

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148360.12 + phase: 0 /pseudo
(102 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
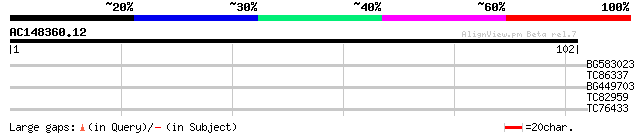
BG583023 similar to PIR|A86220|A86 protein F22O13.29 [imported] ... 29 0.33
TC86337 MtN1 27 1.6
BG449703 weakly similar to PIR|T45588|T455 arm repeat containing... 27 1.6
TC82959 similar to GP|12321948|gb|AAG51005.1 unknown protein; 13... 25 3.7
TC76433 weakly similar to GP|15810483|gb|AAL07129.1 unknown prot... 25 6.3
>BG583023 similar to PIR|A86220|A86 protein F22O13.29 [imported] -
Arabidopsis thaliana, partial (6%)
Length = 827
Score = 28.9 bits (63), Expect = 0.33
Identities = 24/83 (28%), Positives = 35/83 (41%), Gaps = 2/83 (2%)
Frame = -2
Query: 7 PPFDVIWVYSAIKFGCRK--SLKGDILEQISAAKALFQLISACSASVGGSKSIALLAPKA 64
P DVI + + F R L+ D+ S K+ F ++ SV I +
Sbjct: 292 PVSDVISLLPDLMFSSRSMAGLRSDVSSSSSVLKSYFSFDTSTGTSV-----IFDNIGTS 128
Query: 65 MKEVKSLVDMILGFMSICCSKIS 87
+KE+ SL D +SICC S
Sbjct: 127 LKEIGSLGDGQTSLLSICCQSSS 59
>TC86337 MtN1
Length = 925
Score = 26.6 bits (57), Expect = 1.6
Identities = 13/27 (48%), Positives = 18/27 (66%)
Frame = +2
Query: 8 PFDVIWVYSAIKFGCRKSLKGDILEQI 34
PFD+IWV S FG K+L +LE++
Sbjct: 611 PFDLIWVKSDNSFGTYKNL---VLEKV 682
>BG449703 weakly similar to PIR|T45588|T455 arm repeat containing protein
homolog - Arabidopsis thaliana, partial (6%)
Length = 671
Score = 26.6 bits (57), Expect = 1.6
Identities = 12/31 (38%), Positives = 20/31 (63%)
Frame = +3
Query: 57 IALLAPKAMKEVKSLVDMILGFMSICCSKIS 87
++L PK++ L+ ++L FMSICC I+
Sbjct: 468 LSLEGPKSI-----LIHLVLSFMSICCLSIT 545
>TC82959 similar to GP|12321948|gb|AAG51005.1 unknown protein; 13339-10119
{Arabidopsis thaliana}, partial (9%)
Length = 710
Score = 25.4 bits (54), Expect = 3.7
Identities = 12/31 (38%), Positives = 16/31 (50%)
Frame = -1
Query: 18 IKFGCRKSLKGDILEQISAAKALFQLISACS 48
+KF CR S D E +S+ + F ACS
Sbjct: 548 VKFKCR*SHSQDTKESLSSPRKTFNSFFACS 456
>TC76433 weakly similar to GP|15810483|gb|AAL07129.1 unknown protein
{Arabidopsis thaliana}, partial (61%)
Length = 2810
Score = 24.6 bits (52), Expect = 6.3
Identities = 5/18 (27%), Positives = 14/18 (77%)
Frame = +2
Query: 8 PFDVIWVYSAIKFGCRKS 25
P+ ++W++S + F C+++
Sbjct: 1265 PYKMVWMFSTVLFNCQRT 1318
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.135 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,988,257
Number of Sequences: 36976
Number of extensions: 30223
Number of successful extensions: 153
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 153
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 153
length of query: 102
length of database: 9,014,727
effective HSP length: 78
effective length of query: 24
effective length of database: 6,130,599
effective search space: 147134376
effective search space used: 147134376
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 50 (23.9 bits)
Medicago: description of AC148360.12


