

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148241.11 + phase: 0
(390 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
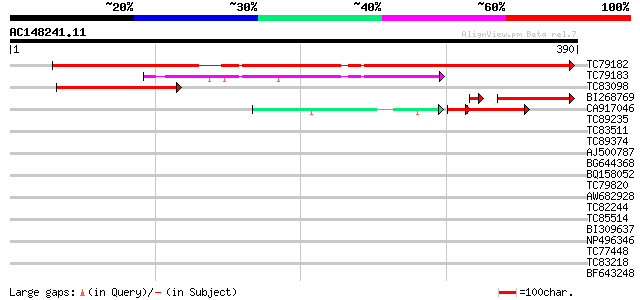
Sequences producing significant alignments: (bits) Value
TC79182 similar to GP|15028159|gb|AAK76703.1 unknown protein {Ar... 401 e-112
TC79183 similar to GP|15028159|gb|AAK76703.1 unknown protein {Ar... 142 3e-34
TC83098 weakly similar to PIR|T04894|T04894 hypothetical protein... 105 3e-23
BI268769 similar to GP|15028159|gb| unknown protein {Arabidopsis... 80 7e-17
CA917046 similar to PIR|T04894|T048 hypothetical protein F18F4.2... 57 9e-14
TC89235 similar to SP|P57758|CTNS_ARATH Cystinosin homolog. [Mou... 35 0.038
TC83511 similar to GP|18252961|gb|AAL62407.1 unknown protein {Ar... 35 0.050
TC89374 similar to PIR|T05879|T05879 hypothetical protein T29A15... 32 0.55
AJ500787 31 0.72
BG644368 similar to GP|12597768|g DNA ligase I putative {Arabid... 30 1.2
BQ158052 30 2.1
TC79820 similar to GP|21537325|gb|AAM61666.1 unknown {Arabidopsi... 30 2.1
AW682928 similar to GP|22507458|gb| Unknown (protein for MGC:303... 29 2.7
TC82244 similar to GP|22507458|gb|AAH19429.1 Unknown (protein fo... 29 2.7
TC85514 homologue to SP|P32869|PSAD_CUCSA Photosystem I reaction... 29 3.6
BI309637 similar to PIR|D86241|D86 protein T16B5.8 [imported] - ... 28 4.6
NP496346 NP496346|AJ418371.1|CAD10810.1 nod region linked recept... 28 6.1
TC77448 SYMRK; MtSYMRK [Medicago truncatula]; nod region linked ... 28 6.1
TC83218 28 7.9
BF643248 weakly similar to PIR|D96595|D96 probable acetyl-CoA sy... 28 7.9
>TC79182 similar to GP|15028159|gb|AAK76703.1 unknown protein {Arabidopsis
thaliana}, partial (61%)
Length = 1205
Score = 401 bits (1030), Expect = e-112
Identities = 217/360 (60%), Positives = 255/360 (70%), Gaps = 1/360 (0%)
Frame = +2
Query: 30 DDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLEPAT 89
D+ISF+ G +SL+ WGVAEIPQIIT FR KSSHG+S+ FLLTWVAGDI NLVGCLLEPAT
Sbjct: 2 DNISFTFGFISLICWGVAEIPQIITNFRAKSSHGVSIVFLLTWVAGDIFNLVGCLLEPAT 181
Query: 90 LPTQFYTALTVINCSFMQALQSYYYCKLYTMITSSDGANIVRMLRVNCYFYNCIILQDNE 149
LPTQ+YTAL + + +QS+YY +Y NI + E
Sbjct: 182 LPTQYYTALLYTITTIVLVVQSFYYDYIYKWCKRRQKINIE---------------ETYE 316
Query: 150 EEKRPLNPKPSQVYSGIAIPNGTQKEAARGEYYYMSARSLAGSATPPSFTHLRAAKSGPS 209
EEK+PL PK + GI I +G + + EYYY SARSLAG+ TPPS T++R AKSGPS
Sbjct: 317 EEKKPLKPK-ERFELGIPIRSGRHRAIPKPEYYYGSARSLAGNVTPPSRTYMRVAKSGPS 493
Query: 210 ALEFIHDSSDDDEASQVTSNISTTKPWSIPRSVDGRYGTFLATAINLPLKGNSMRYGYIG 269
A+ DSS DDEA V + T+P IPRS G YGTFLA +INLP + N+++ GYI
Sbjct: 494 AMGLNEDSSSDDEAHSVPA----TQPRQIPRSA-GSYGTFLAASINLPHQSNALKVGYIA 658
Query: 270 FTGIKLLKVYVVVHSTYGQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIA 329
+G KLL V HS GQ+LGW+MAAIYT RIPQIWLNIKRGSVEGLNPFMF+FALIA
Sbjct: 659 LSGRKLLSQEHVTHSALGQWLGWLMAAIYTGGRIPQIWLNIKRGSVEGLNPFMFIFALIA 838
Query: 330 NTSYVGSILVRTTEFESIKANLPWLLDATVCVALDFFIISQYIYYRYFR-SSESSDDGEY 388
N +YVGSILVRTTE+ESIKAN+PWLLDA VCVALD FII QYI YRY R ++ SSD G Y
Sbjct: 839 NATYVGSILVRTTEWESIKANMPWLLDAIVCVALDLFIILQYINYRYHRKTTTSSDYGNY 1018
>TC79183 similar to GP|15028159|gb|AAK76703.1 unknown protein {Arabidopsis
thaliana}, partial (23%)
Length = 931
Score = 142 bits (357), Expect = 3e-34
Identities = 98/253 (38%), Positives = 127/253 (49%), Gaps = 46/253 (18%)
Frame = +2
Query: 93 QFYTALTVINCSFMQALQSYYYCKLYTMITSSDGANIVRMLRVN-----------CYFYN 141
Q YT T++ +QS+YY +Y NI +L +N + +
Sbjct: 140 QLYTITTIV-----LVVQSFYYDYIYKWCKRRQKINIEEVLSLNQI*EEGKYNDTLFLLS 304
Query: 142 CIILQ-----DNEEEKRPLNPKPSQVYSGIAIPNGTQKEAARGEYYY------------- 183
II+ EEEK+PL PK + GI I +G + + EYYY
Sbjct: 305 YIIIVML***TYEEEKKPLKPK-ERFELGIPIRSGRHRAIPKPEYYYG*VLIFN*IVN** 481
Query: 184 -----------------MSARSLAGSATPPSFTHLRAAKSGPSALEFIHDSSDDDEASQV 226
SARSLAG+ TPPS T++R AKSGPSA+ DSS DDEA V
Sbjct: 482 IKQSSELMSFELID*VNRSARSLAGNVTPPSRTYMRVAKSGPSAMGLNEDSSSDDEAHSV 661
Query: 227 TSNISTTKPWSIPRSVDGRYGTFLATAINLPLKGNSMRYGYIGFTGIKLLKVYVVVHSTY 286
+ T+P IPRS G YGTFLA +INLP + N+++ GYI +G KLL V HS
Sbjct: 662 PA----TQPRQIPRSA-GSYGTFLAASINLPHQSNALKVGYIALSGRKLLSQEHVTHSAL 826
Query: 287 GQYLGWIMAAIYT 299
GQ+LGW+MAAIYT
Sbjct: 827 GQWLGWLMAAIYT 865
>TC83098 weakly similar to PIR|T04894|T04894 hypothetical protein F18F4.200
- Arabidopsis thaliana, partial (29%)
Length = 467
Score = 105 bits (262), Expect = 3e-23
Identities = 48/86 (55%), Positives = 64/86 (73%)
Frame = +2
Query: 33 SFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLEPATLPT 92
S +LG++S++ W +AEIPQ+IT +R KSSHG+S+ FLLTW+ GD+ NL GCLLEPATLPT
Sbjct: 170 SITLGVISVIVWMIAEIPQLITNYREKSSHGLSVTFLLTWIIGDLFNLFGCLLEPATLPT 349
Query: 93 QFYTALTVINCSFMQALQSYYYCKLY 118
Q YTA+ + LQ+ YY +Y
Sbjct: 350 QLYTAVLYTLITLTLCLQATYYGHIY 427
>BI268769 similar to GP|15028159|gb| unknown protein {Arabidopsis thaliana},
partial (13%)
Length = 554
Score = 79.7 bits (195), Expect(2) = 7e-17
Identities = 41/54 (75%), Positives = 45/54 (82%), Gaps = 1/54 (1%)
Frame = +3
Query: 336 SILVRTTEFESIKANLPWLLDATVCVALDFFIISQYIYYRYFR-SSESSDDGEY 388
SILVRTTE+ESIKAN+PWLLDA VCVALD FII QYI YRY R ++ SSD G Y
Sbjct: 222 SILVRTTEWESIKANMPWLLDAIVCVALDLFIILQYINYRYHRKTTTSSDYGNY 383
Score = 25.0 bits (53), Expect(2) = 7e-17
Identities = 9/10 (90%), Positives = 10/10 (100%)
Frame = +2
Query: 317 GLNPFMFVFA 326
GLNPFMF+FA
Sbjct: 62 GLNPFMFIFA 91
>CA917046 similar to PIR|T04894|T048 hypothetical protein F18F4.200 -
Arabidopsis thaliana, partial (19%)
Length = 663
Score = 56.6 bits (135), Expect(3) = 9e-14
Identities = 24/42 (57%), Positives = 33/42 (78%)
Frame = +1
Query: 316 EGLNPFMFVFALIANTSYVGSILVRTTEFESIKANLPWLLDA 357
EG+NP MF+FALI NT+YV SILV + ++ + NLPWL+D+
Sbjct: 538 EGVNPLMFLFALIGNTTYVASILVSSMDWSKLGPNLPWLVDS 663
Score = 33.1 bits (74), Expect(3) = 9e-14
Identities = 40/150 (26%), Positives = 55/150 (36%), Gaps = 19/150 (12%)
Frame = +2
Query: 168 IPNGTQKEAARGEYYYMSARSLAGSATPPSFTHLRAAKS-----------GPSALEFIHD 216
IP QK + YY SAR L+ S TP S R S PS
Sbjct: 65 IPFPAQKSHVETQSYYQSARYLSKSHTPKSELAQRMPSSLILDPIEEPLLVPSVFTKSAP 244
Query: 217 SSDDDEASQVTSNISTTKPWSIPRSVDGRYGTFLATAINLPLKGNSMRYGYIGFTGIKLL 276
S + S ++ ++ S D R + +A R ++ + G KLL
Sbjct: 245 SLKIKNTLCLVSTLTFLGALNLLHSPDTRIHSDVAKP----------RKEFVIYVGRKLL 394
Query: 277 KVY--------VVVHSTYGQYLGWIMAAIY 298
+V V + + G YLGW MA IY
Sbjct: 395 QVSGHKLSDQGVEAYHSIGTYLGWAMAVIY 484
Score = 23.9 bits (50), Expect(3) = 9e-14
Identities = 10/16 (62%), Positives = 13/16 (80%)
Frame = +3
Query: 302 RIPQIWLNIKRGSVEG 317
R+PQI LNI+RG+ G
Sbjct: 495 RLPQICLNIRRGNF*G 542
>TC89235 similar to SP|P57758|CTNS_ARATH Cystinosin homolog. [Mouse-ear
cress] {Arabidopsis thaliana}, partial (92%)
Length = 1103
Score = 35.4 bits (80), Expect = 0.038
Identities = 20/55 (36%), Positives = 31/55 (56%)
Frame = +1
Query: 280 VVVHSTYGQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYV 334
V + TY + LGW +++ S PQ+ LN +R SV GLN + L ++SY+
Sbjct: 40 VSLEVTY-EVLGWFAFIVWSISFYPQVILNFRRKSVVGLNFDFVLLNLTKHSSYL 201
Score = 31.2 bits (69), Expect = 0.72
Identities = 13/35 (37%), Positives = 21/35 (59%)
Frame = +1
Query: 36 LGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLL 70
LG + + W ++ PQ+I FR KS G++ F+L
Sbjct: 67 LGWFAFIVWSISFYPQVILNFRRKSVVGLNFDFVL 171
>TC83511 similar to GP|18252961|gb|AAL62407.1 unknown protein {Arabidopsis
thaliana}, complete
Length = 1014
Score = 35.0 bits (79), Expect = 0.050
Identities = 27/82 (32%), Positives = 41/82 (49%), Gaps = 2/82 (2%)
Frame = +1
Query: 296 AIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESIKANLP--W 353
AI+ C+RIPQI+ N S L + + + G +VR F +I+ N P
Sbjct: 505 AIFLCARIPQIFQNFSNKSTGEL-------SFLTSFMNFGGSMVRV--FTTIQENAPKSV 657
Query: 354 LLDATVCVALDFFIISQYIYYR 375
LL + VA +F I+SQ + Y+
Sbjct: 658 LLGYGIGVATNFTILSQIVIYQ 723
>TC89374 similar to PIR|T05879|T05879 hypothetical protein T29A15.230 -
Arabidopsis thaliana, partial (71%)
Length = 645
Score = 31.6 bits (70), Expect = 0.55
Identities = 12/25 (48%), Positives = 17/25 (68%)
Frame = -2
Query: 80 LVGCLLEPATLPTQFYTALTVINCS 104
L+GCL+ LPTQ +T +T +N S
Sbjct: 386 LLGCLIRSFVLPTQHFTTITAVNIS 312
>AJ500787
Length = 566
Score = 31.2 bits (69), Expect = 0.72
Identities = 16/42 (38%), Positives = 21/42 (49%)
Frame = +3
Query: 35 SLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGD 76
+LG +SL +PQ F+ S G S +LTWV GD
Sbjct: 210 TLGYLSLGIESTVPMPQAYQNFKRHSVSGFSKWIILTWVGGD 335
>BG644368 similar to GP|12597768|g DNA ligase I putative {Arabidopsis
thaliana}, partial (12%)
Length = 641
Score = 30.4 bits (67), Expect = 1.2
Identities = 13/25 (52%), Positives = 16/25 (64%)
Frame = +3
Query: 298 YTCSRIPQIWLNIKRGSVEGLNPFM 322
YT S+ WL +KR VEGLN F+
Sbjct: 567 YTPSKRSDAWLKVKRDYVEGLNDFL 641
>BQ158052
Length = 983
Score = 29.6 bits (65), Expect = 2.1
Identities = 20/84 (23%), Positives = 33/84 (38%)
Frame = +2
Query: 59 KSSHGISLAFLLTWVAGDICNLVGCLLEPATLPTQFYTALTVINCSFMQALQSYYYCKLY 118
K+SH S FLL WV C P F + +N + + + + Y + +
Sbjct: 11 KNSHSGSFLFLLVWV---------CCTGPDMYLILFPRKVPYVNSTMITCMPTSYQNRTF 163
Query: 119 TMITSSDGANIVRMLRVNCYFYNC 142
IVR + +N + +NC
Sbjct: 164 VF--------IVRFILINLFCFNC 211
>TC79820 similar to GP|21537325|gb|AAM61666.1 unknown {Arabidopsis
thaliana}, partial (89%)
Length = 1225
Score = 29.6 bits (65), Expect = 2.1
Identities = 18/56 (32%), Positives = 28/56 (49%), Gaps = 9/56 (16%)
Frame = +3
Query: 80 LVGCLLEPATLPTQFYTALTVIN-----CSFMQALQSYYYCK----LYTMITSSDG 126
L+G LLEP P +F + ++N C F+ A+ YY + LYT ++ G
Sbjct: 375 LIGKLLEPVWGPREFIKFIFIVNILTSLCIFITAIALYYITRQEIYLYTPLSGFHG 542
>AW682928 similar to GP|22507458|gb| Unknown (protein for MGC:30371) {Mus
musculus}, partial (13%)
Length = 641
Score = 29.3 bits (64), Expect = 2.7
Identities = 24/68 (35%), Positives = 31/68 (45%), Gaps = 5/68 (7%)
Frame = +2
Query: 200 HLRAAKSGPSALEFIHDSSDDDEASQ-VTSNISTTKPWS----IPRSVDGRYGTFLATAI 254
H R +A I SSD D A V S ISTT PW+ + T +A+A
Sbjct: 125 HYRRTAEAAAAASSIRYSSDFDIAPPGVASRISTTNPWANAYPSAAAAVAAAATNVASAA 304
Query: 255 NLPLKGNS 262
+L LK +S
Sbjct: 305 SLDLKRSS 328
>TC82244 similar to GP|22507458|gb|AAH19429.1 Unknown (protein for
MGC:30371) {Mus musculus}, partial (13%)
Length = 847
Score = 29.3 bits (64), Expect = 2.7
Identities = 24/68 (35%), Positives = 31/68 (45%), Gaps = 5/68 (7%)
Frame = +3
Query: 200 HLRAAKSGPSALEFIHDSSDDDEASQ-VTSNISTTKPWS----IPRSVDGRYGTFLATAI 254
H R +A I SSD D A V S ISTT PW+ + T +A+A
Sbjct: 360 HYRRTAEAAAAASSIRYSSDFDIAPPGVASRISTTNPWANAFPSAAAAVAAAATNVASAA 539
Query: 255 NLPLKGNS 262
+L LK +S
Sbjct: 540 SLDLKRSS 563
>TC85514 homologue to SP|P32869|PSAD_CUCSA Photosystem I reaction center
subunit II chloroplast precursor (Photosystem I 20 kDa
subunit), partial (77%)
Length = 870
Score = 28.9 bits (63), Expect = 3.6
Identities = 14/46 (30%), Positives = 24/46 (51%)
Frame = +3
Query: 197 SFTHLRAAKSGPSALEFIHDSSDDDEASQVTSNISTTKPWSIPRSV 242
+ TH S S+ FI++ + +AS T +S+ KPW P ++
Sbjct: 15 TLTHTNKHHSSSSSSTFINNMAMATQASLFTPPLSSPKPWKQPSTL 152
>BI309637 similar to PIR|D86241|D86 protein T16B5.8 [imported] - Arabidopsis
thaliana, partial (28%)
Length = 714
Score = 28.5 bits (62), Expect = 4.6
Identities = 20/69 (28%), Positives = 28/69 (39%)
Frame = +2
Query: 181 YYYMSARSLAGSATPPSFTHLRAAKSGPSALEFIHDSSDDDEASQVTSNISTTKPWSIPR 240
Y S SLAG PS +HL E IH+S + N+ T K W +
Sbjct: 311 YGLASWLSLAG----PSLSHLELRMDNLGDNEIIHESPSKLDCIGAAVNVETLKLWGVLI 478
Query: 241 SVDGRYGTF 249
+ ++ TF
Sbjct: 479 KLIPKWETF 505
>NP496346 NP496346|AJ418371.1|CAD10810.1 nod region linked receptor kinase
[Medicago truncatula]
Length = 2706
Score = 28.1 bits (61), Expect = 6.1
Identities = 15/41 (36%), Positives = 22/41 (53%)
Frame = +1
Query: 182 YYMSARSLAGSATPPSFTHLRAAKSGPSALEFIHDSSDDDE 222
+ +S L S TPP L+ A + P LEF+HD + D+
Sbjct: 682 FNVSNVDLKDSVTPP-LQVLQTALTHPERLEFVHDGLETDD 801
>TC77448 SYMRK; MtSYMRK [Medicago truncatula]; nod region linked receptor
kinase [Medicago truncatula]
Length = 3568
Score = 28.1 bits (61), Expect = 6.1
Identities = 15/41 (36%), Positives = 22/41 (53%)
Frame = +1
Query: 182 YYMSARSLAGSATPPSFTHLRAAKSGPSALEFIHDSSDDDE 222
+ +S L S TPP L+ A + P LEF+HD + D+
Sbjct: 1057 FNVSNVDLKDSVTPP-LQVLQTALTHPERLEFVHDGLETDD 1176
>TC83218
Length = 1170
Score = 27.7 bits (60), Expect = 7.9
Identities = 18/54 (33%), Positives = 24/54 (44%)
Frame = +2
Query: 60 SSHGISLAFLLTWVAGDICNLVGCLLEPATLPTQFYTALTVINCSFMQALQSYY 113
S H FLLT C L C+ E +TLP + L+ + SF+ A Y
Sbjct: 512 SQHRTKRCFLLT*HLEAACLLRSCVFEASTLPYNSISDLSWSSRSFLPAYAPAY 673
>BF643248 weakly similar to PIR|D96595|D96 probable acetyl-CoA synthetase
45051-31547 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 690
Score = 27.7 bits (60), Expect = 7.9
Identities = 19/72 (26%), Positives = 36/72 (49%), Gaps = 1/72 (1%)
Frame = +2
Query: 31 DISFSLGLMSLVSWGVAEIPQ-IITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLEPAT 89
D+ +SL L L S E P+ I+ + S HG W++G + N+ C L+P++
Sbjct: 446 DVYWSLVLKEL-SISFIEPPKCILDTSSDPSKHGGK------WLSGSVLNIADCCLQPSS 604
Query: 90 LPTQFYTALTVI 101
P + ++ ++
Sbjct: 605 HPNKPDDSIAIV 640
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.137 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,058,458
Number of Sequences: 36976
Number of extensions: 217323
Number of successful extensions: 1143
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 1124
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1137
length of query: 390
length of database: 9,014,727
effective HSP length: 98
effective length of query: 292
effective length of database: 5,391,079
effective search space: 1574195068
effective search space used: 1574195068
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC148241.11


