

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148405.1 + phase: 0
(149 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
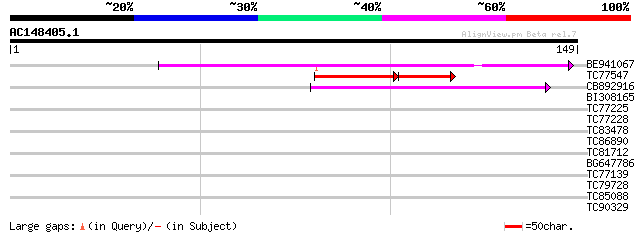
BE941067 weakly similar to GP|6648198|gb| unknown protein {Arabi... 66 6e-12
TC77547 similar to PIR|T48354|T48354 hypothetical protein F12E4.... 43 9e-07
CB892916 similar to GP|6648198|gb|A unknown protein {Arabidopsis... 44 2e-05
BI308165 30 0.48
TC77225 similar to SP|Q9LDU6|ST7R_ARATH 7-dehydrocholesterol red... 28 1.8
TC77228 similar to SP|Q9LDU6|ST7R_ARATH 7-dehydrocholesterol red... 28 1.8
TC83478 similar to PIR|H86304|H86304 hypothetical protein AAF998... 27 3.1
TC86890 weakly similar to GP|4406801|gb|AAD20110.1| unknown prot... 27 3.1
TC81712 similar to GP|23617251|dbj|BAC20918. putative transcript... 26 5.3
BG647786 similar to GP|21593300|gb| unknown {Arabidopsis thalian... 26 5.3
TC77139 weakly similar to GP|12324751|gb|AAG52333.1 unknown prot... 26 6.9
TC79728 similar to GP|22136692|gb|AAM91665.1 unknown protein {Ar... 25 9.0
TC85088 similar to PIR|G84539|G84539 hypothetical protein At2g16... 25 9.0
TC90329 GP|21314227|gb|AAM44081.1 type IIB calcium ATPase MCA5 {... 25 9.0
>BE941067 weakly similar to GP|6648198|gb| unknown protein {Arabidopsis
thaliana}, partial (59%)
Length = 601
Score = 65.9 bits (159), Expect = 6e-12
Identities = 36/115 (31%), Positives = 63/115 (54%), Gaps = 6/115 (5%)
Frame = +2
Query: 40 LTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK------ISWPLVWGQFCLCY 93
+ +TVLKLD++SF V + TAT+ LK A++ + + ISW VW +CL +
Sbjct: 104 MRITVLKLDTTSFDVILMNTATLKDLKLAIKKKVNYMEQSSMGHRHISWRSVWANYCLSF 283
Query: 94 EGQKLVTETDYLRDYGIKDGDQLRFIRHVSNVCNFQRKPKKRTVNLKQHGRYEFS 148
+ KL+ + D L++ G+++ Q+ F+ +V + R+ KR + HG + S
Sbjct: 284 DNNKLLNDDDVLQNLGVRNNSQIHFVPYV--MTKESRRHSKRRKHRFFHGLSKLS 442
>TC77547 similar to PIR|T48354|T48354 hypothetical protein F12E4.60 -
Arabidopsis thaliana, partial (53%)
Length = 2073
Score = 43.1 bits (100), Expect(2) = 9e-07
Identities = 18/22 (81%), Positives = 20/22 (90%)
Frame = +2
Query: 81 SWPLVWGQFCLCYEGQKLVTET 102
S PLVWG+FCLCYEG KLV+ET
Sbjct: 98 SRPLVWGKFCLCYEG*KLVSET 163
Score = 25.0 bits (53), Expect(2) = 9e-07
Identities = 8/15 (53%), Positives = 13/15 (86%)
Frame = +3
Query: 103 DYLRDYGIKDGDQLR 117
+YL++YG K GDQ++
Sbjct: 165 NYLKNYGFKSGDQVK 209
>CB892916 similar to GP|6648198|gb|A unknown protein {Arabidopsis thaliana},
partial (25%)
Length = 388
Score = 43.9 bits (102), Expect = 2e-05
Identities = 18/63 (28%), Positives = 32/63 (50%)
Frame = +2
Query: 80 ISWPLVWGQFCLCYEGQKLVTETDYLRDYGIKDGDQLRFIRHVSNVCNFQRKPKKRTVNL 139
ISW VW +CL ++ KL + D L++ G+++ Q+ F+ V K +R
Sbjct: 74 ISWRSVWANYCLSFDNNKLFNDDDVLQNLGVRNNSQIHFVCRYIFVEYVMTKESRRNSKR 253
Query: 140 KQH 142
++H
Sbjct: 254 RKH 262
>BI308165
Length = 462
Score = 29.6 bits (65), Expect = 0.48
Identities = 20/66 (30%), Positives = 29/66 (43%), Gaps = 2/66 (3%)
Frame = -3
Query: 80 ISWPLVWGQFCLCY--EGQKLVTETDYLRDYGIKDGDQLRFIRHVSNVCNFQRKPKKRTV 137
IS ++ G CLCY EG +L ET + YG K+ F + + C + K R +
Sbjct: 382 ISVGVLKGCLCLCYVIEGARL--ETWVMSKYGEKESWSKAFSIEIKSYCGLSPQDKHRPI 209
Query: 138 NLKQHG 143
G
Sbjct: 208 GFNTFG 191
>TC77225 similar to SP|Q9LDU6|ST7R_ARATH 7-dehydrocholesterol reductase (EC
1.3.1.21) (7-DHC reductase) (Sterol delta-7-reductase),
partial (54%)
Length = 1018
Score = 27.7 bits (60), Expect = 1.8
Identities = 12/36 (33%), Positives = 20/36 (55%)
Frame = -1
Query: 6 SVSNESPRRSLSIQLSSMIIVDCTLPYDKLPSQNLT 41
+ +++P+ I L MII DC + Y K P N++
Sbjct: 700 NTKDKAPKAECGIYLPQMIINDCWVEYPKPPKCNVS 593
>TC77228 similar to SP|Q9LDU6|ST7R_ARATH 7-dehydrocholesterol reductase (EC
1.3.1.21) (7-DHC reductase) (Sterol delta-7-reductase),
partial (30%)
Length = 819
Score = 27.7 bits (60), Expect = 1.8
Identities = 12/36 (33%), Positives = 20/36 (55%)
Frame = -3
Query: 6 SVSNESPRRSLSIQLSSMIIVDCTLPYDKLPSQNLT 41
+ +++P+ I L MII DC + Y K P N++
Sbjct: 724 NTKDKAPKXECGIYLPQMIINDCWVEYPKPPKCNVS 617
>TC83478 similar to PIR|H86304|H86304 hypothetical protein AAF99838.1
[imported] - Arabidopsis thaliana, partial (3%)
Length = 906
Score = 26.9 bits (58), Expect = 3.1
Identities = 30/93 (32%), Positives = 41/93 (43%), Gaps = 13/93 (13%)
Frame = +2
Query: 3 ELTSVSNESPRRSLS-----IQLSSMIIVDCTLPYDKL-PSQNLTLTVLKLDSSSFQVEV 56
+LT V N+ SL+ IQ +M V L D + PSQ + L LKLD +
Sbjct: 365 DLTLVLNQDGVSSLNKNLPEIQRLTMGFVSKMLYADIIHPSQLIGLKYLKLDRVNLDERE 544
Query: 57 AKTATVAVLKQA-------VEAAFRHKPEKISW 82
V+VLK A +E RHK ++ W
Sbjct: 545 ELLYIVSVLKSASNLVELVIEVVIRHK-RRVVW 640
>TC86890 weakly similar to GP|4406801|gb|AAD20110.1| unknown protein
{Arabidopsis thaliana}, partial (54%)
Length = 2446
Score = 26.9 bits (58), Expect = 3.1
Identities = 12/25 (48%), Positives = 14/25 (56%), Gaps = 2/25 (8%)
Frame = +2
Query: 76 KPEKISWPL--VWGQFCLCYEGQKL 98
K + ISWPL VW C C+ G L
Sbjct: 1127 KADSISWPLCMVWSVACYCWIGDVL 1201
>TC81712 similar to GP|23617251|dbj|BAC20918. putative transcription
elongation factor {Oryza sativa (japonica
cultivar-group)}, partial (34%)
Length = 721
Score = 26.2 bits (56), Expect = 5.3
Identities = 21/58 (36%), Positives = 28/58 (48%)
Frame = -2
Query: 41 TLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEKISWPLVWGQFCLCYEGQKL 98
T T LKL SSSF ++ A T VE +F + PL+ FC ++GQ L
Sbjct: 423 TPTTLKLISSSFWLKSALT---------VEQSFLLDFLILLLPLLTSHFC*AHDGQPL 277
>BG647786 similar to GP|21593300|gb| unknown {Arabidopsis thaliana}, partial
(12%)
Length = 801
Score = 26.2 bits (56), Expect = 5.3
Identities = 22/81 (27%), Positives = 36/81 (44%), Gaps = 16/81 (19%)
Frame = -2
Query: 2 SELTSVSNESPRRSLSIQLSSMIIVDCTLPYDKLPSQ---------------NLTLTVLK 46
+EL V NESP SL +++ D LP+ + N+ L +LK
Sbjct: 614 NELPMVVNESPELSLLRAMATWSFEDKRLPFKSAQPEDLASISFSTREHI*LNVFLEILK 435
Query: 47 LDSS-SFQVEVAKTATVAVLK 66
DS+ S +++AK + + K
Sbjct: 434 TDSNFSISLDLAKGPVILLEK 372
>TC77139 weakly similar to GP|12324751|gb|AAG52333.1 unknown protein;
13405-15968 {Arabidopsis thaliana}, partial (55%)
Length = 2340
Score = 25.8 bits (55), Expect = 6.9
Identities = 22/91 (24%), Positives = 41/91 (44%), Gaps = 2/91 (2%)
Frame = +1
Query: 27 DCTLPYDKLPSQNL--TLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEKISWPL 84
+C ++K+ +N+ ++ +LK S ++ + K + L++ ++ FR K EK
Sbjct: 1510 ECYKLWEKVYQENIVASVAILKKLSDDWKEQATKLSPYEPLREILKN-FRQKNEKA---- 1674
Query: 85 VWGQFCLCYEGQKLVTETDYLRDYGIKDGDQ 115
L TETD R +KD D+
Sbjct: 1675 -------------LTTETDAARQVLVKDADK 1728
>TC79728 similar to GP|22136692|gb|AAM91665.1 unknown protein {Arabidopsis
thaliana}, partial (79%)
Length = 1121
Score = 25.4 bits (54), Expect = 9.0
Identities = 10/29 (34%), Positives = 16/29 (54%)
Frame = -2
Query: 117 RFIRHVSNVCNFQRKPKKRTVNLKQHGRY 145
RFI ++N C+F++ +K T K Y
Sbjct: 1105 RFILFINNPCSFRKSVRKHTDECKYKSSY 1019
>TC85088 similar to PIR|G84539|G84539 hypothetical protein At2g16390
[imported] - Arabidopsis thaliana, partial (17%)
Length = 653
Score = 25.4 bits (54), Expect = 9.0
Identities = 8/24 (33%), Positives = 17/24 (70%)
Frame = +1
Query: 103 DYLRDYGIKDGDQLRFIRHVSNVC 126
D++ D ++DG + +F R++ N+C
Sbjct: 403 DFIADLDMRDGVKSKFFRNMLNLC 474
>TC90329 GP|21314227|gb|AAM44081.1 type IIB calcium ATPase MCA5 {Medicago
truncatula}, partial (5%)
Length = 547
Score = 25.4 bits (54), Expect = 9.0
Identities = 18/53 (33%), Positives = 28/53 (51%), Gaps = 6/53 (11%)
Frame = +1
Query: 97 KLVTETDYLRDY--GIKDGDQLR-FIRHVSNVCNFQRKPKKR---TVNLKQHG 143
KL T +YL++ G+K + +R +VC F + PK+R T NL + G
Sbjct: 82 KLRTMENYLQENFGGVKSKNSSEEALRRWRDVCGFVKNPKRRFRFTANLDKRG 240
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.133 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,675,361
Number of Sequences: 36976
Number of extensions: 60632
Number of successful extensions: 318
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 317
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 317
length of query: 149
length of database: 9,014,727
effective HSP length: 87
effective length of query: 62
effective length of database: 5,797,815
effective search space: 359464530
effective search space used: 359464530
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC148405.1


