

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148240.10 - phase: 0 /pseudo
(1012 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
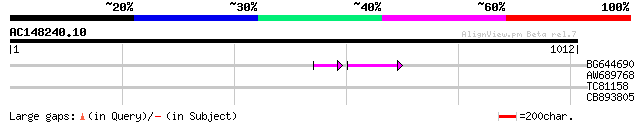
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 47 1e-09
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 42 0.002
TC81158 similar to GP|9280664|gb|AAF86533.1| F21B7.15 {Arabidops... 33 0.42
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 32 0.94
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 46.6 bits (109), Expect(2) = 1e-09
Identities = 32/99 (32%), Positives = 48/99 (48%)
Frame = -3
Query: 603 ETKPDLLLKVTVNKKALITLKHLLQLQDWKQSGYFYPMQLTMA*YCIKWMSKVLFLMVSL 662
E P K T+ KK ++ L +WK + + + C KWM +V LM
Sbjct: 417 EINPSWWCKDTIKKKE*TMMRLFHLLPEWKLLEF**LLLHSWGSSCTKWM*RVHLLMEIS 238
Query: 663 KKKYMSNNLLGLRILSILTMFINLRNHYMA*NKLPELGM 701
K++ +S+NLL L++ M + HYM *+KL E GM
Sbjct: 237 KRRCLSSNLLDLKMQRYQIMCSD*IRHYMV*SKLQEHGM 121
Score = 35.4 bits (80), Expect(2) = 1e-09
Identities = 22/53 (41%), Positives = 30/53 (56%)
Frame = -1
Query: 542 LSLKLLKKLSQMMVGY*PCKKN*ISFREMMCGI*YPNLFRRTLLEQNGYSETS 594
LS ++LKK M G CKKN IS +E+ G + +L + LE G+ ETS
Sbjct: 590 LSPRMLKKHCVMQTGSILCKKNSISLKEVRYGTWFLDLKAKQ*LELGGFLETS 432
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 41.6 bits (96), Expect = 0.002
Identities = 43/129 (33%), Positives = 62/129 (47%)
Frame = +2
Query: 573 GI*YPNLFRRTLLEQNGYSETS*MKKEK*PETKPDLLLKVTVNKKALITLKHLLQLQDWK 632
GI + L R LL +G+ E + + KPD L+V+V +ITLK L +
Sbjct: 149 GILFLFLHTRKLLAASGFIELKKTQMDLLTNLKPD*WLRVSVKHLDVITLKLSLL**NLS 328
Query: 633 QSGYFYPMQLTMA*YCIKWMSKVLFLMVSLKKKYMSNNLLGLRILSILTMFINLRNHYMA 692
SG F P+ + K +S + F MV K+K + NL L++L I + NH MA
Sbjct: 329 LSGSFSPLLSLINGKFNKLISTMPF*MVFFKRKCICLNLRDLKLL-INLWCAS*TNHSMA 505
Query: 693 *NKLPELGM 701
*+K GM
Sbjct: 506 *SKHHVHGM 532
>TC81158 similar to GP|9280664|gb|AAF86533.1| F21B7.15 {Arabidopsis
thaliana}, partial (6%)
Length = 890
Score = 33.5 bits (75), Expect = 0.42
Identities = 21/96 (21%), Positives = 52/96 (53%), Gaps = 11/96 (11%)
Frame = +2
Query: 419 LMIESLEIKLQSKVKVMQVQL-ILKMHQNLINLLILKSIQKLNLVQK----------LKS 467
+ I+ +E+K++ KVKV+ + L +L +H+ + L ++K+ + + ++K L
Sbjct: 401 MRIKMMEVKVKVKVKVLSLNLRVLLLHRRRLVLTVMKTTKTMQKMKKKGR*VMMRRWLLG 580
Query: 468 LQKLSLIQKQKLVQ*NRMKMLLRMFKITLSRLFSQN 503
++ + ++ K++ *+++ +R + S LF N
Sbjct: 581 VRLMVILMMMKML**DQLSQRMRFSLVYSSVLFCSN 688
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 32.3 bits (72), Expect = 0.94
Identities = 58/220 (26%), Positives = 91/220 (41%), Gaps = 16/220 (7%)
Frame = +1
Query: 585 LEQNGYSETS*MKKEK*PETKPDLLLKVTVNKKALITLKHLLQLQDWKQSGY---FYPMQ 641
L+ NG + S*M+ E+ K DL + T+N L T K L Q D +S + P
Sbjct: 166 LDSNGSLKQS*MRMERLRSIKLDLWPRGTLNNMVLTTPKFLHQWLDGTRSEW*LLLLPKL 345
Query: 642 LTMA*YCIKWMSKVLFLMVSLKKKYMSNNLLGL-------RILSILTMFINLRN-HYMA* 693
M C+ K L+ ++ + GL L L M +N + H A
Sbjct: 346 KGME--CVSARCKKRILVWRIE*GSFC*STTGLCEEG**AEGLKGLCMALNKHHEHGTAA 519
Query: 694 NKLPELGMID*VIS*LKMILKEDKLTQHSSERHL-----RRTF*LCKYMLMI*YLVLLMH 748
+KL + ++D + H + L F L +MLMI*YL ++M
Sbjct: 520 SKL--------------ISPRKDSRSVHMNTPCL*S*VKEVKFSLLVFMLMI*YLSVMMR 657
Query: 749 LFAKNSLS*CRMNLK*V*WEN*SSFLEFKSTKVKKEYMFI 788
+ K+ S + NL FLE K +++KE++F+
Sbjct: 658 ICLKSLKSP*KRNLTCQTLARCIIFLELK*LRMRKEFIFV 777
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.359 0.160 0.523
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,157,925
Number of Sequences: 36976
Number of extensions: 378717
Number of successful extensions: 4129
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 1324
Number of HSP's successfully gapped in prelim test: 195
Number of HSP's that attempted gapping in prelim test: 2765
Number of HSP's gapped (non-prelim): 1617
length of query: 1012
length of database: 9,014,727
effective HSP length: 106
effective length of query: 906
effective length of database: 5,095,271
effective search space: 4616315526
effective search space used: 4616315526
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC148240.10


