

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148347.2 + phase: 0 /pseudo
(813 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
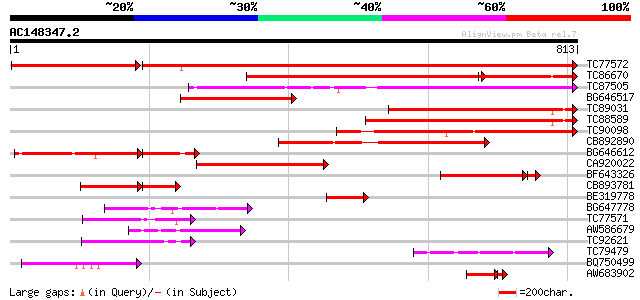
Score E
Sequences producing significant alignments: (bits) Value
TC77572 similar to GP|9558523|dbj|BAB03441.1 ESTs AU078742(C1188... 1211 0.0
TC86670 weakly similar to GP|9758141|dbj|BAB08633.1 disease resi... 459 e-129
TC87505 weakly similar to PIR|T05981|T05981 hypothetical protein... 315 4e-86
BG646517 301 7e-82
TC89031 weakly similar to GP|9758141|dbj|BAB08633.1 disease resi... 300 1e-81
TC88589 weakly similar to GP|9758140|dbj|BAB08632.1 disease resi... 300 1e-81
TC90098 weakly similar to GP|9758140|dbj|BAB08632.1 disease resi... 291 9e-79
CB892890 weakly similar to GP|9758141|dbj| disease resistance pr... 287 1e-77
BG646612 weakly similar to GP|9758140|dbj| disease resistance pr... 280 2e-75
CA920022 similar to PIR|T03031|T030 NBS-LRR type resistance prot... 263 2e-70
BF643326 weakly similar to GP|11994216|db disease resistance com... 200 2e-55
CB893781 homologue to PIR|C64504|C645 hypothetical protein MJ163... 175 5e-44
BE319778 124 2e-28
BG647778 weakly similar to PIR|T48468|T484 disease resistance-li... 122 7e-28
TC77571 homologue to GP|20804846|dbj|BAB92528. putative rust res... 117 1e-26
AW586679 similar to GP|9758140|dbj| disease resistance protein-l... 105 5e-23
TC92621 similar to GP|7107252|gb|AAF36340.1| unknown {Pennisetum... 103 2e-22
TC79479 similar to GP|12056928|gb|AAG48132.1 putative resistance... 86 7e-17
BQ750499 similar to OMNI|NT01MC4764 transport protein HasD puta... 85 1e-16
AW683902 weakly similar to PIR|T48468|T484 disease resistance-li... 68 3e-13
>TC77572 similar to GP|9558523|dbj|BAB03441.1 ESTs AU078742(C11888)
C26225(C11888) correspond to a region of the predicted
gene.~Similar to Oryza, partial (5%)
Length = 2535
Score = 1211 bits (3132), Expect = 0.0
Identities = 618/668 (92%), Positives = 620/668 (92%), Gaps = 45/668 (6%)
Frame = +3
Query: 191 A*KSKIHCWVGYTI*QVENGASEGW*LHTCVDWFRRIGENHSGYKALLG*RS*W------ 244
+*KSKIHCWVGYTI*QVENGASEGW*LHTCVDWFRRIGENHSGYKALLG*+
Sbjct: 447 S*KSKIHCWVGYTI*QVENGASEGW*LHTCVDWFRRIGENHSGYKALLG*KKLMVNSRKT 626
Query: 245 ---------------------------------------LGLLLKKVEGSPLLLVLDDVW 265
LGLLLKKVEGSPLLLVLDDVW
Sbjct: 627 LYLSPSQKHPC*RLFVERIHQHCGYPVPEFQSDEDAVNRLGLLLKKVEGSPLLLVLDDVW 806
Query: 266 PISEPLVEKIQFQISDFKILVTSRVAFPRFSTTCILKPLAHEEAVTLFHHYAQMEKNSSD 325
PISEPLVEKIQFQISDFKILVTSRVAFPRFSTTCILKPLAHEEAVTLFHHYAQMEKNSSD
Sbjct: 807 PISEPLVEKIQFQISDFKILVTSRVAFPRFSTTCILKPLAHEEAVTLFHHYAQMEKNSSD 986
Query: 326 IINKNLVEKVVRSCQGLPLTIKVIATSLKNRPRDMWRKIGKELSQGHSILDSNTDLLTRL 385
IINKNLVEKVVRSCQGLPLTIKVIATSLKNRPRDMWRKIGKELSQGHSILDSNTDLLTRL
Sbjct: 987 IINKNLVEKVVRSCQGLPLTIKVIATSLKNRPRDMWRKIGKELSQGHSILDSNTDLLTRL 1166
Query: 386 QKIFDVLEDNPTIMECFMDIALFPEDHRIPVAALVDMWAELYRLDDNGIQAMEIINKLGI 445
QKIFDVLEDNPTIMECFMDIALFPEDHRIPVAALVDMWAELYRLDDNGIQAMEIINKLGI
Sbjct: 1167 QKIFDVLEDNPTIMECFMDIALFPEDHRIPVAALVDMWAELYRLDDNGIQAMEIINKLGI 1346
Query: 446 MNLANVIIPRKNASDTDNNNYNNHFIILHDILRELGIYQSAKEPFEQRKRLIIDINKNKS 505
MNLANVIIPRKNASDTDNNNYNNHFIILHDILRELGIYQSAKEPFEQRKRLIIDINKNKS
Sbjct: 1347 MNLANVIIPRKNASDTDNNNYNNHFIILHDILRELGIYQSAKEPFEQRKRLIIDINKNKS 1526
Query: 506 GLAEKQQGLMTCILSKFMRLCVKRNPQQLTARILSVSADETCAFDWSQMEPAQVEVLILN 565
GLAEKQQGLMTCILSKFMRLCVKRNPQQLTARILSVSADETCAFDWSQMEPAQVEVLILN
Sbjct: 1527 GLAEKQQGLMTCILSKFMRLCVKRNPQQLTARILSVSADETCAFDWSQMEPAQVEVLILN 1706
Query: 566 LHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLERIYVPSFGT 625
LHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLERIYVPSFGT
Sbjct: 1707 LHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLERIYVPSFGT 1886
Query: 626 LKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLR 685
LKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLR
Sbjct: 1887 LKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLR 2066
Query: 686 VTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPE 745
VTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPE
Sbjct: 2067 VTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPE 2246
Query: 746 EFGNLCNLKNLDMASCASIELPFSVVNLQNLKTITCDEETAATWEDFQHMLPNMKIEVLH 805
EFGNLCNLKNLDMASCASIELPFSVVNLQNLKTITCDEETAATWEDFQHMLPNMKIEVLH
Sbjct: 2247 EFGNLCNLKNLDMASCASIELPFSVVNLQNLKTITCDEETAATWEDFQHMLPNMKIEVLH 2426
Query: 806 VDVNLNWL 813
VDVNLNWL
Sbjct: 2427 VDVNLNWL 2450
Score = 369 bits (946), Expect = e-102
Identities = 184/185 (99%), Positives = 185/185 (99%)
Frame = +1
Query: 3 SATHIITLTTRFQFNDVLNKISQILQKAQKNQSSRRSLRLILKDLSPVVQDIKQYNDHLD 62
+ATHIITLTTRFQFNDVLNKISQILQKAQKNQSSRRSLRLILKDLSPVVQDIKQYNDHLD
Sbjct: 43 AATHIITLTTRFQFNDVLNKISQILQKAQKNQSSRRSLRLILKDLSPVVQDIKQYNDHLD 222
Query: 63 HPREEINSLIEENDAGESACTCSSENDSYGYVVEENQSLTVNDVEETLYKTREILELLNH 122
HPREEINSLIEENDAGESACTCSSENDSYGYVVEENQSLTVNDVEETLYKTREILELLNH
Sbjct: 223 HPREEINSLIEENDAGESACTCSSENDSYGYVVEENQSLTVNDVEETLYKTREILELLNH 402
Query: 123 EFNEDGQLFKRPFNVPENQKFTVGLDIPFSKLKMELLRGGSSTLVLTGLGGLGKTTLATK 182
EFNEDGQLFKRPFNVPENQKFTVGLDIPFSKLKMELLRGGSSTLVLTGLGGLGKTTLATK
Sbjct: 403 EFNEDGQLFKRPFNVPENQKFTVGLDIPFSKLKMELLRGGSSTLVLTGLGGLGKTTLATK 582
Query: 183 LCWDE 187
LCWDE
Sbjct: 583 LCWDE 597
Score = 31.2 bits (69), Expect = 1.7
Identities = 16/44 (36%), Positives = 22/44 (49%)
Frame = -3
Query: 705 LELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFG 748
L L++S L +P S GKL L ++S C +S P E G
Sbjct: 2299 LAQLAMSRFFRLHKLPNSSGKLERLMQFEMSRCFRFNSFPMEVG 2168
>TC86670 weakly similar to GP|9758141|dbj|BAB08633.1 disease resistance
protein-like {Arabidopsis thaliana}, partial (7%)
Length = 2417
Score = 459 bits (1182), Expect = e-129
Identities = 235/346 (67%), Positives = 278/346 (79%), Gaps = 2/346 (0%)
Frame = +3
Query: 340 QGLPLTIKVIATSLKNRPRDMWRKIGKELSQGHSILDSNTDLLTRLQKIFDVLEDNPTIM 399
+GLPL IKVIATS + RP ++W KI KELS+G SILDSNT+LL RLQKI DVLEDN
Sbjct: 3 KGLPLAIKVIATSFRYRPYELWEKIVKELSRGRSILDSNTELLIRLQKILDVLEDNAISK 182
Query: 400 ECFMDIALFPEDHRIPVAALVDMWAELYRLDDNGIQAMEIINKLGIMNLANVIIPRKNAS 459
ECFMD+ALFPED RIPVAAL+DMWAELY LDD+G +AM+IINKL MNLANV+I RKNAS
Sbjct: 183 ECFMDLALFPEDQRIPVAALIDMWAELYGLDDDGKEAMDIINKLDSMNLANVLIARKNAS 362
Query: 460 DTDNNNYNNHFIILHDILRELGIYQSAKEPFEQRKRLIIDINKNK--SGLAEKQQGLMTC 517
DT+N YNNHFI+LHD+LRELG YQ+ +EP EQRKR +I+ N++K L EKQQG M
Sbjct: 363 DTENYYYNNHFIVLHDLLRELGNYQNTQEPIEQRKRQLINTNESKCDQRLREKQQGTMAH 542
Query: 518 ILSKFMRLCVKRNPQQLTARILSVSADETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWI 577
ILSK + K PQ++ AR LS+S DETCA DWSQ++PA EVLILNL TKQY+ PE +
Sbjct: 543 ILSKLIGWFDKPKPQKVPARTLSISIDETCASDWSQVQPALAEVLILNLQTKQYTFPELM 722
Query: 578 AKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLERIYVPSFGTLKNLKKLSLYMC 637
KM+KL+ LI+ N+ PS+LNN+ELL SL NL+RIRLERI VPSFGTLKNLKKLSLYMC
Sbjct: 723 EKMNKLKALIVINHGLRPSELNNLELLSSLSNLKRIRLERISVPSFGTLKNLKKLSLYMC 902
Query: 638 NTILAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKK 683
NT LAFEKGSILISD FPNLE+L++DYCKD+ LP G +L++K
Sbjct: 903 NTRLAFEKGSILISDLFPNLEDLSMDYCKDMTALPNGGV*YYLVEK 1040
Score = 223 bits (569), Expect = 2e-58
Identities = 109/141 (77%), Positives = 123/141 (86%)
Frame = +1
Query: 673 IGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHL 732
+G+CDI LKKL +TNCHKLS LPQ+IGKLENLELLSL SCTDL +P SIG+L NL+ L
Sbjct: 1009 MGVCDIISLKKLSITNCHKLSLLPQEIGKLENLELLSLISCTDLVELPDSIGRLSNLRLL 1188
Query: 733 DISNCISLSSLPEEFGNLCNLKNLDMASCASIELPFSVVNLQNLKTITCDEETAATWEDF 792
DISNCISLSSLPE+FGNLCNL+NLDM SCAS ELPFSVVNLQNLK +TCDEETAA+WE F
Sbjct: 1189 DISNCISLSSLPEDFGNLCNLRNLDMTSCASCELPFSVVNLQNLK-VTCDEETAASWESF 1365
Query: 793 QHMLPNMKIEVLHVDVNLNWL 813
Q M+ N+ IEV HV+VNLNWL
Sbjct: 1366 QSMISNLTIEVPHVEVNLNWL 1428
Score = 29.6 bits (65), Expect = 4.8
Identities = 17/45 (37%), Positives = 24/45 (52%)
Frame = -1
Query: 705 LELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGN 749
L L +S L +P S GKL L L++S+ + SLP E G+
Sbjct: 1280 LAQLVMSRFRRLHRLPKSSGKLERLMQLEMSSSLRFESLPIESGS 1146
>TC87505 weakly similar to PIR|T05981|T05981 hypothetical protein F17M5.60 -
Arabidopsis thaliana, partial (45%)
Length = 2140
Score = 315 bits (807), Expect = 4e-86
Identities = 185/565 (32%), Positives = 320/565 (55%), Gaps = 8/565 (1%)
Frame = +3
Query: 257 LLLVLDDVWPISEPLVEKIQFQISDFKILVTSRVAFPRFSTTCILKPLAHEEAVTLFHHY 316
+L++LDDVW S ++E++ F++ + K +V SR FP F+ T ++ L ++A++LF H+
Sbjct: 396 ILVILDDVW--SPSVLEQLVFRMPNCKFIVVSRFIFPIFNATYKVELLDKDDALSLFCHH 569
Query: 317 AQMEKNSSDIINKNLVEKVVRSCQGLPLTIKVIATSLKNRPRDMWRKIGKELSQGHSILD 376
A +K+ N+NLV++VV C LPL +KVI SL+++ W + LSQG SI +
Sbjct: 570 AFGQKSIPFAANQNLVKQVVAECGNLPLALKVIGASLRDQNEMFWLSVKTRLSQGLSIDE 749
Query: 377 S-NTDLLTRLQKIFDVLEDNPTIMECFMDIALFPEDHRIPVAALVDMWAELYRLDDNGIQ 435
S +L+ R+ + L + I ECF+D+ FPED +IP+ L++MW E++ + + +
Sbjct: 750 SYERNLIDRMAISTNYLPEK--IKECFLDLCSFPEDKKIPLEVLINMWVEIHDIHET--E 917
Query: 436 AMEIINKLGIMNLANVIIPRKNASDTDNNNYNNHF---IILHDILRELGIYQSAKEPFEQ 492
A I+ +L NL ++ + Y++ F + HDILR+L + S + Q
Sbjct: 918 AYAIVVELSNKNLLTLVEEARAGG-----MYSSCFEISVTQHDILRDLALNLSNRGNINQ 1082
Query: 493 RKRLIIDINKNKSGLAEKQQGLMTCILSKFMRLCVKRNPQQLTARILSVSADETCAFDWS 552
R+RL++ ++ L ++ ++ Q A+I+S+ E DW
Sbjct: 1083RRRLVMPKREDNGQLPKEW---------------LRYADQPFEAQIVSIHTGEMRKSDWC 1217
Query: 553 QMEPAQVEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLER 612
+E + EVLI+N + +Y LP +I +M KLR L++ N+ + L+NI + +L NL
Sbjct: 1218NLEFPKAEVLIINFTSSEYFLPPFINRMPKLRALMVINHSTSYACLHNISVFKNLTNLRS 1397
Query: 613 IRLERIYVPSFG--TLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVV 670
+ E++ +P +++L+KL + +C + E I+D FPN+ EL +D+C+D+
Sbjct: 1398LWFEKVSIPHLSGIVMESLRKLFIVLCKINNSLEGKDSNIADIFPNISELTLDHCEDVTE 1577
Query: 671 LPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLK 730
LP IC I L+ L +TNCH L+ LP ++G L LE+L L +C +L +P SI + LK
Sbjct: 1578LPSSICRIQSLQNLSLTNCHSLTRLPIELGSLRYLEILRLYACPNLRTLPPSICGMTRLK 1757
Query: 731 HLDISNCISLSSLPEEFGNLCNLKNLDMASCASI-ELPFSVVNLQNLKTI-TCDEETAAT 788
++DI+ C+ L+S P+ G L NL+ +DM C I +P S ++L ++ CD+E +
Sbjct: 1758YIDIAQCVYLASFPDAIGKLVNLEKIDMRECPMITNIPKSALSLNFFCSL*ICDDEVSWM 1937
Query: 789 WEDFQHMLPNMKIEVLHVDVNLNWL 813
W++ Q + N+ I+V+ ++ +L+WL
Sbjct: 1938WKEVQKVKLNVDIQVVEIEYDLDWL 2012
Score = 35.0 bits (79), Expect = 0.12
Identities = 20/52 (38%), Positives = 33/52 (63%), Gaps = 2/52 (3%)
Frame = +2
Query: 140 NQKFTVGLDIPFSKLKMELLRGGSSTLV--LTGLGGLGKTTLATKLCWDEEV 189
N + GL+ +K+ ME++ G + ++G+GG GKTTLA ++C DE+V
Sbjct: 71 NLDLSFGLEFGKNKV-MEMVVGRKDFCLVGISGIGGSGKTTLAREICRDEQV 223
>BG646517
Length = 668
Score = 301 bits (771), Expect = 7e-82
Identities = 150/167 (89%), Positives = 160/167 (94%)
Frame = +2
Query: 245 LGLLLKKVEGSPLLLVLDDVWPISEPLVEKIQFQISDFKILVTSRVAFPRFSTTCILKPL 304
L LLLKKVEGSPLLLVLDDVWP SE LV+K+QF+ISDFKILVTSRV+FPRF TTCILKPL
Sbjct: 161 LELLLKKVEGSPLLLVLDDVWPTSESLVKKLQFEISDFKILVTSRVSFPRFRTTCILKPL 340
Query: 305 AHEEAVTLFHHYAQMEKNSSDIINKNLVEKVVRSCQGLPLTIKVIATSLKNRPRDMWRKI 364
AHE+AVTLFHHYAQMEKNSSDII+KNLVEKVVRSC+GLPLTIKVIATSL+NRP D+WRKI
Sbjct: 341 AHEDAVTLFHHYAQMEKNSSDIIDKNLVEKVVRSCKGLPLTIKVIATSLRNRPDDLWRKI 520
Query: 365 GKELSQGHSILDSNTDLLTRLQKIFDVLEDNPTIMECFMDIALFPED 411
ELSQGHSILDSNT+LLTRLQKIFDVLEDNPTIMECFMDIALFPED
Sbjct: 521 VMELSQGHSILDSNTELLTRLQKIFDVLEDNPTIMECFMDIALFPED 661
>TC89031 weakly similar to GP|9758141|dbj|BAB08633.1 disease resistance
protein-like {Arabidopsis thaliana}, partial (5%)
Length = 1294
Score = 300 bits (769), Expect = 1e-81
Identities = 155/274 (56%), Positives = 201/274 (72%), Gaps = 4/274 (1%)
Frame = +3
Query: 544 DETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIEL 603
DE+ + DW M+P +VEVL+LNL + QYSLP++ KMSKL+VLI+TNY FH S+L EL
Sbjct: 123 DESFSSDWCDMQPDEVEVLVLNLQSDQYSLPDFTDKMSKLKVLIVTNYGFHRSELIKFEL 302
Query: 604 LGSLQNLERIRLERIYVPSFGTLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNID 663
LG L NL+RIRLE++ VP LKNL+KLSL+MCNT AFE SI ISDA PNL EL+ID
Sbjct: 303 LGFLSNLKRIRLEKVSVPCLSILKNLQKLSLHMCNTRDAFENYSIQISDAMPNLVELSID 482
Query: 664 YCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSI 723
YC DL+ LP G +I LKK+ +TNCHKLS++PQDI KLENLE+L L SC+DL I S+
Sbjct: 483 YCNDLIKLPDGFSNITTLKKISITNCHKLSAIPQDIEKLENLEVLRLCSCSDLVEISESV 662
Query: 724 GKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCASI-ELPFSVVNLQNLK---TI 779
L L+ DIS+C+SLS LP + G+L L+ M C+++ ELP+SV+NL N+K +
Sbjct: 663 SGLNKLRCFDISDCVSLSKLPNDIGDLKKLEKFYMKGCSNLSELPYSVINLGNVKHEIHV 842
Query: 780 TCDEETAATWEDFQHMLPNMKIEVLHVDVNLNWL 813
CDEE AA WE F + +PN+KI++ V++NLNWL
Sbjct: 843 ICDEEGAALWEHFPN-IPNLKIDMPKVEINLNWL 941
>TC88589 weakly similar to GP|9758140|dbj|BAB08632.1 disease resistance
protein-like {Arabidopsis thaliana}, partial (12%)
Length = 1137
Score = 300 bits (768), Expect = 1e-81
Identities = 161/307 (52%), Positives = 210/307 (67%), Gaps = 4/307 (1%)
Frame = +1
Query: 511 QQGLMTCILSKFMRLCVKRNPQQLTARILSVSADETCAFDWSQMEPAQVEVLILNLHTKQ 570
QQG+++ S + VK+ ++ ARIL +S DE + DW M+P + EVL+LNL + Q
Sbjct: 25 QQGIISNWYSFITGMLVKQKQLKVAARILCISTDEIFSSDWCDMQPDKAEVLVLNLRSDQ 204
Query: 571 YSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLERIYVPSFGTLKNLK 630
YSLP++ KM KL+VLI+TNY F S+L ELLGSL NL+RIRLE++ VP LKNL+
Sbjct: 205 YSLPDFTKKMRKLKVLIVTNYGFSRSELTKFELLGSLSNLKRIRLEKVSVPCLCILKNLR 384
Query: 631 KLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCH 690
KLSL+MC+T AFE SI ISDA PNL EL+IDYC DL+ LP C I LKKL +TNCH
Sbjct: 385 KLSLHMCSTNNAFESCSIQISDAMPNLVELSIDYCNDLIKLPGEFCKITTLKKLSITNCH 564
Query: 691 KLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNL 750
K S++PQDIGKL NLE+L L SC+DL+ IP S+ L L+ LDIS+C++L LP GNL
Sbjct: 565 KFSAMPQDIGKLVNLEVLRLCSCSDLKEIPESVADLNKLRCLDISDCVTLHILPNNIGNL 744
Query: 751 CNLKNLDMASCASI-ELPFSVVNLQNLK---TITCDEETAATWEDFQHMLPNMKIEVLHV 806
L+ L M C+++ ELP SV+N NLK + CDEE +A WE + +P +KI + V
Sbjct: 745 QKLEKLYMKGCSNLSELPDSVINFGNLKHEMQVICDEEGSALWEHLSN-IPKLKIYMPKV 921
Query: 807 DVNLNWL 813
+ NL WL
Sbjct: 922 EHNLIWL 942
>TC90098 weakly similar to GP|9758140|dbj|BAB08632.1 disease resistance
protein-like {Arabidopsis thaliana}, partial (16%)
Length = 1128
Score = 291 bits (744), Expect = 9e-79
Identities = 164/351 (46%), Positives = 221/351 (62%), Gaps = 6/351 (1%)
Frame = +1
Query: 469 HFIILHDILRELGIYQSAKEPFEQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMRLCVK 528
H++ H +LR+L I +++E ++R RLIID + N
Sbjct: 1 HYVTQHGLLRDLAIRDNSQEAEDKRNRLIIDTSANN-----------------LPSWWTS 129
Query: 529 RNPQQLTARILSVSADETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLRVLII 588
N + ARILS+S DE W ++P +VEVL+LNL K+++LP ++ KM KL+ LII
Sbjct: 130 ENEYHIKARILSISTDEIFTSKWCNLQPTEVEVLVLNLREKKFALPMFMKKMKKLKALII 309
Query: 589 TNYDFHPSKLNNIELLGSLQNLERIRLERIYVPSFG----TLKNLKKLSLYMCNTILAFE 644
TNYDF P++L N E+L L NL+RIRLE++ VP G LKNL+K S +MCN AFE
Sbjct: 310 TNYDFFPAELENFEVLDHLSNLKRIRLEKVSVPFLGKTVLQLKNLQKCSFFMCNVNKAFE 489
Query: 645 KGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLEN 704
+I S+ PNL E+N DYC D+V LP I +I LKKL +TNCHKL +L + IGKL N
Sbjct: 490 NCTIEDSEMLPNLMEMNFDYC-DMVELPNVISNIVSLKKLSITNCHKLCALHEGIGKLVN 666
Query: 705 LELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCASI 764
LE L LSSC+ L +P SI L LK L+IS CISLS LPE G L L+ L+M C++I
Sbjct: 667 LESLRLSSCSGLLELPNSIRNLHVLKFLNISGCISLSQLPENIGELEKLEKLNMRGCSNI 846
Query: 765 -ELPFSVVNLQNLKTITC-DEETAATWEDFQHMLPNMKIEVLHVDVNLNWL 813
ELP SV+ L+ LK + C DEETA WE ++++L ++K+EV+ D NLN+L
Sbjct: 847 SELPSSVMELEGLKHVVCGDEETAEKWEPYKNILGDLKVEVVQEDFNLNFL 999
>CB892890 weakly similar to GP|9758141|dbj| disease resistance protein-like
{Arabidopsis thaliana}, partial (4%)
Length = 852
Score = 287 bits (735), Expect = 1e-77
Identities = 153/305 (50%), Positives = 204/305 (66%), Gaps = 3/305 (0%)
Frame = +2
Query: 386 QKIFD-VLEDNPTIMECFMDIALFPEDHRIPVAALVDMWAELYRLDDNGIQAMEIINKLG 444
QK+ + L+DN + ECFMD+ LF ED +IPVAAL+DMW ELY LDD+ I+ M I+ KL
Sbjct: 2 QKVLENALQDNHIVKECFMDLGLFFEDKKIPVAALIDMWTELYDLDDDNIKGMNIVRKLA 181
Query: 445 IMNLANVIIPRKNASDTDNNNYNNHFIILHDILRELGIYQSAKEPFEQRKRLIIDINKNK 504
+L +++ R+ + D+ YN+HF+ HD+L+E+ I+Q+ +EPFE RKRLI D+N+N
Sbjct: 182 NWHLVKLVVSREVTTHVDHY-YNHHFLTQHDLLKEISIHQARQEPFELRKRLIFDVNENS 358
Query: 505 SGLAEKQQGLMTCILSKFMRLCVKRNPQQLTARILSVSADETCAFDWSQMEPA-QVEVLI 563
++N Q AR LS+S D+ DWS +E QVEVLI
Sbjct: 359 WD---------------------QQNQQNTIARTLSISPDKILTSDWSNVEKIKQVEVLI 475
Query: 564 LNLHTKQYSLPEWIAKMSKLRVLIITNYD-FHPSKLNNIELLGSLQNLERIRLERIYVPS 622
LNLHT++Y+LPE I KM+KL+VLIITNY FH ++L+N E+LG L NL RIRL ++ VPS
Sbjct: 476 LNLHTEKYTLPECIKKMTKLKVLIITNYKGFHCAELDNFEILGCLPNLRRIRLHQVSVPS 655
Query: 623 FGTLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLK 682
L NL+KLSLY C T AF+ ++ ISD PNL+EL +DYCKDLV LP G+CDI LK
Sbjct: 656 LCKLVNLRKLSLYFCETKQAFQSNTVSISDILPNLKELCVDYCKDLVTLPSGLCDITSLK 835
Query: 683 KLRVT 687
KL +T
Sbjct: 836 KLSIT 850
>BG646612 weakly similar to GP|9758140|dbj| disease resistance protein-like
{Arabidopsis thaliana}, partial (7%)
Length = 735
Score = 280 bits (716), Expect = 2e-75
Identities = 148/188 (78%), Positives = 164/188 (86%), Gaps = 5/188 (2%)
Frame = +2
Query: 8 ITLTTRFQFNDVLNKISQILQKAQKNQSSRRSLRLILKDLSPVVQDIKQYNDHLDHPREE 67
ITLTTRF +DVLNKISQI++ A+K+QS+R+ LR ILKDL+ VVQDIKQYN+HLDHPREE
Sbjct: 23 ITLTTRF--HDVLNKISQIVETARKHQSARQLLRSILKDLTLVVQDIKQYNEHLDHPREE 196
Query: 68 INSLIEENDAGESACTCSSENDSYGYVVEENQSLTVNDVEETLYKTREILELLN-----H 122
I +L+EENDA ESAC CSSENDS YVV+ENQSL VNDVEETLYK REILELLN H
Sbjct: 197 IKTLLEENDAEESACKCSSENDS--YVVDENQSLIVNDVEETLYKAREILELLNYETFEH 370
Query: 123 EFNEDGQLFKRPFNVPENQKFTVGLDIPFSKLKMELLRGGSSTLVLTGLGGLGKTTLATK 182
+FNE FKRPF+VPEN KFTVGLDIPFSKLKMEL+R GSSTLVLTGLGGLGKTTLATK
Sbjct: 371 KFNEAKPPFKRPFDVPENPKFTVGLDIPFSKLKMELIRDGSSTLVLTGLGGLGKTTLATK 550
Query: 183 LCWDEEVN 190
LCWD+EVN
Sbjct: 551 LCWDQEVN 574
Score = 96.7 bits (239), Expect = 3e-20
Identities = 52/82 (63%), Positives = 58/82 (70%)
Frame = +1
Query: 191 A*KSKIHCWVGYTI*QVENGASEGW*LHTCVDWFRRIGENHSGYKALLG*RS*WLGLLLK 250
+*KSKIHCWVGYTI*QVENGA +GW*LH CVD RRI ENH GYKALLG RS*W
Sbjct: 415 S*KSKIHCWVGYTI*QVENGAYKGW*LHPCVDGLRRIRENHFGYKALLGSRS*W------ 576
Query: 251 KVEGSPLLLVLDDVWPISEPLV 272
++G PL L P+ E L+
Sbjct: 577 *IQGKPLYLSPFSKTPMVEDLL 642
>CA920022 similar to PIR|T03031|T030 NBS-LRR type resistance protein - rice
(fragment), partial (8%)
Length = 706
Score = 263 bits (673), Expect = 2e-70
Identities = 134/190 (70%), Positives = 155/190 (81%)
Frame = -1
Query: 268 SEPLVEKIQFQISDFKILVTSRVAFPRFSTTCILKPLAHEEAVTLFHHYAQMEKNSSDII 327
SE LVEK +FQISD+KILVT VAF RF T ILKPLA E++VTLF HY ++EKNSS I
Sbjct: 703 SEDLVEKFKFQISDYKILVTQGVAFSRFDKTFILKPLAQEDSVTLFRHYTEVEKNSSKIP 524
Query: 328 NKNLVEKVVRSCQGLPLTIKVIATSLKNRPRDMWRKIGKELSQGHSILDSNTDLLTRLQK 387
+K+L+EKVV C+GLPL IKVIATS + RP ++W KI KELS+G SILDSNT+LL RLQK
Sbjct: 523 DKDLIEKVVEHCKGLPLAIKVIATSFRYRPYELWEKIVKELSRGRSILDSNTELLIRLQK 344
Query: 388 IFDVLEDNPTIMECFMDIALFPEDHRIPVAALVDMWAELYRLDDNGIQAMEIINKLGIMN 447
I DVLEDN ECFMD+ALFPED RIPVAAL+DMWAELY LDD+G +AM+IINKL MN
Sbjct: 343 ILDVLEDNAISKECFMDLALFPEDQRIPVAALIDMWAELYGLDDDGKEAMDIINKLDSMN 164
Query: 448 LANVIIPRKN 457
LANV+I R N
Sbjct: 163 LANVLIARYN 134
>BF643326 weakly similar to GP|11994216|db disease resistance comples protein
{Arabidopsis thaliana}, partial (2%)
Length = 486
Score = 200 bits (509), Expect(2) = 2e-55
Identities = 101/125 (80%), Positives = 110/125 (87%)
Frame = +2
Query: 618 IYVPSFGTLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICD 677
I VPSFGT+KNLKKLSLYMCNT LAFEKGSILISD FPNLE+L+IDY KD+V LP G+CD
Sbjct: 2 ISVPSFGTMKNLKKLSLYMCNTRLAFEKGSILISDLFPNLEDLSIDYSKDMVALPNGVCD 181
Query: 678 IFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNC 737
I LKKL +TNCHKLSSLPQDIGKL NLELLSL SCTDL +P SIG+LLNL+ LDISNC
Sbjct: 182 IASLKKLSITNCHKLSSLPQDIGKLMNLELLSLISCTDLVELPDSIGRLLNLRLLDISNC 361
Query: 738 ISLSS 742
ISLSS
Sbjct: 362 ISLSS 376
Score = 34.7 bits (78), Expect(2) = 2e-55
Identities = 14/18 (77%), Positives = 17/18 (93%)
Frame = +3
Query: 743 LPEEFGNLCNLKNLDMAS 760
LPE+FGNLCNL+NL M+S
Sbjct: 378 LPEDFGNLCNLRNLYMSS 431
>CB893781 homologue to PIR|C64504|C645 hypothetical protein MJ1637 -
Methanococcus jannaschii, partial (2%)
Length = 473
Score = 175 bits (444), Expect = 5e-44
Identities = 85/89 (95%), Positives = 88/89 (98%)
Frame = +2
Query: 102 TVNDVEETLYKTREILELLNHEFNEDGQLFKRPFNVPENQKFTVGLDIPFSKLKMELLRG 161
TVN+VEETLYKT+EILELLNHEFNEDGQLFKRPFNVPENQKFTVGLDIPF+KLKMELLRG
Sbjct: 2 TVNEVEETLYKTKEILELLNHEFNEDGQLFKRPFNVPENQKFTVGLDIPFTKLKMELLRG 181
Query: 162 GSSTLVLTGLGGLGKTTLATKLCWDEEVN 190
GSSTLVLTGL GLGKTTLATKLCWDEEVN
Sbjct: 182 GSSTLVLTGLRGLGKTTLATKLCWDEEVN 268
Score = 111 bits (278), Expect = 1e-24
Identities = 51/54 (94%), Positives = 53/54 (97%)
Frame = +1
Query: 191 A*KSKIHCWVGYTI*QVENGASEGW*LHTCVDWFRRIGENHSGYKALLG*RS*W 244
+*KSKIHCWVGYTI QVENGASEGW*LHTCVDWF+RIGENHSGYKALLG*RS*W
Sbjct: 109 S*KSKIHCWVGYTIYQVENGASEGW*LHTCVDWFKRIGENHSGYKALLG*RS*W 270
>BE319778
Length = 378
Score = 124 bits (310), Expect = 2e-28
Identities = 60/60 (100%), Positives = 60/60 (100%)
Frame = +2
Query: 455 RKNASDTDNNNYNNHFIILHDILRELGIYQSAKEPFEQRKRLIIDINKNKSGLAEKQQGL 514
RKNASDTDNNNYNNHFIILHDILRELGIYQSAKEPFEQRKRLIIDINKNKSGLAEKQQGL
Sbjct: 197 RKNASDTDNNNYNNHFIILHDILRELGIYQSAKEPFEQRKRLIIDINKNKSGLAEKQQGL 376
Score = 41.6 bits (96), Expect = 0.001
Identities = 20/20 (100%), Positives = 20/20 (100%)
Frame = +1
Query: 436 AMEIINKLGIMNLANVIIPR 455
AMEIINKLGIMNLANVIIPR
Sbjct: 1 AMEIINKLGIMNLANVIIPR 60
>BG647778 weakly similar to PIR|T48468|T484 disease resistance-like protein -
Arabidopsis thaliana, partial (4%)
Length = 660
Score = 122 bits (305), Expect = 7e-28
Identities = 90/219 (41%), Positives = 119/219 (54%), Gaps = 7/219 (3%)
Frame = +2
Query: 136 NVPENQKFTVGLDIPFSKLKMELLRGGSSTLVLTGLGGLGKTTLATKLCWDEEVNA*KSK 195
+ P TVGLD P +LK LL G S LVLTGL G GKTTLAT LCWD++V +
Sbjct: 5 DAPVKPDLTVGLDFPLIQLKSWLLGSGVSVLVLTGLAGSGKTTLATLLCWDDKVRGKFGE 184
Query: 196 IHCWVGYTI*QVENGASEGW*LHTCVDW-FRRIGENH----SGYKALLG*RS*WLGLLLK 250
+ +T+ ++ N L V FR G + A+ RS LL K
Sbjct: 185 NILF--FTVSKIPN-------LKNIVQTLFRHCGHDEPCLIDDDDAVKHLRS----LLTK 325
Query: 251 KVEGSPLLLVLDDVWPISEPLVEKIQFQISDFKILVTSRVAFPRFSTTCILKPLAHEEAV 310
E P++LVLD+V P SE LVE Q Q+ D KIL+TSRV FPRFS + LKPL ++AV
Sbjct: 326 IGESCPMMLVLDNVCPGSESLVEDFQVQVPDCKILITSRVEFPRFS-SLFLKPLRDDDAV 502
Query: 311 TLFHHYA--QMEKNSSDIINKNLVEKVVRSCQGLPLTIK 347
TLF +A ++ + + V++V + C G PL +K
Sbjct: 503 TLFGSFALPNDATRATYVPAEKYVKQVAKGCWGSPLALK 619
>TC77571 homologue to GP|20804846|dbj|BAB92528. putative rust resistance
protein {Oryza sativa (japonica cultivar-group)},
partial (1%)
Length = 954
Score = 117 bits (294), Expect = 1e-26
Identities = 77/166 (46%), Positives = 97/166 (58%), Gaps = 4/166 (2%)
Frame = +2
Query: 105 DVEETLYKTREILELLNHEFNEDGQLFKRPFNVPENQKFTVGLDIPFSKLKMELLRGGSS 164
DVE L + +IL+ N G + K PF+VP N +FTVGLD+ KLK+E+LR G S
Sbjct: 491 DVEGKLRELLKILDKENFGKKISGSILKGPFDVPANPEFTVGLDLQLIKLKVEILREGRS 670
Query: 165 TLVLTGLGGLGKTTLATKLCWDEEVNA*KSKIHCWVGYTI*QVENGASEGW*LHTCVDWF 224
TL+LTGLGG+GKTTLATKLC D++V K K EN + H C
Sbjct: 671 TLLLTGLGGMGKTTLATKLCLDDQV---KGKFK----------ENIIFVTFSKHPC*RLL 811
Query: 225 RRIGENHSGYKAL----LG*RS*WLGLLLKKVEGSPLLLVLDDVWP 266
R N + + L + R LGLLL+K+EGSP+LLVLDDVWP
Sbjct: 812 *RDYLNIADIRCLSTKVMKMRENGLGLLLRKIEGSPILLVLDDVWP 949
>AW586679 similar to GP|9758140|dbj| disease resistance protein-like
{Arabidopsis thaliana}, partial (4%)
Length = 481
Score = 105 bits (263), Expect = 5e-23
Identities = 69/169 (40%), Positives = 99/169 (57%), Gaps = 1/169 (0%)
Frame = +3
Query: 171 LGGLGKTTLATKLCWDEEVNA*KSKIHCWVGY-TI*QVENGASEGW*LHTCVDWFRRIGE 229
LGG GKTTLA KLCW+++ K K + + T+ + N + + T + R
Sbjct: 3 LGGSGKTTLAKKLCWEQDF---KGKFGGNIFFVTVSETPNLKNI---VKTLFEQCGRPVP 164
Query: 230 NHSGYKALLG*RS*WLGLLLKKVEGSPLLLVLDDVWPISEPLVEKIQFQISDFKILVTSR 289
+ + + + LG LL + E S +LLVLDDVWP SE LVEK F++ D+KILVTSR
Sbjct: 165 DFTNEEDAIN----QLGHLLSQFERSQILLVLDDVWPGSESLVEKFTFKLPDYKILVTSR 332
Query: 290 VAFPRFSTTCILKPLAHEEAVTLFHHYAQMEKNSSDIINKNLVEKVVRS 338
V F RF T C L PL + A +LF HYAQ+ S + + +LV+++V++
Sbjct: 333 VGFRRFGTPCQLNPLDEDPAASLFRHYAQLHHIISYMPDGDLVDEIVKA 479
>TC92621 similar to GP|7107252|gb|AAF36340.1| unknown {Pennisetum glaucum},
partial (10%)
Length = 827
Score = 103 bits (258), Expect = 2e-22
Identities = 70/163 (42%), Positives = 92/163 (55%)
Frame = +2
Query: 104 NDVEETLYKTREILELLNHEFNEDGQLFKRPFNVPENQKFTVGLDIPFSKLKMELLRGGS 163
+D +T Y+ +EIL +FK PF VPEN +FTVGLD K KMELLR G
Sbjct: 344 HDKPKTFYEFKEILFFDKKISGYGVSIFKGPFGVPENPEFTVGLDSQLIKPKMELLRDGR 523
Query: 164 STLVLTGLGGLGKTTLATKLCWDEEVNA*KSKIHCWVGYTI*QVENGASEGW*LHTCVDW 223
STL+LTGLGG+GKTTLATKLC D++V + +V ++ + + H
Sbjct: 524 STLLLTGLGGMGKTTLATKLCLDQQVKGKFKENIIFVTFSKTPMLKIMVQRLFEHGGYP- 700
Query: 224 FRRIGENHSGYKALLG*RS*WLGLLLKKVEGSPLLLVLDDVWP 266
+ E S A+ G +G LL+K+EGSP+L VLDDV P
Sbjct: 701 ---VPEYQSDEXAVNG-----MGNLLRKIEGSPILXVLDDVGP 805
>TC79479 similar to GP|12056928|gb|AAG48132.1 putative resistance protein
{Glycine max}, partial (21%)
Length = 1087
Score = 85.5 bits (210), Expect = 7e-17
Identities = 66/201 (32%), Positives = 101/201 (49%)
Frame = +1
Query: 579 KMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLERIYVPSFGTLKNLKKLSLYMCN 638
KM+ L+ LII + F N L SL+ LE +PS+ K L L L +
Sbjct: 97 KMTGLQTLIIRSLCFAEGPKN---LPNSLRVLEWWGYPSQSLPSYFYPKKLAVLKLPHSS 267
Query: 639 TILAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQD 698
F + S F N+ LN D CK + +P + L++L + +C L +
Sbjct: 268 ----FMSLELSKSKKFVNMTLLNFDECKIITHIP-DVSGAPNLERLSLDSCENLVEIHDS 432
Query: 699 IGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDM 758
+G L+ LE+L+L SC L +P L +L+HL++S+C SL S PE GN+ N+ +L +
Sbjct: 433 VGFLDKLEILNLGSCAKLRNLPPI--HLTSLQHLNLSHCSSLVSFPEILGNMKNITSLSL 606
Query: 759 ASCASIELPFSVVNLQNLKTI 779
A E P+S+ NL LK++
Sbjct: 607 EYTAIREFPYSIGNLPRLKSL 669
>BQ750499 similar to OMNI|NT01MC4764 transport protein HasD putative
{Magnetococcus sp. MC-1}, partial (3%)
Length = 800
Score = 84.7 bits (208), Expect = 1e-16
Identities = 63/198 (31%), Positives = 101/198 (50%), Gaps = 26/198 (13%)
Frame = +1
Query: 18 DVLNKISQILQKAQKNQSSRRSLRLILKDLSPVVQDIKQYNDHLDHPREEINSLIEENDA 77
+V+ K Q ++K ++ + L +L+P+V++IK YND LD P EEI L +
Sbjct: 67 EVVKKALQTIKKGREFGPTLERNIETLNNLAPLVKEIKVYNDFLDRPSEEIERLEKHIRE 246
Query: 78 GESACTCSSENDSYGYV---------------VEENQSLTVN-----DVEETLYKTREI- 116
GE S + + ++ ++ + S+ V D+ + + K EI
Sbjct: 247 GEELVRKSKKLTLWNFLSFPGYQGKLKKKDEGLQRHLSVNVQLENKKDLIKLVAKVDEIS 426
Query: 117 --LELLNHEFNE---DGQLFKRPFNVPENQKFTVGLDIPFSKLKMELLRGGSSTLVLTGL 171
E+L + N DG + PE + +G+D P +KLK++LL+ G S LVLTGL
Sbjct: 427 KIFEILMRKENLGQFDGSQIRGLCGAPEEPQ-CMGVDEPLNKLKIQLLKDGVSVLVLTGL 603
Query: 172 GGLGKTTLATKLCWDEEV 189
GG GK+TLA KLCW+ ++
Sbjct: 604 GGSGKSTLAKKLCWNPQI 657
Score = 30.8 bits (68), Expect = 2.2
Identities = 16/32 (50%), Positives = 18/32 (56%)
Frame = +3
Query: 207 VENGASEGW*LHTCVDWFRRIGENHSGYKALL 238
VE+ E W TC DWFR I HS +ALL
Sbjct: 549 VEDSIIERWCFCTCFDWFRWIW*KHSC*EALL 644
>AW683902 weakly similar to PIR|T48468|T484 disease resistance-like protein -
Arabidopsis thaliana, partial (4%)
Length = 262
Score = 67.8 bits (164), Expect(2) = 3e-13
Identities = 30/48 (62%), Positives = 38/48 (78%)
Frame = +1
Query: 655 PNLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKL 702
PNL EL+IDYC DL+ LP +C+I LKKL +TNCHKLS +P+DI K+
Sbjct: 1 PNLVELSIDYCNDLIKLPDELCNITTLKKLSITNCHKLSLMPRDIWKV 144
Score = 36.2 bits (82), Expect = 0.052
Identities = 17/43 (39%), Positives = 24/43 (55%)
Frame = +1
Query: 704 NLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEE 746
NL LS+ C DL +P + + LK L I+NC LS +P +
Sbjct: 4 NLVELSIDYCNDLIKLPDELCNITTLKKLSITNCHKLSLMPRD 132
Score = 25.8 bits (55), Expect(2) = 3e-13
Identities = 11/14 (78%), Positives = 12/14 (85%)
Frame = +2
Query: 700 GKLENLELLSLSSC 713
GKLENLE+L L SC
Sbjct: 137 GKLENLEVLRLCSC 178
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,134,223
Number of Sequences: 36976
Number of extensions: 385509
Number of successful extensions: 2544
Number of sequences better than 10.0: 312
Number of HSP's better than 10.0 without gapping: 2205
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2401
length of query: 813
length of database: 9,014,727
effective HSP length: 104
effective length of query: 709
effective length of database: 5,169,223
effective search space: 3664979107
effective search space used: 3664979107
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 62 (28.5 bits)
Medicago: description of AC148347.2


