

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148241.10 - phase: 1 /pseudo
(611 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
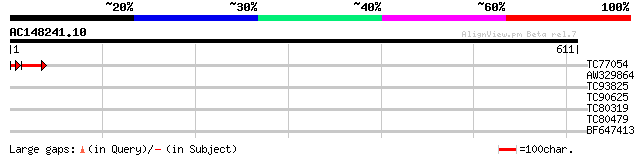
TC77054 similar to GP|5882739|gb|AAD55292.1| Contains PF|00249 M... 47 8e-06
AW329864 39 0.004
TC93825 similar to GP|23498747|emb|CAD50817. hypothetical protei... 39 0.004
TC90625 weakly similar to GP|10177856|dbj|BAB11208. phloem-speci... 34 0.14
TC80319 similar to GP|20259261|gb|AAM14366.1 putative HhoA prote... 33 0.42
TC80479 homologue to PIR|S12720|HHPM17 heat shock protein 17.7 -... 31 1.2
BF647413 similar to GP|13877625|gb Unknown protein {Arabidopsis ... 30 2.7
>TC77054 similar to GP|5882739|gb|AAD55292.1| Contains PF|00249 Myb-like
DNA-binding domain. EST gb|Z18152 comes from this
gene., partial (11%)
Length = 643
Score = 47.0 bits (110), Expect(2) = 8e-06
Identities = 25/27 (92%), Positives = 26/27 (95%)
Frame = +1
Query: 13 GR*YSKR*SYNFFIRT*TCS*ESKN*R 39
GR*YSKR*SYNFFIRT*T S*ESKN*+
Sbjct: 289 GR*YSKR*SYNFFIRT*TSS*ESKN*K 369
Score = 20.8 bits (42), Expect(2) = 8e-06
Identities = 9/11 (81%), Positives = 9/11 (81%)
Frame = +2
Query: 1 EVSKN*FYFQE 11
E SKN FYFQE
Sbjct: 257 EDSKNRFYFQE 289
>AW329864
Length = 401
Score = 39.3 bits (90), Expect = 0.004
Identities = 21/27 (77%), Positives = 22/27 (80%)
Frame = +2
Query: 1 EVSKN*FYFQEEGR*YSKR*SYNFFIR 27
E SKN FYFQ+EG *Y KR*S NFFIR
Sbjct: 200 EDSKNRFYFQKEGC*Y*KR*SCNFFIR 280
>TC93825 similar to GP|23498747|emb|CAD50817. hypothetical protein
{Plasmodium falciparum 3D7}, partial (0%)
Length = 948
Score = 39.3 bits (90), Expect = 0.004
Identities = 21/27 (77%), Positives = 22/27 (80%)
Frame = -1
Query: 1 EVSKN*FYFQEEGR*YSKR*SYNFFIR 27
E SKN FYFQ+EG *Y KR*S NFFIR
Sbjct: 561 EDSKNRFYFQKEGC*Y*KR*SCNFFIR 481
>TC90625 weakly similar to GP|10177856|dbj|BAB11208. phloem-specific
lectin-like protein {Arabidopsis thaliana}, partial (9%)
Length = 892
Score = 34.3 bits (77), Expect = 0.14
Identities = 14/26 (53%), Positives = 20/26 (76%)
Frame = +3
Query: 428 KEIENLLETGEIVTGKGKNQVGTVKR 453
+E+E LLE EI++GKG Q+G +KR
Sbjct: 69 EEVEYLLEIDEIISGKGAKQIGALKR 146
>TC80319 similar to GP|20259261|gb|AAM14366.1 putative HhoA protease
precursor {Arabidopsis thaliana}, partial (47%)
Length = 1195
Score = 32.7 bits (73), Expect = 0.42
Identities = 19/53 (35%), Positives = 30/53 (55%), Gaps = 1/53 (1%)
Frame = -2
Query: 73 STKTNGCNTKSLSKMGSLSIKLRKLSH-VRYRESTKAISTQLV*LVFFVARIF 124
S+ + C ++S S+ G +KLRKLSH R K ++ +LV + V R+F
Sbjct: 297 SSSCSCCESESESESGCAELKLRKLSHEATTRLELKTMALRLVAVTVVVGRVF 139
>TC80479 homologue to PIR|S12720|HHPM17 heat shock protein 17.7 - garden
pea, partial (96%)
Length = 764
Score = 31.2 bits (69), Expect = 1.2
Identities = 14/39 (35%), Positives = 22/39 (55%)
Frame = +2
Query: 52 SGRH*NFFRKGSRKTYSNLEISTKTNGCNTKSLSKMGSL 90
+G H R+ S++T +NL K+NGCNT ++ L
Sbjct: 179 NGPHRRHNREESKRTNTNLRSRRKSNGCNTSRRERVSKL 295
>BF647413 similar to GP|13877625|gb Unknown protein {Arabidopsis thaliana},
partial (41%)
Length = 667
Score = 30.0 bits (66), Expect = 2.7
Identities = 28/112 (25%), Positives = 45/112 (40%), Gaps = 6/112 (5%)
Frame = +1
Query: 402 FEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETG------EIVTGKGKNQVGTVKRAG 455
F LIF+ + ++T + + + KEI N + + +GKGK V
Sbjct: 115 FTSLIFISHFASAATLKLNTQEVKALKEIGNKIGKKDWDFGVDPCSGKGKWNV------S 276
Query: 456 DTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFD 507
D+R G IC+ + ++C V+ I K + S HLK D
Sbjct: 277 DSRKGFESAVICNCSFNHNSSCHVVSIFLKAQNLSGTLSPEFSKLPHLKILD 432
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.359 0.157 0.535
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,745,859
Number of Sequences: 36976
Number of extensions: 235705
Number of successful extensions: 2490
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 1037
Number of HSP's successfully gapped in prelim test: 133
Number of HSP's that attempted gapping in prelim test: 1443
Number of HSP's gapped (non-prelim): 1205
length of query: 611
length of database: 9,014,727
effective HSP length: 102
effective length of query: 509
effective length of database: 5,243,175
effective search space: 2668776075
effective search space used: 2668776075
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148241.10


