804
>BQ143898 similar to GP|14596171|gb| H+-transporting ATP synthase-like
protein {Arabidopsis thaliana}, partial (11%)
Length = 721
Score = 92.0 bits (227), Expect = 1e-19
Identities = 57/128 (44%), Positives = 79/128 (61%), Gaps = 3/128 (2%)
Frame = +1
Query: 85 FLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEV--KELEEQANAVLRAARA-EI 141
FLMFAL P + + + NS + + + +EQA A+ + ++
Sbjct: 13 FLMFALRYTLLNPFRN-LHGPPRRLISEANSAASPAHPKN*RSWKEQAYALCESMTVLKL 189
Query: 142 AVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETVKALDSQIAALSQDIV 201
VALN MK++T A+VEE +AEGR V+AEL++ALAN+ + + ALDSQIAALSQDIV
Sbjct: 190 QVALNPMKRDTHADVEETMAEGRNTVEAELRQALANMHNRTDNADNALDSQIAALSQDIV 369
Query: 202 NKVLPVSR 209
N+VLPVSR
Sbjct: 370 NRVLPVSR 393
>BQ145998 similar to GP|14596171|gb| H+-transporting ATP synthase-like
protein {Arabidopsis thaliana}, partial (15%)
Length = 279
Score = 72.4 bits (176), Expect = 1e-13
Identities = 38/46 (82%), Positives = 41/46 (88%)
Frame = +3
Query: 164 RKKVDAELQEALANLEKQKEETVKALDSQIAALSQDIVNKVLPVSR 209
R + + +EALANLEKQKEETVKALDSQIAALSQDIVNKVLPVSR
Sbjct: 6 RPRAEFGTREALANLEKQKEETVKALDSQIAALSQDIVNKVLPVSR 143
>TC80825 similar to PIR|F84614|F84614 probable kinesin heavy chain
[imported] - Arabidopsis thaliana, partial (16%)
Length = 1771
Score = 39.7 bits (91), Expect = 8e-04
Identities = 24/97 (24%), Positives = 51/97 (51%), Gaps = 5/97 (5%)
Frame = +2
Query: 102 MDNRDAEIRGKLNSVTNTSEEVKELE---EQANAVLRAARAEIAVALNQM--KKETQAEV 156
+ N +++ +GK N N E++KELE E ++ + +++ ++ K+ET +
Sbjct: 755 LQNIESKAKGKDNIHKNLQEKIKELEGQIELKTSMQNQSEKQVSQLCEKLKGKEETCCTL 934
Query: 157 EEKIAEGRKKVDAELQEALANLEKQKEETVKALDSQI 193
+ K+ E +K+ +LQ AN +++ + K L Q+
Sbjct: 935 QHKVKELERKIKEQLQTETANFQQKAWDLEKKLKDQL 1045
>TC80245 similar to PIR|T48581|T48581 hypothetical protein T31B5.160 -
Arabidopsis thaliana, partial (60%)
Length = 895
Score = 37.7 bits (86), Expect = 0.003
Identities = 27/76 (35%), Positives = 43/76 (56%), Gaps = 3/76 (3%)
Frame = +2
Query: 114 NSVTNTSEEVKELEEQ-ANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKV--DAE 170
+S EE+K LEE+ A + A R + LN +E + E+E ++AEG KK+ D E
Sbjct: 275 SSKRQQDEELKLLEEETARRIEEAIRKNVEEKLNS--EEVKVEIERRVAEGVKKLFDDVE 448
Query: 171 LQEALANLEKQKEETV 186
+Q LEK+K++ +
Sbjct: 449 VQ-----LEKEKQDAL 481
>TC91376 weakly similar to GP|17065054|gb|AAL32681.1 putative protein
{Arabidopsis thaliana}, partial (41%)
Length = 997
Score = 35.4 bits (80), Expect = 0.016
Identities = 18/65 (27%), Positives = 37/65 (56%)
Frame = +1
Query: 134 LRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETVKALDSQI 193
L+ ++EI + + Q + + E+EE+ + + + A +QEA+ KQKEE ++ ++ Q
Sbjct: 442 LQNQQSEIDLFIAQHTERVRMEIEEQRLKQSRMLQAAIQEAVTKKLKQKEEEIQRMEKQN 621
Query: 194 AALSQ 198
L +
Sbjct: 622 LMLQE 636
>TC85143 similar to PIR|T11852|T11852 lipoxygenase (EC 1.13.11.12) - kidney
bean, partial (93%)
Length = 3009
Score = 34.3 bits (77), Expect = 0.035
Identities = 24/74 (32%), Positives = 35/74 (46%), Gaps = 5/74 (6%)
Frame = -2
Query: 16 PTNSLPSTPKLPSLQTITPNLKLQLSKLKSMTLAATSLSFVFAPPS-----LAFEKAALF 70
PTN P L +L P L+ +++ SMT TSL+ PS ++F+ A
Sbjct: 278 PTNPFPILSVLVALINWRPTLRPKIAVAPSMTPPTTSLTRPNPAPSEGPALISFKSNAFL 99
Query: 71 DFNLTLPIIVVEFL 84
N T+P+I FL
Sbjct: 98 HINTTVPLIFCFFL 57
>BQ752218 similar to GP|19170914|emb hypothetical protein {Encephalitozoon
cuniculi}, partial (5%)
Length = 637
Score = 32.3 bits (72), Expect = 0.13
Identities = 21/76 (27%), Positives = 37/76 (48%)
Frame = -2
Query: 124 KELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKE 183
K+ E+A + AR + KKE + + +E+ + KK+ E +EA +K K+
Sbjct: 576 KKAREEAAKKAKKAREAAYKKAKEEKKEAEKKAKEEKRQAEKKIKEEKKEAEKKAKKDKK 397
Query: 184 ETVKALDSQIAALSQD 199
E + D++ A QD
Sbjct: 396 EQDRK-DAESALSRQD 352
>TC82954 weakly similar to GP|8809591|dbj|BAA97142.1
emb|CAB82814.1~gene_id:MNB8.8~similar to unknown protein
{Arabidopsis thaliana}, partial (7%)
Length = 783
Score = 32.0 bits (71), Expect = 0.17
Identities = 18/88 (20%), Positives = 44/88 (49%)
Frame = +3
Query: 102 MDNRDAEIRGKLNSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIA 161
++ +D +I + E+ K+ E+ A + +++ E + ++K A +E +
Sbjct: 525 VEEKDKKIEEEEKKRKELEEKAKKAEKDAEELRESSKREGQEHSSDLRKHKTAFIE--LV 698
Query: 162 EGRKKVDAELQEALANLEKQKEETVKAL 189
++ ++AEL A+ +L+ KEE + +
Sbjct: 699 SNQRHLEAELGRAVKHLDAAKEELIAVM 782
>TC80007
Length = 1599
Score = 31.2 bits (69), Expect = 0.30
Identities = 27/85 (31%), Positives = 42/85 (48%), Gaps = 3/85 (3%)
Frame = +1
Query: 114 NSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDA---E 170
N + + KELEEQ NA+ +A I+V Q K T + + ++ E K + A E
Sbjct: 1063 NQLATLEQRKKELEEQINAI----KANISVF--QSAKITATKRKREVFEEAKTLKAQRDE 1224
Query: 171 LQEALANLEKQKEETVKALDSQIAA 195
L+E + +L K + E K + I A
Sbjct: 1225 LREQVPHL-KDEREVAKKIQENIQA 1296
>CA990754 similar to GP|21397269|g Unknown protein {Oryza sativa (japonica
cultivar-group)}, partial (15%)
Length = 700
Score = 30.8 bits (68), Expect = 0.39
Identities = 30/118 (25%), Positives = 57/118 (47%), Gaps = 7/118 (5%)
Frame = +3
Query: 79 IVVEFLFLMFALDK-LYFTPLGTFMDNRDAEIRGKLNSVTNTSEE--VKELEEQANAVLR 135
IV +F +D+ L PL T +DN A +R +L + + E + + ++QA+ V+
Sbjct: 168 IVYQFRKAQNEIDRYLRLVPLITLVDN--ARVRERLEVIEHDQREYTLDDEDQQAHTVIL 341
Query: 136 AA---RAEIAVALNQMK-KETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETVKAL 189
+ + AV + E I + ++K++ ELQ + ANL+ + E ++ L
Sbjct: 342 KPDPDKEDTAVLKKTLSCSYPNCTFPEAIKKEKEKLNLELQRSQANLDMNQCEVIQRL 515
>TC93344 weakly similar to GP|19310663|gb|AAL85062.1 unknown protein
{Arabidopsis thaliana}, partial (18%)
Length = 420
Score = 30.8 bits (68), Expect = 0.39
Identities = 25/63 (39%), Positives = 30/63 (46%)
Frame = +1
Query: 1 MASMIMASSKPIIRVPTNSLPSTPKLPSLQTITPNLKLQLSKLKSMTLAATSLSFVFAPP 60
MA I A S + P SLPS LP TIT K QL ++ T SL +F PP
Sbjct: 43 MAMEITAMSS--LPTPKPSLPSPSTLP--HTITTFSKPQLRRISLPTSTTISLLTLFTPP 210
Query: 61 SLA 63
+ A
Sbjct: 211 NEA 219
>TC83868
Length = 828
Score = 30.0 bits (66), Expect = 0.66
Identities = 24/67 (35%), Positives = 39/67 (57%), Gaps = 3/67 (4%)
Frame = +3
Query: 120 SEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAE---LQEALA 176
S+EVK E + + A + ALNQ KE++AE+E + RKK D E L + +A
Sbjct: 303 SKEVKVDHE----IEKDAHLKEIEALNQRLKESEAEIESSREQLRKK-DFEITALHKMVA 467
Query: 177 NLEKQKE 183
N++K+++
Sbjct: 468 NIQKRQK 488
>TC77763 similar to GP|15215674|gb|AAK91382.1 AT4g27500/F27G19_100
{Arabidopsis thaliana}, partial (38%)
Length = 2297
Score = 30.0 bits (66), Expect = 0.66
Identities = 28/93 (30%), Positives = 46/93 (49%), Gaps = 1/93 (1%)
Frame = +1
Query: 96 TPLGTFMDNRDAEIRGKLNSVTNTSEEVKELEEQANAVLRAARAEIAVA-LNQMKKETQA 154
+P+ T ++ + +G+ + + E + ++A A A EI A L +MK+E +
Sbjct: 1453 SPVETLPAQKETKSKGRDLAAIDDDYEFENPHKEAPA---AKEPEIDPAKLKEMKREEEI 1623
Query: 155 EVEEKIAEGRKKVDAELQEALANLEKQKEETVK 187
+ K+A RKK AE A A L+ QKE K
Sbjct: 1624 -AKAKLAAERKKKMAEKAAAKAALKAQKEAEKK 1719
>TC79049 weakly similar to PIR|T10798|T10798 pherophorin-S - Volvox carteri,
partial (6%)
Length = 957
Score = 28.9 bits (63), Expect = 1.5
Identities = 17/42 (40%), Positives = 25/42 (59%), Gaps = 2/42 (4%)
Frame = -3
Query: 142 AVALNQMKKETQAEVEE--KIAEGRKKVDAELQEALANLEKQ 181
A L ++ K+ + VEE K EG KKV AE + + +LEK+
Sbjct: 913 AEGLKEVLKDKEGRVEELVKEVEGLKKVKAESEARVKDLEKR 788
>TC88377 similar to GP|8778315|gb|AAF79324.1| F14J16.29 {Arabidopsis
thaliana}, partial (23%)
Length = 644
Score = 28.9 bits (63), Expect = 1.5
Identities = 17/46 (36%), Positives = 23/46 (49%)
Frame = +1
Query: 16 PTNSLPSTPKLPSLQTITPNLKLQLSKLKSMTLAATSLSFVFAPPS 61
P S+P+ P LPSL P L L S++ +S+ F PPS
Sbjct: 316 PLPSIPTNPTLPSL-NFPPFPSNSLPNLPSISTIISSIPFFSPPPS 450
>TC78572 similar to GP|9801280|emb|CAC03581.1 putative 1-deoxy-D-xylulose
5-phosphate reductoisomerase {Zea mays}, partial (88%)
Length = 1654
Score = 28.9 bits (63), Expect = 1.5
Identities = 20/67 (29%), Positives = 37/67 (54%)
Frame = +2
Query: 117 TNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALA 176
T T + V E ++ V AA + I + ++Q+K+ + +A + + AELQEAL
Sbjct: 329 TQTLDIVAENPDKFRVVALAAGSNITLLVDQVKRFKP----QLVAVRNESLLAELQEALK 496
Query: 177 NLEKQKE 183
++E++ E
Sbjct: 497 DVEEKPE 517
>TC88376 similar to GP|8778315|gb|AAF79324.1| F14J16.29 {Arabidopsis
thaliana}, partial (25%)
Length = 706
Score = 28.1 bits (61), Expect = 2.5
Identities = 16/46 (34%), Positives = 23/46 (49%)
Frame = +1
Query: 16 PTNSLPSTPKLPSLQTITPNLKLQLSKLKSMTLAATSLSFVFAPPS 61
P S+P+ P LPSL P L + S++ +S+ F PPS
Sbjct: 334 PLPSIPTNPTLPSL-NFPPFPSNSLPNIPSISTIISSIPFFSPPPS 468
>AW773748 similar to GP|10176892|dbj Na+-dependent inorganic phosphate
cotransporter-like protein {Arabidopsis thaliana},
partial (20%)
Length = 596
Score = 27.7 bits (60), Expect = 3.3
Identities = 28/105 (26%), Positives = 49/105 (46%), Gaps = 15/105 (14%)
Frame = +1
Query: 5 IMASSKPIIRVPTNSLPS------------TPKLPSLQTITPNLKLQLS---KLKSMTLA 49
++ SK + RVP + + S T K P ++ N LQLS + +S LA
Sbjct: 286 MIQQSKNVTRVPQSKILSPVENNKNTRELKTHKSPGMK----NTSLQLSGVERSESDALA 453
Query: 50 ATSLSFVFAPPSLAFEKAALFDFNLTLPIIVVEFLFLMFALDKLY 94
A+S V + P++ + A+F+ +T+P + L+ +LY
Sbjct: 454 ASSNFPVRSTPTIPAKVPAVFETPITIPAYLGATSILIHCKPQLY 588
>BI270589 similar to PIR|C96829|C96 unknown protein F19K16.21 [imported] -
Arabidopsis thaliana, partial (20%)
Length = 651
Score = 27.7 bits (60), Expect = 3.3
Identities = 20/79 (25%), Positives = 39/79 (49%)
Frame = +3
Query: 109 IRGKLNSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVD 168
+R + ++ T E+ ++ ++ AA+ E + + + EE+I + R+K
Sbjct: 384 LRAEQTQLSRTLEKERQRAAESRQEYLAAKEEADTQEGRAR-----QFEEEIRDIRQKHK 548
Query: 169 AELQEALANLEKQKEETVK 187
ELQEAL + E ++E K
Sbjct: 549 QELQEALIHRELLQQEIEK 605
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.128 0.329
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,086,479
Number of Sequences: 36976
Number of extensions: 45684
Number of successful extensions: 351
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 345
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 349
length of query: 209
length of database: 9,014,727
effective HSP length: 92
effective length of query: 117
effective length of database: 5,612,935
effective search space: 656713395
effective search space used: 656713395
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC148405.2


