

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148404.10 + phase: 0
(400 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
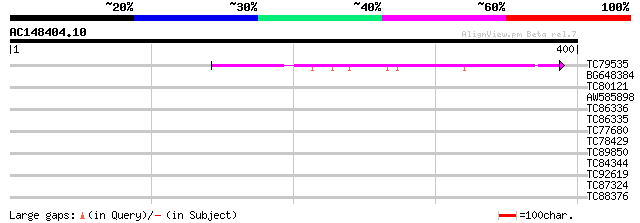
Score E
Sequences producing significant alignments: (bits) Value
TC79535 weakly similar to GP|10178054|dbj|BAB11418. gene_id:MUB3... 86 3e-17
BG648384 similar to GP|14718995|gb ceramide glucosyltransferase ... 29 2.8
TC80121 similar to SP|Q43111|PME3_PHAVU Pectinesterase 3 precurs... 29 2.8
AW585898 weakly similar to GP|15983422|gb| At2g19880/F6F22.9 {Ar... 29 2.8
TC86336 similar to GP|14626761|gb|AAK71662.1 mature anther-speci... 29 3.7
TC86335 similar to GP|14626761|gb|AAK71662.1 mature anther-speci... 29 3.7
TC77680 similar to GP|18650600|gb|AAL75900.1 AT4g38220/F20D10_34... 28 4.8
TC78429 similar to GP|16604607|gb|AAL24096.1 putative AtBgamma p... 28 4.8
TC89850 similar to GP|22748338|gb|AAN05340.1 Unknown protein {Or... 28 6.3
TC84344 28 6.3
TC92619 similar to PIR|T47763|T47763 hypothetical protein F24I3.... 28 6.3
TC87324 homologue to PIR|E86217|E86217 protein T27G7.10 [importe... 28 8.2
TC88376 similar to GP|8778315|gb|AAF79324.1| F14J16.29 {Arabidop... 28 8.2
>TC79535 weakly similar to GP|10178054|dbj|BAB11418.
gene_id:MUB3.3~pir||T03813~similar to unknown protein
{Arabidopsis thaliana}, partial (86%)
Length = 1356
Score = 85.9 bits (211), Expect = 3e-17
Identities = 75/286 (26%), Positives = 132/286 (45%), Gaps = 37/286 (12%)
Frame = +2
Query: 143 ESHHLSLSLPMNVSYNGLKHIIVGKGITVEVRRAREISFYYQSDLDLQRNGSVICSNQKN 202
++ + LSLP +V LK +++ G V V+ AR +S + L L N S +N
Sbjct: 446 DAKDMRLSLPHDVDAGVLKKVVLADGAVVTVKGARSVSLRHPLTLPLPLNRS------QN 607
Query: 203 EFWPFLQSMC-----------VPLIPIRIIGSASL----IAYVARNPYVQI-----GTAL 242
F L ++ PL+ +RI+G SL A + N +++ G
Sbjct: 608 GFAAGLLTLAEHLRHASRGQDAPLLSLRIVGPTSLEAPSSASTSSNNRLKLKRLAPGLVE 787
Query: 243 ISEDAVELLPEKCYHGCVFRKQA------------CPVASLN---LRLILLEKILRSLLG 287
+S + L + +++A P+ASLN L+ E++L S+LG
Sbjct: 788 LSSQSKSKLVDTSLSTVDLQEEAPTLLTPTQFTALWPLASLNGSNANLLGFERLLSSVLG 967
Query: 288 HKILQDRLSGLIKANIKAYAGVKFPLELERDV--GNNATLSTLPDWRTRPSVERVWFEVM 345
K + L+KA++ A VK + E+ + G+ + P+WRT+P R+ FEV+
Sbjct: 968 PKANEKGSFRLLKADVSAQTFVKIGFQAEKKLKEGDGISFEGFPEWRTKPDTVRLHFEVL 1147
Query: 346 ARVEDSRLKPLSIKKVKPFIESDSVSWANLMSNLSYTKLRPVLLPP 391
A+V+ ++ P + +V P + DSV+ N+++N P++ PP
Sbjct: 1148AKVDGDKVIPERVMQVNPVVTEDSVA-PNMLTNNGTMSKMPLVQPP 1282
>BG648384 similar to GP|14718995|gb ceramide glucosyltransferase {Gossypium
arboreum}, partial (26%)
Length = 784
Score = 29.3 bits (64), Expect = 2.8
Identities = 11/19 (57%), Positives = 15/19 (78%)
Frame = +2
Query: 135 GPFELRVHESHHLSLSLPM 153
GP ELR+H H++LSLP+
Sbjct: 113 GPVELRIHPLPHINLSLPL 169
>TC80121 similar to SP|Q43111|PME3_PHAVU Pectinesterase 3 precursor (EC
3.1.1.11) (Pectin methylesterase 3) (PE 3)., partial
(5%)
Length = 792
Score = 29.3 bits (64), Expect = 2.8
Identities = 18/54 (33%), Positives = 28/54 (51%)
Frame = -3
Query: 34 SSTHSNITHIFQDILKAISSRQKWDLNDVRVFNFDVAKIRFGTSQNYLFRIGSS 87
SST T++FQ I + RQ+ + + NF + + R+GT N+ RI S
Sbjct: 616 SSTFQRNTNLFQPINTRLLIRQRSTNPRLNIINFTICQ-RYGTVSNHRHRIVES 458
>AW585898 weakly similar to GP|15983422|gb| At2g19880/F6F22.9 {Arabidopsis
thaliana}, partial (12%)
Length = 622
Score = 29.3 bits (64), Expect = 2.8
Identities = 11/19 (57%), Positives = 15/19 (78%)
Frame = +1
Query: 135 GPFELRVHESHHLSLSLPM 153
GP ELR+H H++LSLP+
Sbjct: 253 GPVELRIHPLPHINLSLPL 309
>TC86336 similar to GP|14626761|gb|AAK71662.1 mature anther-specific
protein LAT61 {Lycopersicon esculentum}, partial (37%)
Length = 778
Score = 28.9 bits (63), Expect = 3.7
Identities = 15/29 (51%), Positives = 17/29 (57%)
Frame = +1
Query: 14 FFIIFFLQLFTAFSNASSSSSSTHSNITH 42
FF IF L T F+ SSS S HSN +H
Sbjct: 25 FFFIFSSSLLTNFNIIFSSSFSRHSNNSH 111
>TC86335 similar to GP|14626761|gb|AAK71662.1 mature anther-specific
protein LAT61 {Lycopersicon esculentum}, partial (60%)
Length = 1595
Score = 28.9 bits (63), Expect = 3.7
Identities = 15/29 (51%), Positives = 17/29 (57%)
Frame = +2
Query: 14 FFIIFFLQLFTAFSNASSSSSSTHSNITH 42
FF IF L T F+ SSS S HSN +H
Sbjct: 41 FFFIFSSSLLTNFNIIFSSSFSRHSNNSH 127
>TC77680 similar to GP|18650600|gb|AAL75900.1 AT4g38220/F20D10_340
{Arabidopsis thaliana}, partial (86%)
Length = 1433
Score = 28.5 bits (62), Expect = 4.8
Identities = 17/43 (39%), Positives = 24/43 (55%)
Frame = +2
Query: 7 LRLHQLPFFIIFFLQLFTAFSNASSSSSSTHSNITHIFQDILK 49
++L++ F+I FL +SSSSSS SNI FQ L+
Sbjct: 59 MKLNRGSIFLIAFLLSTLTSLQSSSSSSSEESNIISRFQQYLQ 187
>TC78429 similar to GP|16604607|gb|AAL24096.1 putative AtBgamma protein
{Arabidopsis thaliana}, partial (62%)
Length = 1596
Score = 28.5 bits (62), Expect = 4.8
Identities = 16/29 (55%), Positives = 20/29 (68%), Gaps = 3/29 (10%)
Frame = +3
Query: 15 FIIFFLQLFTAF---SNASSSSSSTHSNI 40
F+ FFL LFT N+SSSSSS+ S+I
Sbjct: 186 FLFFFLSLFTTSPPNPNSSSSSSSSSSSI 272
>TC89850 similar to GP|22748338|gb|AAN05340.1 Unknown protein {Oryza sativa
(japonica cultivar-group)}, partial (34%)
Length = 1093
Score = 28.1 bits (61), Expect = 6.3
Identities = 12/37 (32%), Positives = 20/37 (53%)
Frame = +1
Query: 15 FIIFFLQLFTAFSNASSSSSSTHSNITHIFQDILKAI 51
FI+FF Q+F A S+ S + + H+ ++K I
Sbjct: 766 FIVFFNQIFDALCKLSADSDANVQSAAHLLDRLVKDI 876
>TC84344
Length = 769
Score = 28.1 bits (61), Expect = 6.3
Identities = 14/52 (26%), Positives = 25/52 (47%), Gaps = 2/52 (3%)
Frame = +3
Query: 65 FNFDVAKIRFGTSQNYLFRIGSSKNNFTVKFSDEISSWNHN--KFTTTPKPD 114
F+F ++ RF T+ + F + NNF + ++N+ +F T PD
Sbjct: 177 FSFPISLTRFCTTTTHPFAVSYLVNNFGFSTESTLKAFNNKQVRFNTPDNPD 332
>TC92619 similar to PIR|T47763|T47763 hypothetical protein F24I3.110 -
Arabidopsis thaliana, partial (43%)
Length = 682
Score = 28.1 bits (61), Expect = 6.3
Identities = 15/27 (55%), Positives = 18/27 (66%)
Frame = +3
Query: 18 FFLQLFTAFSNASSSSSSTHSNITHIF 44
FFLQLF+ +S SSS STH + H F
Sbjct: 60 FFLQLFS-YSPPSSSPYSTHKTLFHQF 137
>TC87324 homologue to PIR|E86217|E86217 protein T27G7.10 [imported] -
Arabidopsis thaliana, partial (37%)
Length = 1897
Score = 27.7 bits (60), Expect = 8.2
Identities = 19/59 (32%), Positives = 27/59 (45%), Gaps = 6/59 (10%)
Frame = +2
Query: 114 DLASLVDQLSS------IAFLDYIKLEGPFELRVHESHHLSLSLPMNVSYNGLKHIIVG 166
DL L D+ S IA++DY+ L + H +SL L + V Y H+I G
Sbjct: 425 DLMRLFDEYGSPSTAGDIAYIDYLFLGDYVDRGQHSLETISLLLALKVEYPNNIHLIRG 601
>TC88376 similar to GP|8778315|gb|AAF79324.1| F14J16.29 {Arabidopsis
thaliana}, partial (25%)
Length = 706
Score = 27.7 bits (60), Expect = 8.2
Identities = 12/25 (48%), Positives = 17/25 (68%)
Frame = +3
Query: 20 LQLFTAFSNASSSSSSTHSNITHIF 44
LQ++T S+ +SSS + NI HIF
Sbjct: 6 LQIYTLISSINSSSHGLNENIHHIF 80
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,195,127
Number of Sequences: 36976
Number of extensions: 192431
Number of successful extensions: 1402
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 1357
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1387
length of query: 400
length of database: 9,014,727
effective HSP length: 98
effective length of query: 302
effective length of database: 5,391,079
effective search space: 1628105858
effective search space used: 1628105858
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC148404.10


