

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148360.3 - phase: 0
(301 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
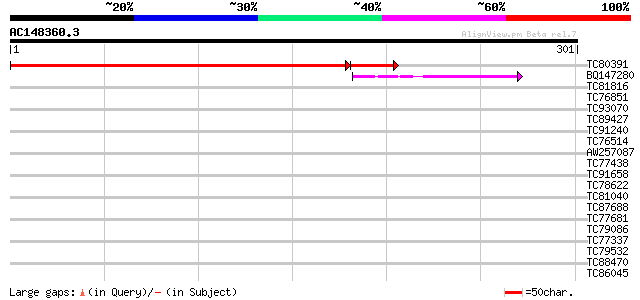
Sequences producing significant alignments: (bits) Value
TC80391 similar to GP|21553422|gb|AAM62515.1 unknown {Arabidopsi... 360 e-108
BQ147280 similar to PIR|C86333|C863 hypothetical protein AAF7991... 40 8e-04
TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana taba... 37 0.012
TC76851 homologue to PIR|S40122|S40122 high mobility group prote... 36 0.016
TC93070 weakly similar to GP|17065230|gb|AAL32769.1 Unknown prot... 35 0.046
TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed pr... 33 0.10
TC91240 homologue to PIR|T48518|T48518 transcription factor like... 33 0.10
TC76514 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - ... 33 0.10
AW257087 homologue to GP|6683126|dbj KIAA0339 protein {Homo sapi... 33 0.10
TC77438 homologue to PIR|T06807|T06807 nucleosome assembly prote... 33 0.13
TC91658 similar to GP|14329812|emb|CAC40753. putative nucleosome... 33 0.13
TC78622 weakly similar to GP|14587220|dbj|BAB61154. putative GTP... 33 0.13
TC81040 33 0.13
TC87688 similar to GP|10177535|dbj|BAB10930. gene_id:K1F13.21~un... 33 0.13
TC77681 similar to GP|22597156|gb|AAN03465.1 nucleolar histone d... 33 0.18
TC79086 similar to GP|23499033|emb|CAD51113. ubiquitin-protein l... 33 0.18
TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Atel... 33 0.18
TC79532 weakly similar to GP|8164030|gb|AAF73960.1| SGS3 {Arabid... 32 0.23
TC88470 weakly similar to GP|15146334|gb|AAK83650.1 At1g07080/F1... 32 0.30
TC86045 similar to PIR|T46225|T46225 alpha NAC-like protein - Ar... 32 0.30
>TC80391 similar to GP|21553422|gb|AAM62515.1 unknown {Arabidopsis
thaliana}, partial (52%)
Length = 708
Score = 360 bits (923), Expect(2) = e-108
Identities = 176/181 (97%), Positives = 177/181 (97%)
Frame = +1
Query: 1 MKGVFWFLVVSVVIASWIPLSHCAKKPVGIARKEDVPYIKCQVCEILAKQLYQQVQSKKA 60
MKGVFWFLVVSVVIASWIPLSHCAKKPVGIARKEDVPYIKCQVCEILAKQLYQQVQSKKA
Sbjct: 88 MKGVFWFLVVSVVIASWIPLSHCAKKPVGIARKEDVPYIKCQVCEILAKQLYQQVQSKKA 267
Query: 61 EISPKKISEYQIIEIAENVCNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTV 120
EISPKKISEYQIIEIAENVCNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSE TV
Sbjct: 268 EISPKKISEYQIIEIAENVCNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSE**TV 447
Query: 121 ERACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPFVA 180
ERACQEV+GYSDTDVAEYLYSSKPDIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPF
Sbjct: 448 ERACQEVLGYSDTDVAEYLYSSKPDIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPFCC 627
Query: 181 K 181
K
Sbjct: 628 K 630
Score = 49.7 bits (117), Expect(2) = e-108
Identities = 24/25 (96%), Positives = 24/25 (96%)
Frame = +3
Query: 182 SSKEAEMEKLLKSMEGMPGAPGMKM 206
SSKEAEMEKLLKSMEGMP APGMKM
Sbjct: 633 SSKEAEMEKLLKSMEGMPRAPGMKM 707
>BQ147280 similar to PIR|C86333|C863 hypothetical protein AAF79914.1
[imported] - Arabidopsis thaliana, partial (10%)
Length = 666
Score = 40.4 bits (93), Expect = 8e-04
Identities = 31/90 (34%), Positives = 46/90 (50%)
Frame = +2
Query: 183 SKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLG 242
SK ++++KLLK E P + SR + K E +DDD+D DE++D+ K+
Sbjct: 176 SKGSKLKKLLKE-ETEPISRATSA-SRSKVKK-----ELKDDDDDSDEDDDDKPIAKKIS 334
Query: 243 KILKSKENEKGDWKQKIRKGIVDTSTTLKK 272
K KE K K K + +V T T +KK
Sbjct: 335 KTKVVKEEVKKKKKVKKEEEVVVTETKVKK 424
>TC81816 similar to GP|8096269|dbj|BAA95789.1 KED {Nicotiana tabacum},
partial (13%)
Length = 663
Score = 36.6 bits (83), Expect = 0.012
Identities = 36/127 (28%), Positives = 55/127 (42%), Gaps = 2/127 (1%)
Frame = +3
Query: 170 KDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAP--GMKMYSRDDLMKKNFGAENEDDDED 227
++ GE K+ E +K K +G G + S+ D KK +NEDDDE
Sbjct: 279 EENVKGEEEDGDEKKDKEKKKKEKKEKGKEDKDKDGEEKKSKKDKEKKK--DKNEDDDEG 452
Query: 228 EDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKG 287
ED + + +K K K +E+EK + K +R +D T K+ K + K
Sbjct: 453 EDGSKKK---KNKDKKEKKKEEDEKEEGKVSVRD--IDIEETAKEGKEKKKKKEDKEEKK 617
Query: 288 KKTTSSK 294
KK+ K
Sbjct: 618 KKSGKDK 638
>TC76851 homologue to PIR|S40122|S40122 high mobility group protein HMG-1 -
garden pea, partial (92%)
Length = 863
Score = 36.2 bits (82), Expect = 0.016
Identities = 23/67 (34%), Positives = 35/67 (51%), Gaps = 1/67 (1%)
Frame = +2
Query: 177 PFVAKSSK-EAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEA 235
P+ AK+ K +AE EK +K+ A G ++ K +E++D+DED E+E
Sbjct: 377 PYAAKAEKRKAEYEKTMKAYNKKQ-AEGPAAVEEEESGKSESEVHDENEDDDEDGSEEEE 553
Query: 236 DFPSKLG 242
D KLG
Sbjct: 554 DDE*KLG 574
>TC93070 weakly similar to GP|17065230|gb|AAL32769.1 Unknown protein
{Arabidopsis thaliana}, partial (19%)
Length = 679
Score = 34.7 bits (78), Expect = 0.046
Identities = 16/47 (34%), Positives = 28/47 (59%)
Frame = +1
Query: 190 KLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEAD 236
K+LK+ + Y DD +++ ++ED+DEDEDE++D+ D
Sbjct: 166 KILKTNRKSTYGKFLSPYDSDDEIEEMDFEDDEDEDEDEDEDDDDDD 306
Score = 31.6 bits (70), Expect = 0.39
Identities = 27/106 (25%), Positives = 48/106 (44%), Gaps = 6/106 (5%)
Frame = +1
Query: 177 PFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEAD 236
P + K+++++ K L + M DD + E++DDD+D+D+E+D+ D
Sbjct: 163 PKILKTNRKSTYGKFLSPYDSDDEIEEMDF--EDDEDEDEDEDEDDDDDDDDDDEDDDDD 336
Query: 237 ---FPSKLG---KILKSKENEKGDWKQKIRKGIVDTSTTLKKHATK 276
P+ L K LKSK +++ KG+ + K K
Sbjct: 337 GFAEPTDLNAKDKRLKSKTVPDRQQEKEKEKGVRSLNNGQSKRLPK 474
>TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed protein
{Arabidopsis thaliana}, partial (50%)
Length = 753
Score = 33.5 bits (75), Expect = 0.10
Identities = 18/44 (40%), Positives = 24/44 (53%)
Frame = +1
Query: 200 GAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGK 243
GA G DD + + ED+DEDE+E++DE PSK K
Sbjct: 433 GAGGSDDDDEDDDDDDDDNDDGEDEDEDEEEDDDEDQPPSKKKK 564
Score = 28.5 bits (62), Expect = 3.3
Identities = 9/27 (33%), Positives = 18/27 (66%)
Frame = +1
Query: 210 DDLMKKNFGAENEDDDEDEDEEEDEAD 236
DD + G ++DD++D+D+++D D
Sbjct: 415 DDPVPNGAGGSDDDDEDDDDDDDDNDD 495
>TC91240 homologue to PIR|T48518|T48518 transcription factor like protein -
Arabidopsis thaliana, partial (7%)
Length = 680
Score = 33.5 bits (75), Expect = 0.10
Identities = 17/62 (27%), Positives = 35/62 (56%), Gaps = 1/62 (1%)
Frame = +3
Query: 222 EDDDEDEDEEEDEADF-PSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNH 280
ED++E+E+EEE++ADF P L + L + ++ +VD+ ++ K++++
Sbjct: 270 EDNEEEEEEEEEDADFNPLFLKETLSDASSSLSSEGDRLDHNVVDSGPSMDIDLEKITDN 449
Query: 281 IQ 282
Q
Sbjct: 450 EQ 455
>TC76514 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - alfalfa,
partial (61%)
Length = 1288
Score = 33.5 bits (75), Expect = 0.10
Identities = 40/153 (26%), Positives = 60/153 (39%), Gaps = 3/153 (1%)
Frame = +3
Query: 105 EEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCKD---LSKAC 161
EE DSE + + K +A G + A S + D D ++ +D +KA
Sbjct: 528 EEEDSEESSDEDKKPAAKAVPSKNGSAPAKKAA---SDEEDTDESSDEDEEDEKPAAKAV 698
Query: 162 NTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAEN 221
+K VP + E S E +K P A K S KK + +
Sbjct: 699 PSKNGSVPAKKADTESSDEDSESSDEEDK-------KPAAKASKNVSAPT--KKAASSSD 851
Query: 222 EDDDEDEDEEEDEADFPSKLGKILKSKENEKGD 254
E+ DE+ DE+ED A SK + K + + D
Sbjct: 852 EESDEESDEDED-AKPVSKPAAVAKKSKKDSSD 947
>AW257087 homologue to GP|6683126|dbj KIAA0339 protein {Homo sapiens},
partial (2%)
Length = 466
Score = 33.5 bits (75), Expect = 0.10
Identities = 16/32 (50%), Positives = 23/32 (71%)
Frame = -3
Query: 220 ENEDDDEDEDEEEDEADFPSKLGKILKSKENE 251
E+ED+DEDEDE+EDE D ++L+ E+E
Sbjct: 368 EDEDEDEDEDEDEDEDD-----EELLEESESE 288
>TC77438 homologue to PIR|T06807|T06807 nucleosome assembly protein 1 - garden
pea, partial (93%)
Length = 1607
Score = 33.1 bits (74), Expect = 0.13
Identities = 16/31 (51%), Positives = 19/31 (60%)
Frame = +2
Query: 222 EDDDEDEDEEEDEADFPSKLGKILKSKENEK 252
EDDDEDEDE+EDE D + K K + K
Sbjct: 1070 EDDDEDEDEDEDEDDDEDEEETKTKKKSSAK 1162
Score = 32.0 bits (71), Expect = 0.30
Identities = 14/35 (40%), Positives = 22/35 (62%)
Frame = +2
Query: 219 AENEDDDEDEDEEEDEADFPSKLGKILKSKENEKG 253
AE++D+DEDEDE+ED+ + + KS + G
Sbjct: 1067 AEDDDEDEDEDEDEDDDEDEEETKTKKKSSAKKSG 1171
Score = 29.6 bits (65), Expect = 1.5
Identities = 13/35 (37%), Positives = 22/35 (62%)
Frame = +2
Query: 220 ENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGD 254
E+EDDDEDE+E + + +K I + E ++G+
Sbjct: 1100 EDEDDDEDEEETKTKKKSSAKKSGIAQLGEGQQGE 1204
Score = 29.6 bits (65), Expect = 1.5
Identities = 13/31 (41%), Positives = 20/31 (63%)
Frame = +2
Query: 210 DDLMKKNFGAENEDDDEDEDEEEDEADFPSK 240
DD + + E+ED+DED+DE+E+E K
Sbjct: 1058 DDDAEDDDEDEDEDEDEDDDEDEEETKTKKK 1150
Score = 29.6 bits (65), Expect = 1.5
Identities = 21/75 (28%), Positives = 36/75 (48%), Gaps = 3/75 (4%)
Frame = +2
Query: 165 PPPVPKDRTP-GEPFVAKSSKEAEMEKLLKSMEGMPGAP-GMKMYSRDDLMKKNFG-AEN 221
PP VP+D E + + E + + S P + ++ + + FG ++
Sbjct: 863 PPEVPEDDEDIDEDMAEELQNQMEQDYDIGSTIRDKIIPHAVSWFTGEAAQGEEFGDLDD 1042
Query: 222 EDDDEDEDEEEDEAD 236
EDDDED+D E+D+ D
Sbjct: 1043EDDDEDDDAEDDDED 1087
>TC91658 similar to GP|14329812|emb|CAC40753. putative nucleosome assembly
protein 1 {Atropa belladonna}, partial (46%)
Length = 583
Score = 33.1 bits (74), Expect = 0.13
Identities = 15/43 (34%), Positives = 26/43 (59%)
Frame = +2
Query: 220 ENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKG 262
+++DDD+D+D+EEDE D +E ++G K K ++G
Sbjct: 116 DDDDDDDDDDDEEDEDD-----------EEEDEGKGKSKSKRG 211
Score = 28.5 bits (62), Expect = 3.3
Identities = 15/30 (50%), Positives = 21/30 (70%), Gaps = 1/30 (3%)
Frame = +2
Query: 220 ENEDDDEDE-DEEEDEADFPSKLGKILKSK 248
+++DD+EDE DEEEDE SK + K+K
Sbjct: 134 DDDDDEEDEDDEEEDEGKGKSKSKRGSKAK 223
>TC78622 weakly similar to GP|14587220|dbj|BAB61154. putative GTPase {Oryza
sativa (japonica cultivar-group)}, partial (32%)
Length = 1025
Score = 33.1 bits (74), Expect = 0.13
Identities = 27/75 (36%), Positives = 36/75 (48%), Gaps = 6/75 (8%)
Frame = +1
Query: 211 DLMKKNFGAENEDD-DEDEDEEEDEADFPSKLGKILKSKENEK-----GDWKQKIRKGIV 264
+L K++ G EN DD DEDE E + D + K SK+NEK G K+R+
Sbjct: 415 ELAKQDEGPENVDDNDEDESMECGKEDVSKEKVKSASSKQNEKLYTAEGILNPKLRR--- 585
Query: 265 DTSTTLKKHATKVSN 279
KK A K S+
Sbjct: 586 -AEKKKKKKANKASS 627
>TC81040
Length = 478
Score = 33.1 bits (74), Expect = 0.13
Identities = 15/16 (93%), Positives = 16/16 (99%)
Frame = +2
Query: 286 KGKKTTSSKKNSKSEL 301
+GKKTTSSKKNSKSEL
Sbjct: 11 RGKKTTSSKKNSKSEL 58
>TC87688 similar to GP|10177535|dbj|BAB10930. gene_id:K1F13.21~unknown
protein {Arabidopsis thaliana}, partial (46%)
Length = 2077
Score = 33.1 bits (74), Expect = 0.13
Identities = 21/88 (23%), Positives = 43/88 (48%), Gaps = 1/88 (1%)
Frame = +2
Query: 214 KKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIR-KGIVDTSTTLKK 272
K G+E EDD E+ED+E++E E+++ D K+K++ GI D + +
Sbjct: 449 KAKGGSEGEDDFEEEDDEDEEG----------SEDEDDEEDEKEKVKGGGIEDKFLKIDE 598
Query: 273 HATKVSNHIQRWWKGKKTTSSKKNSKSE 300
+ + KG++ + ++S+ +
Sbjct: 599 LTEYLEKEEDNYEKGEERDEADEDSEED 682
Score = 32.7 bits (73), Expect = 0.18
Identities = 26/96 (27%), Positives = 46/96 (47%), Gaps = 6/96 (6%)
Frame = +2
Query: 210 DDLMKK--NFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKG----DWKQKIRKGI 263
DD ++K F ++EDDD+D++E+E+ D + + +NEKG D ++
Sbjct: 680 DDELEKAGEFEMDDEDDDDDDEEDEEAEDMGNVRYEDFFGGKNEKGSKRKDQLLEVSGDE 859
Query: 264 VDTSTTLKKHATKVSNHIQRWWKGKKTTSSKKNSKS 299
D +T +K T + QR GK + + +S
Sbjct: 860 DDMESTKQKKRTASTYEKQR---GKNSIKDSADGES 958
>TC77681 similar to GP|22597156|gb|AAN03465.1 nucleolar histone deacetylase
HD2-P39 {Glycine max}, partial (47%)
Length = 1233
Score = 32.7 bits (73), Expect = 0.18
Identities = 26/92 (28%), Positives = 36/92 (38%)
Frame = +2
Query: 208 SRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTS 267
S DD M+ ++DD DED+EE+ GK ++ K K K
Sbjct: 614 SSDDEMENADSDSEDEDDSDEDDEEETPVKKVDQGKKRPNESASKTPISSKKSKNATPEK 793
Query: 268 TTLKKHATKVSNHIQRWWKGKKTTSSKKNSKS 299
T KK + H ++ GK S K KS
Sbjct: 794 TDGKKAGHTATPHPKK--GGKTPNSDAKTPKS 883
Score = 27.3 bits (59), Expect = 7.4
Identities = 14/53 (26%), Positives = 27/53 (50%), Gaps = 2/53 (3%)
Frame = -2
Query: 22 HCAKKPVGIARKEDVPYIKCQVCEILAKQLYQQVQSKKAEIS--PKKISEYQI 72
HC + I R E C++ +I ++ YQ++ + S PK++S +Q+
Sbjct: 374 HCLHSKLCIHRNEH*QLWNCEIAQIPFQEQYQEISEEFCPWSGFPKQVSVHQL 216
>TC79086 similar to GP|23499033|emb|CAD51113. ubiquitin-protein ligase 1
putative {Plasmodium falciparum 3D7}, partial (0%)
Length = 1418
Score = 32.7 bits (73), Expect = 0.18
Identities = 24/75 (32%), Positives = 31/75 (41%)
Frame = +2
Query: 226 EDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWW 285
EDEDEEE + D K ++ K K I T + + TK SN
Sbjct: 254 EDEDEEETDTDEDEPPPKTKPIEQTLKSGTKPSIAPARSGTKRPAENNDTKQSN------ 415
Query: 286 KGKKTTSSKKNSKSE 300
KK T+ +KN K E
Sbjct: 416 --KKKTTEEKNKKKE 454
>TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Ateline
herpesvirus 3}, partial (8%)
Length = 986
Score = 32.7 bits (73), Expect = 0.18
Identities = 15/35 (42%), Positives = 20/35 (56%)
Frame = +3
Query: 202 PGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEAD 236
P K S+ K AE ED ++DEDEE+D+ D
Sbjct: 360 PTTKGDSKTQSQDKQHDAEGEDGNDDEDEEDDDGD 464
Score = 28.9 bits (63), Expect = 2.5
Identities = 12/35 (34%), Positives = 22/35 (62%)
Frame = +3
Query: 220 ENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGD 254
E++D+DED+D+EE+ + + G + E E+ D
Sbjct: 630 EDDDEDEDDDDEEEGGEEDEEEGVDEEDNEEEEED 734
Score = 28.9 bits (63), Expect = 2.5
Identities = 15/42 (35%), Positives = 24/42 (56%)
Frame = +3
Query: 193 KSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDE 234
K+ EG G DD ++ ++EDDDEDED++++E
Sbjct: 552 KAPEGGAGGADENGEEEDD---EDGDDQDEDDDEDEDDDDEE 668
>TC79532 weakly similar to GP|8164030|gb|AAF73960.1| SGS3 {Arabidopsis
thaliana}, partial (47%)
Length = 1558
Score = 32.3 bits (72), Expect = 0.23
Identities = 17/80 (21%), Positives = 40/80 (49%)
Frame = +1
Query: 207 YSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDT 266
++R++++ + F +N+ +++D+DEEE+E D ++ D + D+
Sbjct: 607 HARNEIVPEEFEQKNDGEEDDDDEEEEEDDC------------DDLEDTDDDLMSDEYDS 750
Query: 267 STTLKKHATKVSNHIQRWWK 286
+ K H T+ + +W+K
Sbjct: 751 DASQKSHETRKKS---KWFK 801
>TC88470 weakly similar to GP|15146334|gb|AAK83650.1 At1g07080/F10K1_15
{Arabidopsis thaliana}, partial (32%)
Length = 962
Score = 32.0 bits (71), Expect = 0.30
Identities = 17/38 (44%), Positives = 26/38 (67%)
Frame = -1
Query: 226 EDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGI 263
E+ DEEE + SK+G+ +K K+NE+G + K R+GI
Sbjct: 119 EEVDEEEGATNSNSKIGRRMKQKKNERG*IEPK-REGI 9
>TC86045 similar to PIR|T46225|T46225 alpha NAC-like protein - Arabidopsis
thaliana, partial (69%)
Length = 1166
Score = 32.0 bits (71), Expect = 0.30
Identities = 22/97 (22%), Positives = 47/97 (47%)
Frame = +1
Query: 168 VPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDED 227
+ K PG P + + E+E+E+ + S++ DD +++ DDED
Sbjct: 82 ISKTMAPG-PVIDAPATESEVEQQIPSVDETT-LKKKPQPEEDDAPIIEDVKDDDKDDED 255
Query: 228 EDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIV 264
+DE++D+ D K + + +++ ++K RK ++
Sbjct: 256 DDEDDDDEDDDDKEDALAGADGSKQSRSEKKSRKAML 366
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.311 0.129 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,809,966
Number of Sequences: 36976
Number of extensions: 137382
Number of successful extensions: 1980
Number of sequences better than 10.0: 137
Number of HSP's better than 10.0 without gapping: 1081
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1490
length of query: 301
length of database: 9,014,727
effective HSP length: 96
effective length of query: 205
effective length of database: 5,465,031
effective search space: 1120331355
effective search space used: 1120331355
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148360.3


