

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148236.3 + phase: 0 /pseudo
(116 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
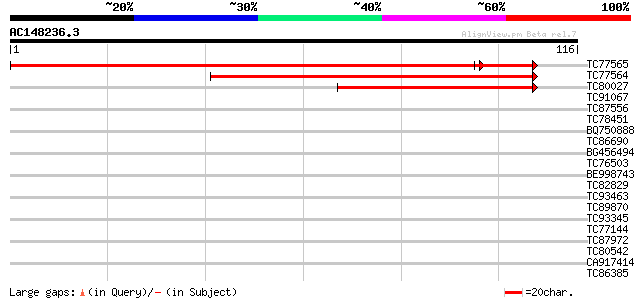
TC77565 similar to GP|21592665|gb|AAM64614.1 Isp4-like protein {... 172 2e-46
TC77564 similar to GP|20466754|gb|AAM20694.1 Isp4-like protein {... 127 6e-31
TC80027 similar to GP|15451020|gb|AAK96781.1 Unknown protein {Ar... 64 7e-12
TC91067 similar to PIR|H96681|H96681 protein F1E22.10 [imported]... 35 0.004
TC87556 similar to GP|14517530|gb|AAK62655.1 At1g48370/F11A17_27... 34 0.009
TC78451 similar to GP|9279625|dbj|BAB01083.1 gb|AAF23830.1~gene_... 30 0.10
BQ750888 GP|13992574|em glutenin HMW subunit 1Ax {Triticum timop... 28 0.40
TC86690 similar to GP|6041851|gb|AAF02160.1| unknown protein {Ar... 27 1.5
BG456494 homologue to GP|2645999|gb| chlorophyll a/b binding pro... 27 1.5
TC76503 homologue to GP|506629|gb|AAA50172.1|| photosystem II ty... 27 1.5
BE998743 similar to GP|22128591|gb| fertility restorer-like prot... 27 1.5
TC82829 weakly similar to GP|14334592|gb|AAK59474.1 unknown prot... 26 2.0
TC93463 similar to GP|15145367|gb|AAB54239.2 Hypothetical protei... 26 2.6
TC89870 similar to PIR|A86387|A86387 probable cytochrome b-561 [... 25 3.4
TC93345 similar to SP|Q9SJK3|AGOL_ARATH Argonaute-like protein A... 25 3.4
TC77144 similar to GP|20160594|dbj|BAB89541. contains ESTs AU031... 25 3.4
TC87972 weakly similar to PIR|F96751|F96751 unknown protein F28P... 25 4.4
TC80542 similar to GP|18087595|gb|AAL58928.1 unknown protein {Ar... 24 7.5
CA917414 similar to GP|6648966|gb| seed maturation protein PM25 ... 24 7.5
TC86385 similar to SP|P25469|H2A_LYCES Histone H2A. [Tomato] {Ly... 24 9.8
>TC77565 similar to GP|21592665|gb|AAM64614.1 Isp4-like protein {Arabidopsis
thaliana}, partial (71%)
Length = 1698
Score = 172 bits (437), Expect(2) = 2e-46
Identities = 76/97 (78%), Positives = 89/97 (91%)
Frame = +1
Query: 1 MPWWALIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGY 60
MPWW L+FA LA FTLP++IITATTNQTPGLN+ITEY+ G+I PGRPIANVCFKTYGY
Sbjct: 1282 MPWWGLLFAGALAFAFTLPISIITATTNQTPGLNIITEYVFGLIYPGRPIANVCFKTYGY 1461
Query: 61 MSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIA 97
+SM+QA+SFLSDFKLGHYMKIPPRSMF+VQ +GT++A
Sbjct: 1462 ISMAQAVSFLSDFKLGHYMKIPPRSMFLVQFIGTMLA 1572
Score = 27.3 bits (59), Expect(2) = 2e-46
Identities = 8/13 (61%), Positives = 12/13 (91%)
Frame = +2
Query: 96 IAGTVDVGVAWWL 108
+ GT+++GVAWWL
Sbjct: 1568 LPGTINIGVAWWL 1606
>TC77564 similar to GP|20466754|gb|AAM20694.1 Isp4-like protein {Arabidopsis
thaliana}, partial (37%)
Length = 1275
Score = 127 bits (319), Expect = 6e-31
Identities = 54/67 (80%), Positives = 65/67 (96%)
Frame = +1
Query: 42 GVILPGRPIANVCFKTYGYMSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVD 101
G+I PGRPIANVCFKTYGY+SM+QA+SFLSDFKLGHYMKIPPRSMF+VQ +GT++AGT++
Sbjct: 79 GLIYPGRPIANVCFKTYGYISMAQAVSFLSDFKLGHYMKIPPRSMFLVQFIGTMLAGTIN 258
Query: 102 VGVAWWL 108
+GVAWWL
Sbjct: 259 IGVAWWL 279
>TC80027 similar to GP|15451020|gb|AAK96781.1 Unknown protein {Arabidopsis
thaliana}, partial (30%)
Length = 926
Score = 64.3 bits (155), Expect = 7e-12
Identities = 25/41 (60%), Positives = 34/41 (81%)
Frame = +1
Query: 68 SFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWL 108
+FL + KLGHYMKIPPR M+ Q++GTL+AG V++ VAWW+
Sbjct: 1 AFLQNLKLGHYMKIPPRCMYTAQLVGTLVAGVVNLSVAWWM 123
>TC91067 similar to PIR|H96681|H96681 protein F1E22.10 [imported] -
Arabidopsis thaliana, partial (46%)
Length = 1223
Score = 35.0 bits (79), Expect = 0.004
Identities = 15/44 (34%), Positives = 24/44 (54%)
Frame = +3
Query: 63 MSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAW 106
+S A + DFK G+ P+SMF+ Q++GT + + V W
Sbjct: 540 VSTASDLMQDFKTGYMTLASPKSMFVSQVIGTAMGCIISPCVFW 671
>TC87556 similar to GP|14517530|gb|AAK62655.1 At1g48370/F11A17_27
{Arabidopsis thaliana}, partial (48%)
Length = 1309
Score = 33.9 bits (76), Expect = 0.009
Identities = 15/44 (34%), Positives = 24/44 (54%)
Frame = +1
Query: 63 MSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAW 106
++ A + DFK G+ PRS+F+ QI+GT + + V W
Sbjct: 517 VATASDLMQDFKTGYLTLASPRSVFVSQIIGTTMGCIISPCVFW 648
>TC78451 similar to GP|9279625|dbj|BAB01083.1
gb|AAF23830.1~gene_id:MOJ10.9~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (88%)
Length = 2452
Score = 30.4 bits (67), Expect = 0.10
Identities = 17/62 (27%), Positives = 30/62 (47%)
Frame = +2
Query: 47 GRPIANVCFKTYGYMSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAW 106
G IA V ++ A + DFK G+ +SMF+ Q++GT + G + + +
Sbjct: 1568 GGVIAGVASCAVMMSIVATAADLMQDFKTGYLTLSSAKSMFVSQLIGTAM-GCIIAPLTF 1744
Query: 107 WL 108
W+
Sbjct: 1745 WM 1750
>BQ750888 GP|13992574|em glutenin HMW subunit 1Ax {Triticum timopheevii},
partial (1%)
Length = 721
Score = 28.5 bits (62), Expect = 0.40
Identities = 12/39 (30%), Positives = 22/39 (55%)
Frame = +2
Query: 66 AISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGV 104
A + +SDF++G ++ PP F Q +G ++ + GV
Sbjct: 479 ATALVSDFRVGFLLRTPPHLQFYAQAIGGAVSIFLAPGV 595
>TC86690 similar to GP|6041851|gb|AAF02160.1| unknown protein {Arabidopsis
thaliana}, partial (63%)
Length = 1592
Score = 26.6 bits (57), Expect = 1.5
Identities = 12/40 (30%), Positives = 21/40 (52%)
Frame = -2
Query: 7 IFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILP 46
+ +GLA +F L A+++A +PG V ++LP
Sbjct: 430 LVGTGLARVFLLTAALLSAAVTISPGFAVPEASTTSLVLP 311
>BG456494 homologue to GP|2645999|gb| chlorophyll a/b binding protein of
LHCII type I precursor {Panax ginseng}, partial (45%)
Length = 535
Score = 26.6 bits (57), Expect = 1.5
Identities = 9/16 (56%), Positives = 15/16 (93%)
Frame = +1
Query: 80 KIPPRSMFIVQILGTL 95
K PPRS+F++++LGT+
Sbjct: 25 KQPPRSLFLLEVLGTV 72
>TC76503 homologue to GP|506629|gb|AAA50172.1|| photosystem II type I
chlorophyll a/b-binding protein {Glycine max}, partial
(98%)
Length = 1148
Score = 26.6 bits (57), Expect = 1.5
Identities = 9/16 (56%), Positives = 15/16 (93%)
Frame = +2
Query: 80 KIPPRSMFIVQILGTL 95
K PPRS+F++++LGT+
Sbjct: 161 KQPPRSLFLLEVLGTV 208
>BE998743 similar to GP|22128591|gb| fertility restorer-like protein {Petunia
x hybrida}, partial (7%)
Length = 829
Score = 26.6 bits (57), Expect = 1.5
Identities = 15/50 (30%), Positives = 24/50 (48%)
Frame = -2
Query: 13 ALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGYMS 62
A++ +A+ T+ T Q P +N + G ILP IAN + + S
Sbjct: 258 AIVLKTCLALFTSLTRQYPSINELYVTTSGFILPLIIIANAFLAPFTFPS 109
>TC82829 weakly similar to GP|14334592|gb|AAK59474.1 unknown protein
{Arabidopsis thaliana}, partial (12%)
Length = 703
Score = 26.2 bits (56), Expect = 2.0
Identities = 15/59 (25%), Positives = 28/59 (47%)
Frame = +3
Query: 1 MPWWALIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYG 59
MP++ + LAL++ + T ++ P L +I+G++L G + TYG
Sbjct: 228 MPFFLFLVGISLALVYKNKRSRPTQSSTWKPLLRSFQLFILGILLQGGYFHGIHSFTYG 404
>TC93463 similar to GP|15145367|gb|AAB54239.2 Hypothetical protein K07B1.3
{Caenorhabditis elegans}, partial (5%)
Length = 754
Score = 25.8 bits (55), Expect = 2.6
Identities = 11/23 (47%), Positives = 14/23 (60%)
Frame = +3
Query: 8 FASGLALIFTLPVAIITATTNQT 30
F LA IFT+PV++IT T
Sbjct: 534 FVGALAQIFTMPVSVITTRQQAT 602
>TC89870 similar to PIR|A86387|A86387 probable cytochrome b-561 [imported] -
Arabidopsis thaliana, partial (77%)
Length = 858
Score = 25.4 bits (54), Expect = 3.4
Identities = 19/89 (21%), Positives = 40/89 (44%), Gaps = 6/89 (6%)
Frame = +1
Query: 27 TNQTPGLNVITEYIMGVILPGRPIANVCFKTYGYMSMSQAISFLSDFKLGHYMKIPPRSM 86
TN + + T + +LP +A VC + ++ + A++F S F L +P + +
Sbjct: 82 TNTFRSVVMATATVFPSLLPLLFLARVCGLSVAFLVLFWALTFKSSF-LNTSNSLPQQDL 258
Query: 87 F------IVQILGTLIAGTVDVGVAWWLP 109
++ ++G ++ + V WLP
Sbjct: 259 IYAVLHPVLMVIGFILLSGEAILVHRWLP 345
>TC93345 similar to SP|Q9SJK3|AGOL_ARATH Argonaute-like protein At2g27880.
[Mouse-ear cress] {Arabidopsis thaliana}, partial (12%)
Length = 639
Score = 25.4 bits (54), Expect = 3.4
Identities = 10/27 (37%), Positives = 18/27 (66%)
Frame = -3
Query: 47 GRPIANVCFKTYGYMSMSQAISFLSDF 73
G+ N C ++G++ +S A+ FLS+F
Sbjct: 631 GKITINSCSLSFGFVLLSFALQFLSNF 551
>TC77144 similar to GP|20160594|dbj|BAB89541. contains ESTs AU031508(E61760)
AU068322(C20031) AU068323(C20031)~similar to Arabidopsis
thaliana, partial (46%)
Length = 817
Score = 25.4 bits (54), Expect = 3.4
Identities = 13/29 (44%), Positives = 18/29 (61%)
Frame = -1
Query: 61 MSMSQAISFLSDFKLGHYMKIPPRSMFIV 89
MSMS A+ FL KLG + I P + F++
Sbjct: 382 MSMSLALMFLGFTKLGFSISITP*TPFVL 296
>TC87972 weakly similar to PIR|F96751|F96751 unknown protein F28P22.12
[imported] - Arabidopsis thaliana, partial (81%)
Length = 901
Score = 25.0 bits (53), Expect = 4.4
Identities = 10/24 (41%), Positives = 15/24 (61%)
Frame = -1
Query: 18 LPVAIITATTNQTPGLNVITEYIM 41
+P I+ TTN P ++ IT Y+M
Sbjct: 742 IPTENISNTTNSNPSISKITVYLM 671
>TC80542 similar to GP|18087595|gb|AAL58928.1 unknown protein {Arabidopsis
thaliana}, partial (66%)
Length = 1262
Score = 24.3 bits (51), Expect = 7.5
Identities = 14/48 (29%), Positives = 21/48 (43%)
Frame = +3
Query: 8 FASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCF 55
F AL+F + I A + GL + I G +L +P+A F
Sbjct: 636 FTDMKALVFLVVANAIAAGYSLIQGLRCVVSMIKGSVLFNKPLAWAIF 779
>CA917414 similar to GP|6648966|gb| seed maturation protein PM25 {Glycine
max}, partial (93%)
Length = 781
Score = 24.3 bits (51), Expect = 7.5
Identities = 9/23 (39%), Positives = 14/23 (60%)
Frame = +1
Query: 23 ITATTNQTPGLNVITEYIMGVIL 45
+T T Q PG +ITE + G ++
Sbjct: 286 VTVTETQVPGRRIITETVGGQVV 354
>TC86385 similar to SP|P25469|H2A_LYCES Histone H2A. [Tomato] {Lycopersicon
esculentum}, partial (95%)
Length = 679
Score = 23.9 bits (50), Expect = 9.8
Identities = 10/29 (34%), Positives = 17/29 (58%)
Frame = -2
Query: 67 ISFLSDFKLGHYMKIPPRSMFIVQILGTL 95
I ++DF LG +PP F+V +L ++
Sbjct: 144 IDLVTDFFLGPPPFLPPAPFFVVLVLASI 58
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.328 0.142 0.459
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,349,495
Number of Sequences: 36976
Number of extensions: 34739
Number of successful extensions: 259
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 259
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 259
length of query: 116
length of database: 9,014,727
effective HSP length: 92
effective length of query: 24
effective length of database: 5,612,935
effective search space: 134710440
effective search space used: 134710440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 50 (23.9 bits)
Medicago: description of AC148236.3


