

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148348.6 + phase: 0
(552 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
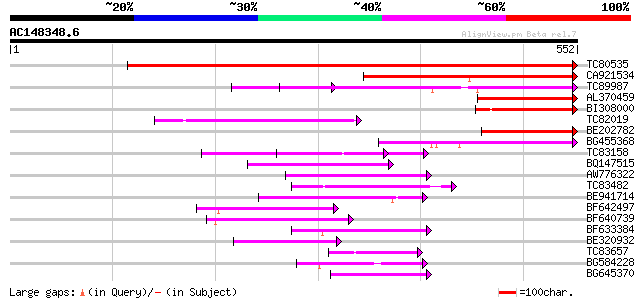
Score E
Sequences producing significant alignments: (bits) Value
TC80535 weakly similar to GP|14335136|gb|AAK59848.1 At2g15690/F9... 908 0.0
CA921534 similar to PIR|D96516|D96 F16N3.14 [imported] - Arabido... 288 3e-78
TC89987 weakly similar to PIR|D86443|D86443 probable PPR-repeat ... 186 3e-47
AL370459 weakly similar to GP|20146256|d hypothetical protein {O... 143 2e-34
BI308000 similar to GP|7363286|dbj| Similar to Arabidopsis thali... 136 2e-32
TC82019 weakly similar to GP|11994382|dbj|BAB02341. gb|AAB82628.... 129 2e-30
BE202782 weakly similar to GP|22093801|dbj hypothetical protein~... 125 6e-29
BG455368 weakly similar to GP|9759524|dbj selenium-binding prote... 120 2e-27
TC83158 weakly similar to PIR|H86191|H86191 hypothetical protein... 113 2e-25
BQ147515 weakly similar to PIR|T05355|T05 hypothetical protein F... 94 2e-19
AW776322 weakly similar to PIR|H84859|H84 hypothetical protein A... 90 3e-18
TC83482 weakly similar to GP|9758453|dbj|BAB08982.1 selenium-bin... 88 1e-17
BE941714 weakly similar to GP|11994734|dbj selenium-binding prot... 80 2e-15
BF642497 weakly similar to PIR|H96713|H96 hypothetical protein T... 78 1e-14
BF640739 weakly similar to GP|7363286|dbj Similar to Arabidopsis... 76 4e-14
BF633384 similar to PIR|T46179|T46 hypothetical protein T8H10.30... 73 3e-13
BE320932 weakly similar to PIR|T04548|T045 hypothetical protein ... 69 6e-12
TC83657 weakly similar to GP|11994382|dbj|BAB02341. gb|AAB82628.... 67 1e-11
BG584228 weakly similar to PIR|F71411|F71 hypothetical protein -... 67 2e-11
BG645370 weakly similar to PIR|T00405|T004 hypothetical protein ... 66 4e-11
>TC80535 weakly similar to GP|14335136|gb|AAK59848.1 At2g15690/F9O13.24
{Arabidopsis thaliana}, partial (55%)
Length = 1727
Score = 908 bits (2347), Expect = 0.0
Identities = 438/438 (100%), Positives = 438/438 (100%)
Frame = +2
Query: 115 PPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQA 174
PPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQA
Sbjct: 5 PPPPQNPNFRQPIRQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQA 184
Query: 175 PPPPSIVDLTRFCQEGKVKEALELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFL 234
PPPPSIVDLTRFCQEGKVKEALELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFL
Sbjct: 185 PPPPSIVDLTRFCQEGKVKEALELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFL 364
Query: 235 QSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQ 294
QSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQ
Sbjct: 365 QSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQ 544
Query: 295 LFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVL 354
LFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVL
Sbjct: 545 LFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVL 724
Query: 355 GQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKAVANK 414
GQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKAVANK
Sbjct: 725 GQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKAVANK 904
Query: 415 IPTPPPKKYTAISMLDGKNRIIEYKNPTLYKDDEKLIAMNSMKDAGYVPDTRYVLHDIDQ 474
IPTPPPKKYTAISMLDGKNRIIEYKNPTLYKDDEKLIAMNSMKDAGYVPDTRYVLHDIDQ
Sbjct: 905 IPTPPPKKYTAISMLDGKNRIIEYKNPTLYKDDEKLIAMNSMKDAGYVPDTRYVLHDIDQ 1084
Query: 475 EAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRD 534
EAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRD
Sbjct: 1085EAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRD 1264
Query: 535 NKRFHHFKDGKCSCGDYW 552
NKRFHHFKDGKCSCGDYW
Sbjct: 1265NKRFHHFKDGKCSCGDYW 1318
>CA921534 similar to PIR|D96516|D96 F16N3.14 [imported] - Arabidopsis
thaliana, partial (18%)
Length = 790
Score = 288 bits (738), Expect = 3e-78
Identities = 140/211 (66%), Positives = 172/211 (81%), Gaps = 3/211 (1%)
Frame = -1
Query: 345 EHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVS 404
E Y+G++++ G + L +A EF+ +P EP V ++ETL+N+ARIHGD++ ED +EL+
Sbjct: 784 EPYLGVVNIFGCAVGLMKAHEFMRNMPIEPGVELWETLRNFARIHGDLEREDCADELLTV 605
Query: 405 LDPSKAVANKIPTPPPKKYTAISMLDGKNRIIEYKNPTLYKD--DEKLIAMNS-MKDAGY 461
LDPSKA A+K+P P KK +AI+ML+ KNR+ EY+ YK+ D KL + M++AGY
Sbjct: 604 LDPSKAAADKVPLPQRKKQSAINMLEEKNRVSEYRCNMPYKEEGDVKLRGLTGQMREAGY 425
Query: 462 VPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKI 521
VPDTRYVLHDID+E KE+AL YHSERLAIAYGLISTPPRT LRIIKNLR+CGDCHNAIKI
Sbjct: 424 VPDTRYVLHDIDEEEKEKALQYHSERLAIAYGLISTPPRTTLRIIKNLRICGDCHNAIKI 245
Query: 522 MSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
MS+IVGRELIVRDNKRFHHFKDGKCSCGDYW
Sbjct: 244 MSKIVGRELIVRDNKRFHHFKDGKCSCGDYW 152
>TC89987 weakly similar to PIR|D86443|D86443 probable PPR-repeat protein
[imported] - Arabidopsis thaliana, partial (62%)
Length = 1559
Score = 186 bits (471), Expect = 3e-47
Identities = 107/322 (33%), Positives = 169/322 (52%), Gaps = 32/322 (9%)
Frame = +2
Query: 263 RVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAE 322
RVF +M +N S+ +MI G A G E L++F +M E GL + V SAC A
Sbjct: 578 RVFKNMSEKNRYSYTVMISGLAIHGRGKEALKVFSEMIEEGLAPDDVVYVGVFSACSHAG 757
Query: 323 AVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETL 382
VE+ +SM+ ++ IEP V+HY ++D+LG+ G LKEA E I+ + +P ++ +L
Sbjct: 758 LVEEGLQCFKSMQFEHKIEPTVQHYGCMVDLLGRFGMLKEAYELIKSMSIKPNDVIWRSL 937
Query: 383 KNYARIHGDVDLEDHVEELIVSLDPSKA---------------------VANKIPTPPPK 421
+ ++H ++++ E + L+ + + + K+
Sbjct: 938 LSACKVHHNLEIGKIAAENLFMLNQNNSGDYLVLANMYAKAQKWDDVAKIRTKLAERNLV 1117
Query: 422 KYTAISMLDGKNRIIEYKNPTLYKDDEKLIAMN-----------SMKDAGYVPDTRYVLH 470
+ S+++ K ++ ++ + D+ + N +K GY+PDT VL
Sbjct: 1118QTPGFSLIEAKRKVYKFVS-----QDKSIPQWNIIYEMIHQMEWQVKFEGYIPDTSQVLL 1282
Query: 471 DIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGREL 530
D+D E K++ L +HS++LAIA+GLI T PLRI +NLR+C DCH K +S I RE+
Sbjct: 1283DVDDEEKKERLKFHSQKLAIAFGLIHTSEGXPLRITRNLRMCSDCHTYTKYISMIYEREI 1462
Query: 531 IVRDNKRFHHFKDGKCSCGDYW 552
VRD RF HFK+G CSC DYW
Sbjct: 1463TVRDRXRFXHFKNGSCSCKDYW 1528
Score = 54.3 bits (129), Expect = 1e-07
Identities = 31/103 (30%), Positives = 52/103 (50%), Gaps = 1/103 (0%)
Frame = +1
Query: 217 CGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSW 276
C V++ +VH + + D + N +I MYG C + +A VF+ M +++ SW
Sbjct: 133 CSLLGVVDEGIQVHGHVFKMGLEGDVIVQNSLINMYGKCGEIKNACDVFNGMDEKSVASW 312
Query: 277 HMMIRGYANSTMGDEGLQLFEQMNELG-LEITSETMLAVLSAC 318
+I +A M +E L L +M+ G + T++ VLSAC
Sbjct: 313 SAIIGAHACVEMWNECLMLLGKMSSEGRCRVEESTLVNVLSAC 441
>AL370459 weakly similar to GP|20146256|d hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (11%)
Length = 477
Score = 143 bits (360), Expect = 2e-34
Identities = 61/97 (62%), Positives = 76/97 (77%)
Frame = +1
Query: 456 MKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDC 515
+K+ GYVP+T +VLHD++ E KEQ L HSE+LA+A+ LISTP P+R+ KNLRVCGDC
Sbjct: 91 IKNVGYVPNTDFVLHDVEDEQKEQYLFQHSEKLAVAFALISTPNPKPIRVFKNLRVCGDC 270
Query: 516 HNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
H AIK +S + GRE++VRD RFHH KDG CSC DYW
Sbjct: 271 HTAIKYISMVSGREIVVRDANRFHHMKDGTCSCNDYW 381
>BI308000 similar to GP|7363286|dbj| Similar to Arabidopsis thaliana DNA
chromosome 4 BAC clone F4I10; vegetative storage
protein., partial (12%)
Length = 771
Score = 136 bits (342), Expect = 2e-32
Identities = 59/99 (59%), Positives = 79/99 (79%)
Frame = +3
Query: 454 NSMKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCG 513
+ ++DAGY+PDT + HD++++ KEQ L HSERLAIA+GL++T TP+ + KNLRVCG
Sbjct: 42 DKIRDAGYIPDTNSI-HDVEEKVKEQLLSSHSERLAIAFGLLNTNHGTPIHVRKNLRVCG 218
Query: 514 DCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
DCH+ K +S + GRE+IVRD +RFHHFK+G CSCGDYW
Sbjct: 219 DCHDVTKYISLVTGREIIVRDLRRFHHFKNGICSCGDYW 335
>TC82019 weakly similar to GP|11994382|dbj|BAB02341.
gb|AAB82628.1~gene_id:MIL23.2~similar to unknown protein
{Arabidopsis thaliana}, partial (7%)
Length = 737
Score = 129 bits (325), Expect = 2e-30
Identities = 75/201 (37%), Positives = 117/201 (57%)
Frame = +1
Query: 142 GINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPPSIVDLTRFCQEGKVKEALELMEK 201
G N +++ P+ + NP+ Q + + QAP V+ F QEG V + LELM +
Sbjct: 142 GTNNSRFTPRKTQPLRMGNPSIQPKLNH--HQAPHQHKNVNFAHFLQEGNVNQVLELMGQ 315
Query: 202 GIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDA 261
G AD + F L LC KS+E K+VH++ +S F + ++ N++I +Y C S+ DA
Sbjct: 316 GAFADYSDFLSLLKLCEDLKSLELGKRVHEFLRRSKFGGNVELCNRLIGLYVKCGSVKDA 495
Query: 262 RRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSA 321
R+VFD MP+RN+ S ++MI GY + + +GL +F+QM + G+ ET VL+ C
Sbjct: 496 RKVFDKMPDRNVGSLNLMIGGYNVNGLRIDGLLVFKQMRQQGVVPDEETFALVLAVCALV 675
Query: 322 EAVEDAYIYLESMKSKYGIEP 342
+ VE++ + ESMK +YGI P
Sbjct: 676 DGVEESLMQFESMK-EYGIVP 735
>BE202782 weakly similar to GP|22093801|dbj hypothetical protein~predicted by
GlimmerM~similar to Arabidopsis thaliana chromosome3
At3g23330, partial (13%)
Length = 548
Score = 125 bits (313), Expect = 6e-29
Identities = 53/93 (56%), Positives = 71/93 (75%)
Frame = +2
Query: 460 GYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAI 519
G +P T+ VL D++++ KEQ L HSE+LA+ GLI+T P PL++IKNLR+C DCH I
Sbjct: 2 GCLPMTKSVLQDVEEQDKEQILCGHSEKLAVVLGLINTSPGQPLQVIKNLRICDDCHAVI 181
Query: 520 KIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
K++SR+ GRE+ VRD RFHHFK+G CSC D+W
Sbjct: 182 KVISRLEGREIFVRDTNRFHHFKEGVCSCADFW 280
>BG455368 weakly similar to GP|9759524|dbj selenium-binding protein-like
{Arabidopsis thaliana}, partial (33%)
Length = 687
Score = 120 bits (300), Expect = 2e-27
Identities = 72/221 (32%), Positives = 115/221 (51%), Gaps = 28/221 (12%)
Frame = +1
Query: 360 LKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSK---------- 409
+++A + ++ +P EP V+ L + IHG+ D+ + + L+P
Sbjct: 7 VEKALQLVQTMPMEPNGGVWGALLGASHIHGNPDVAEIASRSLFELEPDNLGNYLLLSKT 186
Query: 410 -AVANK----------IPTPPPKKYTAISMLDGKNRII------EYKNPTLYKDDEKLI- 451
A+A K + +K S ++ KN II + K+P + + + L
Sbjct: 187 YALAAKWDDVSRVRKLMREKQLRKNPGCSWVEAKNGIIHEFFAGDVKHPEINEIKKALDD 366
Query: 452 AMNSMKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRV 511
+ +K GY P V +DID E K L+ HSE+LA+AYGL+ST + ++I+KNLR+
Sbjct: 367 LLQRLKCTGYQPKLNSVPYDIDDEGKRCLLVSHSEKLALAYGLLSTDAGSTIKIMKNLRI 546
Query: 512 CGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
C DCH + S + GR++IVRDN FHHF +G CSC ++W
Sbjct: 547 CEDCHIVMCGASXLTGRKIIVRDNMXFHHFLNGACSCNNFW 669
>TC83158 weakly similar to PIR|H86191|H86191 hypothetical protein [imported]
- Arabidopsis thaliana, partial (42%)
Length = 1194
Score = 113 bits (283), Expect = 2e-25
Identities = 66/221 (29%), Positives = 110/221 (48%)
Frame = +2
Query: 187 CQEGKVKEALELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHN 246
C E ++ E+ G+ D + C ++ VH ++ FR + K+ N
Sbjct: 98 CYEEALECFREMQLAGVVPDFVTVIAIISACANLGALGLGLWVHRLVMKKEFRDNVKVLN 277
Query: 247 KVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEI 306
+I+MY C + AR+VFD M RN+ SW+ +I G+A + + D+ L F M + GLE
Sbjct: 278 SLIDMYARCGCIELARQVFDGMSQRNLVSWNSIIVGFAVNGLADKALSFFRSMKKEGLEP 457
Query: 307 TSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEF 366
+ + L+AC A +++ +K + P +EHY L+D+ ++G LKEA +
Sbjct: 458 NGVSYTSALTACSHAGLIDEGLKIFADIKRDHRNSPRIEHYGCLVDLYSRAGRLKEAWDV 637
Query: 367 IEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDP 407
I+++P P V +L R GDV+L + V + V L P
Sbjct: 638 IKKMPMMPNEVVLGSLLAACRTQGDVELAEKVMKYQVELYP 760
Score = 50.1 bits (118), Expect = 2e-06
Identities = 27/109 (24%), Positives = 57/109 (51%)
Frame = +2
Query: 260 DARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACG 319
DA ++FD +P +N+ SW ++I G+ +E L+ F +M G+ T++A++SAC
Sbjct: 14 DALKLFDKLPVKNVVSWTVVIGGFVKKECYEEALECFREMQLAGVVPDFVTVIAIISACA 193
Query: 320 SAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIE 368
+ A+ +++ + K V+ L+D+ + G ++ A + +
Sbjct: 194 NLGAL-GLGLWVHRLVMKKEFRDNVKVLNSLIDMYARCGCIELARQVFD 337
>BQ147515 weakly similar to PIR|T05355|T05 hypothetical protein F8B4.150 -
Arabidopsis thaliana, partial (23%)
Length = 427
Score = 93.6 bits (231), Expect = 2e-19
Identities = 48/142 (33%), Positives = 76/142 (52%)
Frame = +2
Query: 232 YFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDE 291
Y LQ N ++EMY C S+ DA VF +M R++ + +MI+ A + ++
Sbjct: 2 YVLQHLSPLKVSTCNGILEMYFQCGSVDDAVNVFKNMTERDLTTICIMIKQLAKNGFAED 181
Query: 292 GLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLL 351
+ LF Q GL+ + + V AC + + ++ ESM Y I P +EHY+ L+
Sbjct: 182 SIDLFTQFKRSGLKPDGQMFIGVFGACXMLGDIVEGMLHFESMSRDYDIVPTMEHYVSLV 361
Query: 352 DVLGQSGYLKEAEEFIEQLPFE 373
D++G G+L EA EFIE++P E
Sbjct: 362 DMIGSIGHLDEALEFIEKMPME 427
>AW776322 weakly similar to PIR|H84859|H84 hypothetical protein At2g42920
[imported] - Arabidopsis thaliana, partial (27%)
Length = 633
Score = 89.7 bits (221), Expect = 3e-18
Identities = 48/143 (33%), Positives = 79/143 (54%), Gaps = 1/143 (0%)
Frame = +1
Query: 269 PNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGL-EITSETMLAVLSACGSAEAVEDA 327
P R + W+ +I G A + E + F ++ L + S + + VL+AC A+ A
Sbjct: 10 PRRGLSCWNSIIIGLAMNGHEREAFEFFSKLESSKLLKPDSVSFIGVLTACKHLGAINKA 189
Query: 328 YIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYAR 387
Y E M +KY IEP ++HY ++DVLGQ+G L+EAEE I+ +P +P ++ +L + R
Sbjct: 190 RDYFELMMNKYEIEPSIKHYTCIVDVLGQAGLLEEAEELIKGMPLKPDAIIWGSLLSSCR 369
Query: 388 IHGDVDLEDHVEELIVSLDPSKA 410
H +V + + + L+PS A
Sbjct: 370 KHRNVQIARRAAQRVYELNPSDA 438
>TC83482 weakly similar to GP|9758453|dbj|BAB08982.1 selenium-binding
protein-like {Arabidopsis thaliana}, partial (10%)
Length = 895
Score = 87.8 bits (216), Expect = 1e-17
Identities = 50/161 (31%), Positives = 85/161 (52%)
Frame = +2
Query: 275 SWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESM 334
SW+ +I GYA+ + L+ F++M +G T + VLSAC A VE+ + M
Sbjct: 11 SWNAIIGGYASHGLATRALEEFDRMKVVGTP-DEVTFVNVLSACVHAGLVEEGEKHFTDM 187
Query: 335 KSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDL 394
+KYGI+ +EHY ++D+ G++G EAE I+ +PFEP V ++ L +H +++L
Sbjct: 188 LTKYGIQAEMEHYSCMVDLYGRAGRFDEAENLIKNMPFEPDVVLWGALLAACGLHSNLEL 367
Query: 395 EDHVEELIVSLDPSKAVANKIPTPPPKKYTAISMLDGKNRI 435
++ E I L+ S P Y+ +S + G+ +
Sbjct: 368 GEYAAERIRRLESSH----------PVSYSVLSKIQGEKGV 460
>BE941714 weakly similar to GP|11994734|dbj selenium-binding protein-like
{Arabidopsis thaliana}, partial (10%)
Length = 619
Score = 80.5 bits (197), Expect = 2e-15
Identities = 49/169 (28%), Positives = 86/169 (49%), Gaps = 5/169 (2%)
Frame = +2
Query: 243 KMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNEL 302
++ +++MY C ++ A+R+FD M R++ W+ MI G A G L+LF M ++
Sbjct: 17 RLSTSLLDMYAKCGNLELAKRLFDSMNMRDVVCWNAMISGMAMHGDGKGALKLFYDMEKV 196
Query: 303 GLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKE 362
G++ T +AV +AC + + + L+ M S Y I P EHY L+D+L ++G +E
Sbjct: 197 GVKPDDITFIAVFTACSYSGMAYEGLMLLDKMCSVYNIVPKSEHYGCLVDLLSRAGLFEE 376
Query: 363 AEEFIEQLP-----FEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLD 406
A I ++ E T+ + + HG+ L + E ++ LD
Sbjct: 377 AMVMIRKITNSWNGSEETL-AWRAFLSACCNHGETQLAELAAEKVLQLD 520
>BF642497 weakly similar to PIR|H96713|H96 hypothetical protein T6L1.11
[imported] - Arabidopsis thaliana, partial (20%)
Length = 461
Score = 77.8 bits (190), Expect = 1e-14
Identities = 36/142 (25%), Positives = 78/142 (54%), Gaps = 4/142 (2%)
Frame = +3
Query: 183 LTRFCQEGKVKEALELMEK----GIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTF 238
+T F Q G ++A+++ + ++ D F + CG ++++ K+VH Y +++ +
Sbjct: 24 ITGFTQNGLDRDAIDIFREMKLENLQMDQYTFGSVLTACGGVMALQEGKQVHAYIIRTDY 203
Query: 239 RSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQ 298
+ + + + +++MY CK++ A VF M +N+ SW M+ GY + +E ++ F
Sbjct: 204 KDNIFVASALVDMYCKCKNIKSAEAVFKKMTCKNVVSWTAMLVGYGQNGYSEEAVKTFSD 383
Query: 299 MNELGLEITSETMLAVLSACGS 320
M + G+E T+ +V+S+C +
Sbjct: 384 MQKYGIEPDDFTLGSVISSCAN 449
>BF640739 weakly similar to GP|7363286|dbj Similar to Arabidopsis thaliana
DNA chromosome 4 BAC clone F4I10; vegetative storage
protein., partial (14%)
Length = 444
Score = 75.9 bits (185), Expect = 4e-14
Identities = 45/148 (30%), Positives = 73/148 (48%), Gaps = 5/148 (3%)
Frame = +1
Query: 192 VKEALELM----EKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNK 247
V EAL L +GIK D+ F + AK +H +++ ++ +
Sbjct: 1 VNEALNLFCTMQSQGIKPDSFTFVSVITALADLSVTRQAKWIHGLAIRTNMDTNVFVATA 180
Query: 248 VIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQM-NELGLEI 306
+++MY C ++ AR +FD M R++ +W+ MI GY +G L LF+ M NE L+
Sbjct: 181 LVDMYAKCGAIETARELFDMMQERHVITWNAMIDGYGTHGLGKAALDLFDDMQNEASLKP 360
Query: 307 TSETMLAVLSACGSAEAVEDAYIYLESM 334
T L+V+SAC + VE+ Y + M
Sbjct: 361 NDITFLSVISACSHSGFVEEGLYYFKIM 444
>BF633384 similar to PIR|T46179|T46 hypothetical protein T8H10.30 -
Arabidopsis thaliana, partial (22%)
Length = 595
Score = 72.8 bits (177), Expect = 3e-13
Identities = 41/142 (28%), Positives = 78/142 (54%), Gaps = 6/142 (4%)
Frame = +1
Query: 275 SWHMMIRGYANSTMGDEGLQLFEQMNELG-----LEITSETMLAVLSACGSAEAVEDAYI 329
+W+++I Y G+E L+LF +M E G + T +A+ ++ + V++
Sbjct: 4 TWNVLIMAYGMHGKGEEALKLFRRMVEEGDNNREIRPNEVTYIAIFASLSHSGMVDEGLN 183
Query: 330 YLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFE-PTVTVFETLKNYARI 388
+MK+K+GIEP +HY L+D+LG+SG ++EA I+ +P V + +L +I
Sbjct: 184 LFYTMKAKHGIEPTSDHYACLVDLLGRSGQIEEAYNLIKTMPSNMKKVDAWSSLLGACKI 363
Query: 389 HGDVDLEDHVEELIVSLDPSKA 410
H ++++ + + + LDP+ A
Sbjct: 364 HQNLEIGEIAAKNLFVLDPNVA 429
>BE320932 weakly similar to PIR|T04548|T045 hypothetical protein F28J12.180 -
Arabidopsis thaliana, partial (14%)
Length = 354
Score = 68.6 bits (166), Expect = 6e-12
Identities = 30/105 (28%), Positives = 58/105 (54%)
Frame = +3
Query: 219 KSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHM 278
++K+ + ++H ++ +SD + +I+MY C + +++VFD M RN +W
Sbjct: 12 ENKAFKCGTQLHGAIVKKICKSDVFIGTSLIDMYAKCGEIVSSKKVFDRMKVRNTATWTS 191
Query: 279 MIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEA 323
+I GYA + G+E L F M + + T++ V++ACG+ +A
Sbjct: 192 IISGYARNGFGEEALNFFRLMKRKKVYVNKSTLVCVMTACGTIKA 326
>TC83657 weakly similar to GP|11994382|dbj|BAB02341.
gb|AAB82628.1~gene_id:MIL23.2~similar to unknown protein
{Arabidopsis thaliana}, partial (10%)
Length = 1299
Score = 67.4 bits (163), Expect = 1e-11
Identities = 34/92 (36%), Positives = 56/92 (59%)
Frame = +3
Query: 311 MLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQL 370
+L VLSAC + +A + M+ +YGIE G+ HY ++D+LG++G LKEA E I+++
Sbjct: 75 LLXVLSACAHGGLMSEALEVISKME-EYGIEMGIRHYGCMVDLLGRAGKLKEAYELIKRM 251
Query: 371 PFEPTVTVFETLKNYARIHGDVDLEDHVEELI 402
P +P TV + IH D+ + + V ++I
Sbjct: 252 PMKPNETVLGAMIGACWIHSDMKMAEQVMKMI 347
>BG584228 weakly similar to PIR|F71411|F71 hypothetical protein - Arabidopsis
thaliana, partial (19%)
Length = 827
Score = 66.6 bits (161), Expect = 2e-11
Identities = 41/129 (31%), Positives = 65/129 (49%), Gaps = 2/129 (1%)
Frame = -3
Query: 280 IRGYANSTMGDEGLQLFEQMN--ELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSK 337
+RGYA+ L+ FE+M G+ ++ T++++LSACG A V ESM+
Sbjct: 684 VRGYAHQGNVYMALRQFEEMTFGSRGIRLSYVTLVSILSACGRAGVVLRGMQIFESMRLN 505
Query: 338 YGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDH 397
Y IEPG HY ++ +LG FI+ +P +PT++ + L R+HG +
Sbjct: 504 YAIEPGTGHYACVVGLLG------SVC*FIQNMPIQPTISEWGVLLGAFRMHGKTKMGKI 343
Query: 398 VEELIVSLD 406
E + LD
Sbjct: 342 TAEKLFELD 316
Score = 28.9 bits (63), Expect = 5.4
Identities = 11/14 (78%), Positives = 11/14 (78%)
Frame = -1
Query: 534 DNKRFHHFKDGKCS 547
DN RFH FKDG CS
Sbjct: 56 DNHRFHRFKDGCCS 15
>BG645370 weakly similar to PIR|T00405|T004 hypothetical protein At2g44880
[imported] - Arabidopsis thaliana, partial (11%)
Length = 784
Score = 65.9 bits (159), Expect = 4e-11
Identities = 27/98 (27%), Positives = 57/98 (57%)
Frame = -2
Query: 313 AVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPF 372
AVL+AC A ++ Y M + YG+ P ++ Y L+D+ ++G L++A + +E++P+
Sbjct: 777 AVLTACNHAGFIDKGEEYFNKMITNYGLSPDIDIYACLIDLYARNGNLRKARDLMEEMPY 598
Query: 373 EPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKA 410
+P ++ + + +I+GDV+L ++ ++P A
Sbjct: 597 DPNCIIWSSFLSACKIYGDVELGREAAIQLIKMEPCNA 484
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,007,613
Number of Sequences: 36976
Number of extensions: 525003
Number of successful extensions: 18887
Number of sequences better than 10.0: 813
Number of HSP's better than 10.0 without gapping: 4735
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 9372
length of query: 552
length of database: 9,014,727
effective HSP length: 101
effective length of query: 451
effective length of database: 5,280,151
effective search space: 2381348101
effective search space used: 2381348101
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148348.6


