

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148289.16 + phase: 0
(200 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
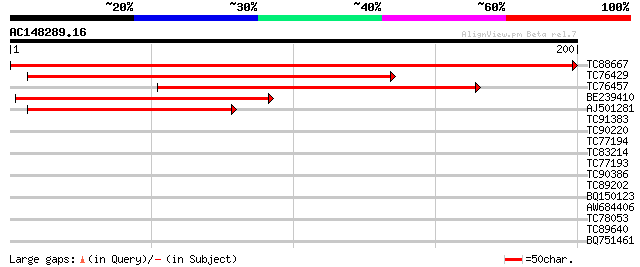
Score E
Sequences producing significant alignments: (bits) Value
TC88667 similar to GP|6728982|gb|AAF26980.1| unknown protein {Ar... 409 e-115
TC76429 similar to GP|4056425|gb|AAC97999.1| ESTs gb|H36249 gb|... 163 4e-41
TC76457 similar to GP|4056425|gb|AAC97999.1| ESTs gb|H36249 gb|... 162 1e-40
BE239410 similar to GP|4056425|gb|A ESTs gb|H36249 gb|AA59732 a... 103 6e-23
AJ501281 similar to GP|4056425|gb|A ESTs gb|H36249 gb|AA59732 a... 89 1e-18
TC91383 similar to GP|9858782|gb|AAG01129.1| BAC19.14 {Lycopersi... 32 0.16
TC90220 weakly similar to PIR|T01152|T01152 probable pectinester... 29 1.1
TC77194 homologue to GP|21593495|gb|AAM65462.1 glycine rich prot... 29 1.1
TC83214 similar to SP|O49204|KAPS_CATRO Adenylylsulfate kinase ... 28 3.1
TC77193 homologue to GP|21593495|gb|AAM65462.1 glycine rich prot... 27 4.0
TC90386 weakly similar to GP|5882724|gb|AAD55277.1| F25A4.4 {Ara... 27 5.3
TC89202 weakly similar to SP|O48923|C7DA_SOYBN Cytochrome P450 7... 27 6.9
BQ150123 similar to GP|21109992|gb| TonB-dependent receptor {Xan... 27 6.9
AW684406 weakly similar to GP|21536774|gb| unknown {Arabidopsis ... 26 9.0
TC78053 similar to GP|5923675|gb|AAD56326.1| putative mRNA cappi... 26 9.0
TC89640 similar to GP|16648927|gb|AAL24315.1 formyltetrahydrofol... 26 9.0
BQ751461 weakly similar to GP|18376015|em conserved hypothetical... 26 9.0
>TC88667 similar to GP|6728982|gb|AAF26980.1| unknown protein {Arabidopsis
thaliana}, partial (79%)
Length = 1018
Score = 409 bits (1050), Expect = e-115
Identities = 200/200 (100%), Positives = 200/200 (100%)
Frame = +3
Query: 1 MIVEVENIDEDDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPE 60
MIVEVENIDEDDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPE
Sbjct: 189 MIVEVENIDEDDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPE 368
Query: 61 ENLDFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTG 120
ENLDFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTG
Sbjct: 369 ENLDFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTG 548
Query: 121 CLVGCFRKLQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLRQCLYSIIYQYQGAS 180
CLVGCFRKLQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLRQCLYSIIYQYQGAS
Sbjct: 549 CLVGCFRKLQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLRQCLYSIIYQYQGAS 728
Query: 181 KKRRLMYQDENIQKPRLTSF 200
KKRRLMYQDENIQKPRLTSF
Sbjct: 729 KKRRLMYQDENIQKPRLTSF 788
>TC76429 similar to GP|4056425|gb|AAC97999.1| ESTs gb|H36249 gb|AA59732 and
gb|AA651219 come from this gene. {Arabidopsis thaliana},
partial (55%)
Length = 666
Score = 163 bits (413), Expect = 4e-41
Identities = 74/130 (56%), Positives = 97/130 (73%)
Frame = +3
Query: 7 NIDEDDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFL 66
N D+ D+ +PP NF+MV++ I+RS P ++F F+++L LRS+I LCPEPYPE +FL
Sbjct: 276 NSDDADEAFVPPLNFAMVDNGIFRSGFPDSANFGFMKSLRLRSVICLCPEPYPEATAEFL 455
Query: 67 KEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCF 126
IRL+QFGI+G E + D I EALKV++DVRNHP+L+HCK+GKHRTGCLVGC
Sbjct: 456 NANGIRLYQFGIDGCKEPFVNIPNDKIREALKVVLDVRNHPVLIHCKRGKHRTGCLVGCI 635
Query: 127 RKLQNWCLSS 136
R+LQ CLSS
Sbjct: 636 RRLQRRCLSS 665
>TC76457 similar to GP|4056425|gb|AAC97999.1| ESTs gb|H36249 gb|AA59732 and
gb|AA651219 come from this gene. {Arabidopsis thaliana},
partial (57%)
Length = 741
Score = 162 bits (409), Expect = 1e-40
Identities = 73/114 (64%), Positives = 88/114 (77%)
Frame = +1
Query: 53 LCPEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHC 112
LCPEPYPE +FL IRL+QFGI+G E + D I EALKV++DVRNHP+L+HC
Sbjct: 211 LCPEPYPEATAEFLNANGIRLYQFGIDGCKEPFVNIPNDKIREALKVVLDVRNHPVLIHC 390
Query: 113 KQGKHRTGCLVGCFRKLQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLR 166
K+GKHRTGCLVGC R+LQ WCLSS F+EYQRFAG K+R +D FIE FD+ SL+
Sbjct: 391 KRGKHRTGCLVGCIRRLQRWCLSSIFDEYQRFAGAKARVSDQRFIELFDISSLK 552
>BE239410 similar to GP|4056425|gb|A ESTs gb|H36249 gb|AA59732 and
gb|AA651219 come from this gene. {Arabidopsis thaliana},
partial (34%)
Length = 540
Score = 103 bits (256), Expect = 6e-23
Identities = 50/91 (54%), Positives = 66/91 (71%)
Frame = +2
Query: 3 VEVENIDEDDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEEN 62
V+V + + +D+ IPP NF+MV++ I+RS P+PS+F FLQTL L SIIYLCPEPYPE N
Sbjct: 266 VDVCDKIDGEDLFIPPLNFAMVDNGIFRSGFPEPSNFSFLQTLGLGSIIYLCPEPYPEAN 445
Query: 63 LDFLKEQNIRLFQFGIEGKTEVSLPALRDSI 93
L+FLK I+L+ FGIEG E + D+I
Sbjct: 446 LEFLKSNGIKLYHFGIEGHKEPFVNIPEDTI 538
>AJ501281 similar to GP|4056425|gb|A ESTs gb|H36249 gb|AA59732 and
gb|AA651219 come from this gene. {Arabidopsis thaliana},
partial (30%)
Length = 675
Score = 88.6 bits (218), Expect = 1e-18
Identities = 38/74 (51%), Positives = 54/74 (72%)
Frame = +2
Query: 7 NIDEDDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFL 66
N D+ D+ +PP NF+MV++ I+RS P ++F F+++L LRS+I LCPEPYPE +FL
Sbjct: 206 NSDDADEAFVPPLNFAMVDNGIFRSGFPDSANFGFMKSLRLRSVICLCPEPYPEATAEFL 385
Query: 67 KEQNIRLFQFGIEG 80
IRL+QFGI+G
Sbjct: 386 NANGIRLYQFGIDG 427
>TC91383 similar to GP|9858782|gb|AAG01129.1| BAC19.14 {Lycopersicon
esculentum}, partial (11%)
Length = 819
Score = 32.0 bits (71), Expect = 0.16
Identities = 29/97 (29%), Positives = 43/97 (43%), Gaps = 4/97 (4%)
Frame = +2
Query: 17 PPPNFSMVEDCIYRSSLPKPSSFPFLQTL--NLRSIIYLCPEPYPEENLDFLKEQNIRLF 74
P P + ++ + P P+ P TL RS++ LC P PE+ Q +
Sbjct: 122 PQPKLTTTQNIVVNVPTPNPTPTPTTTTLCYEPRSVLELCRSPSPEKK----PSQQEQEH 289
Query: 75 QFGIEGKTEVSLPALR--DSIMEALKVLVDVRNHPIL 109
+E + SLP L DSIM+ L L D + PI+
Sbjct: 290 NPEVEQEDHASLPNLDWWDSIMKDLG-LQDDSSTPII 397
>TC90220 weakly similar to PIR|T01152|T01152 probable pectinesterase
[imported] - Arabidopsis thaliana, partial (27%)
Length = 737
Score = 29.3 bits (64), Expect = 1.1
Identities = 16/48 (33%), Positives = 26/48 (53%), Gaps = 1/48 (2%)
Frame = +1
Query: 70 NIRLFQ-FGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGK 116
NI LF+ F EG T V R + + +L+D+ H I+++C Q +
Sbjct: 574 NITLFRNFSYEGVTVVKNERKRVMFILVMSLLIDLMFH*IIIYCNQNE 717
>TC77194 homologue to GP|21593495|gb|AAM65462.1 glycine rich protein-like
{Arabidopsis thaliana}, partial (96%)
Length = 700
Score = 29.3 bits (64), Expect = 1.1
Identities = 18/51 (35%), Positives = 26/51 (50%), Gaps = 1/51 (1%)
Frame = +3
Query: 76 FGIEGKTEVSLPALRDSI-MEALKVLVDVRNHPILVHCKQGKHRTGCLVGC 125
F I+G + P + +I M L +L + HP+LV K G+ G LV C
Sbjct: 105 FIIDGT*SSNTPIINININMLPLSLLKTAQGHPMLVELKNGETYNGHLVNC 257
>TC83214 similar to SP|O49204|KAPS_CATRO Adenylylsulfate kinase chloroplast
precursor (EC 2.7.1.25) (APS kinase), partial (23%)
Length = 603
Score = 27.7 bits (60), Expect = 3.1
Identities = 14/66 (21%), Positives = 28/66 (42%)
Frame = +3
Query: 126 FRKLQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLRQCLYSIIYQYQGASKKRRL 185
++ LQ C F+ + + + F RF+ V LR+C S++ ++
Sbjct: 72 YKHLQPQCGGVTFQNIECGTSTVEKLGIVNFRHRFNAVELRRCRKSMLKPIMAQDERESS 251
Query: 186 MYQDEN 191
+ D+N
Sbjct: 252 IADDDN 269
>TC77193 homologue to GP|21593495|gb|AAM65462.1 glycine rich protein-like
{Arabidopsis thaliana}, partial (85%)
Length = 989
Score = 27.3 bits (59), Expect = 4.0
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = +2
Query: 94 MEALKVLVDVRNHPILVHCKQGKHRTGCLVGC 125
M L +L + HP+LV K G+ G LV C
Sbjct: 107 MLPLSLLKTAQGHPMLVELKNGETYNGHLVNC 202
>TC90386 weakly similar to GP|5882724|gb|AAD55277.1| F25A4.4 {Arabidopsis
thaliana}, partial (54%)
Length = 674
Score = 26.9 bits (58), Expect = 5.3
Identities = 11/33 (33%), Positives = 19/33 (57%)
Frame = -1
Query: 92 SIMEALKVLVDVRNHPILVHCKQGKHRTGCLVG 124
++ME K + ++RNH + H + G HR + G
Sbjct: 182 TVMEM*KNMAEMRNHHVAPHDRAGLHRNETIDG 84
>TC89202 weakly similar to SP|O48923|C7DA_SOYBN Cytochrome P450 71D10 (EC
1.14.-.-). [Soybean] {Glycine max}, partial (38%)
Length = 1040
Score = 26.6 bits (57), Expect = 6.9
Identities = 12/30 (40%), Positives = 16/30 (53%)
Frame = -1
Query: 2 IVEVENIDEDDDVLIPPPNFSMVEDCIYRS 31
I ++ I DDV +P PN S + D I S
Sbjct: 968 IADIAQITASDDVSVPAPNMSWMIDFILSS 879
>BQ150123 similar to GP|21109992|gb| TonB-dependent receptor {Xanthomonas
axonopodis pv. citri str. 306}, partial (2%)
Length = 1131
Score = 26.6 bits (57), Expect = 6.9
Identities = 15/52 (28%), Positives = 24/52 (45%), Gaps = 2/52 (3%)
Frame = -1
Query: 29 YRSSLPKPSSFPFLQTLNLRSIIYLCPE--PYPEENLDFLKEQNIRLFQFGI 78
+ S LP P S ++ RS+++ CP P P ++ FL R F +
Sbjct: 183 FASPLPAPRSLGAF*IVSFRSLLFFCPSFPPPPFPSVLFLPSLPFRFSVFSV 28
>AW684406 weakly similar to GP|21536774|gb| unknown {Arabidopsis thaliana},
partial (13%)
Length = 535
Score = 26.2 bits (56), Expect = 9.0
Identities = 19/56 (33%), Positives = 27/56 (47%), Gaps = 5/56 (8%)
Frame = -3
Query: 133 CLSSAFEEYQRFAGVKSRAAD-LTFIE--RFDLVSLRQCL--YSIIYQYQGASKKR 183
CLS A Y+R AG+ D + F++ + L CL YSI+ Q+ S R
Sbjct: 425 CLSYAIHSYRR*AGIGKITIDSMIFLQSWTYRLARSNICLSIYSILVQFNNRSNPR 258
>TC78053 similar to GP|5923675|gb|AAD56326.1| putative mRNA capping enzyme
RNA guanylyltransferase {Arabidopsis thaliana}, partial
(71%)
Length = 2017
Score = 26.2 bits (56), Expect = 9.0
Identities = 16/61 (26%), Positives = 31/61 (50%), Gaps = 3/61 (4%)
Frame = +1
Query: 66 LKEQNIRLFQFGIEGKTEV-SLPALRDSIMEALKVLVDVRNHP--ILVHCKQGKHRTGCL 122
LK++ I+ + +G+ V ++ + E ++ L + ILVHC G +RTG +
Sbjct: 703 LKKEGIKHLKIQCKGRDSVPENSSVNQFVYEVIQFLSRQKQSKKYILVHCTHGHNRTGYM 882
Query: 123 V 123
+
Sbjct: 883 I 885
>TC89640 similar to GP|16648927|gb|AAL24315.1 formyltetrahydrofolate
deformylase-like {Arabidopsis thaliana}, partial (83%)
Length = 1126
Score = 26.2 bits (56), Expect = 9.0
Identities = 14/33 (42%), Positives = 19/33 (57%)
Frame = +2
Query: 25 EDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEP 57
EDC+ S+PK S FP T+ + +I L P P
Sbjct: 98 EDCVL-VSVPKLSDFPIEITVTILLMIILLPVP 193
>BQ751461 weakly similar to GP|18376015|em conserved hypothetical protein
{Neurospora crassa}, partial (19%)
Length = 773
Score = 26.2 bits (56), Expect = 9.0
Identities = 7/13 (53%), Positives = 9/13 (68%)
Frame = -1
Query: 121 CLVGCFRKLQNWC 133
C+ GC+ KL WC
Sbjct: 44 CMAGCWNKLDRWC 6
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.141 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,867,305
Number of Sequences: 36976
Number of extensions: 109708
Number of successful extensions: 603
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 598
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 602
length of query: 200
length of database: 9,014,727
effective HSP length: 91
effective length of query: 109
effective length of database: 5,649,911
effective search space: 615840299
effective search space used: 615840299
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC148289.16


