

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
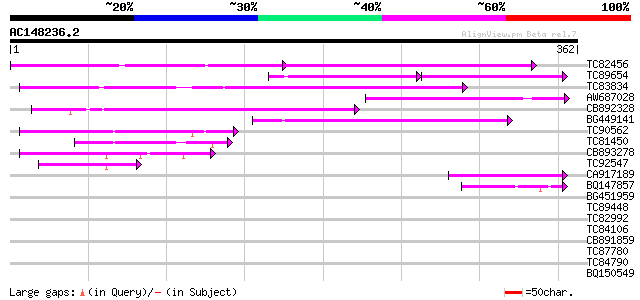
Query= AC148236.2 - phase: 0 /pseudo
(362 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far... 119 2e-38
TC89654 weakly similar to PIR|T05644|T05644 hypothetical protein... 84 5e-29
TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabid... 119 1e-27
AW687028 weakly similar to GP|9909176|dbj| hypothetical protein~... 116 2e-26
CB892328 GP|15075873|em PROBABLE GLYCOSYL HYDROLASE PROTEIN {Sin... 102 3e-22
BG449141 similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidop... 93 1e-19
TC90562 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20... 65 4e-11
TC81450 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20... 57 1e-08
CB893278 weakly similar to PIR|T05645|T056 hypothetical protein ... 54 7e-08
TC92547 similar to PIR|A56235|A56235 transcription activator Maf... 50 1e-06
CA917189 similar to PIR|T05645|T05 hypothetical protein F20D10.3... 44 1e-04
BQ147857 weakly similar to GP|15983442|gb At1g10240/F14N23_12 {A... 43 2e-04
BG451959 homologue to GP|6175165|gb| Mutator-like transposase {A... 39 0.002
TC89448 similar to PIR|C84864|C84864 hypothetical protein At2g43... 35 0.003
TC82992 homologue to PIR|T05645|T05645 hypothetical protein F20D... 37 0.012
TC84106 similar to GP|9502168|gb|AAF88018.1| contains simlarity ... 35 0.045
CB891859 similar to GP|6175165|gb| Mutator-like transposase {Ara... 34 0.077
TC87780 similar to GP|6403499|gb|AAF07839.1| hypothetical protei... 29 3.2
TC84790 similar to GP|13676624|gb|AAK38197.1 phosphate transport... 29 3.2
BQ150549 28 4.2
>TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far-red
impaired response protein {Oryza sativa}, partial (7%)
Length = 1244
Score = 119 bits (299), Expect(2) = 2e-38
Identities = 58/162 (35%), Positives = 88/162 (53%)
Frame = +2
Query: 175 LVQGEAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDS 234
L +G+ + +F R +N FY A +D + I N+FWADAR Y YFG+V++LD+
Sbjct: 704 LWRGDVDAIHSFFHRMQKQNSQFYCAMDMDDKRNIQNLFWADARCRAAYEYFGEVITLDT 883
Query: 235 TYCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFT 294
TY T+ PL G NHH ++ G A+L + A++ WLF +LE P + T
Sbjct: 884 TYLTSKYDLPLVPFVGVNHHGQTILLGCAILSNLDAKTLTWLFTRWLECMHGHAPNGIIT 1063
Query: 295 DQAKSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGES 336
++ K+M A+ P+A H C WH+M+ + LG + ES
Sbjct: 1064EEDKAMKNAIEVAFPKARHRWCLWHIMKKVPEMLGKYSRNES 1189
Score = 57.0 bits (136), Expect(2) = 2e-38
Identities = 45/178 (25%), Positives = 75/178 (41%), Gaps = 1/178 (0%)
Frame = +1
Query: 1 MNEDWNPKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMR 60
++ED P+ M F S + ++ Y + GFGV K + DG C
Sbjct: 190 VSEDEEPRTGMTFRSEEEVIRYYMNYANRVGFGVTKISSKNADDGKKYFTLACNCARRYV 369
Query: 61 KPDKRDYKTKNPRLETRTNCEARLGL-KNVDGKLMVHDFVEDHNHELLLPETTHMLSSQR 119
K K L ++T C ARL +DG + V +HNHEL + + +
Sbjct: 370 STSKNPSKQY---LTSKTQCRARLNACVALDGSSTISRIVLEHNHELGQTKGRYFRLDES 540
Query: 120 KVSEIHCQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQ 177
+ + + +EL D G+ ++ + E+ G NL D +N+++K R LV+
Sbjct: 541 SGTHVE-RNLELNDQGGINVDRNLQSVGLEMNGCGNLTSGENDCRNFVQKVRRLRLVE 711
>TC89654 weakly similar to PIR|T05644|T05644 hypothetical protein F20D10.290
- Arabidopsis thaliana, partial (65%)
Length = 1358
Score = 84.0 bits (206), Expect(2) = 5e-29
Identities = 42/99 (42%), Positives = 59/99 (59%), Gaps = 1/99 (1%)
Frame = +2
Query: 166 YLRKKRERSLVQGEAGY-LLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYS 224
++ R+R+L G G+ +L Y +R EN F++A Q D+++ N+FWAD +YS
Sbjct: 83 HMSSTRQRTL--GGGGHQVLDYLKRMRAENHAFFYAVQSDVDNAGGNIFWADETCRTNYS 256
Query: 225 YFGDVVSLDSTYCTNSSHRPLAIISGFNHHRGAVIFGAA 263
YFGD V D+TY TN P A +GFNHH V+FG A
Sbjct: 257 YFGDTVIFDTTYKTNQYRVPFASFTGFNHHGQPVLFGCA 373
Score = 61.6 bits (148), Expect(2) = 5e-29
Identities = 28/93 (30%), Positives = 52/93 (55%)
Frame = +3
Query: 264 LLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALAEVMPEAYHGLCTWHLMQN 323
L+ +E+ SY WLF+T+L A + P ++ TD + A+A+V+P H C W + +
Sbjct: 375 LILNESEPSYIWLFKTWLRAVSGRPPVSITTDLDPVIQVAVAQVLPPTRHRFCKWSIFRE 554
Query: 324 GIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFE 356
L +L + F ++F+KC++ +++FE
Sbjct: 555 NRSKLAHLYQSNPTFDNEFKKCVHESVTIDEFE 653
>TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabidopsis
thaliana, partial (24%)
Length = 1032
Score = 119 bits (299), Expect = 1e-27
Identities = 78/287 (27%), Positives = 129/287 (44%), Gaps = 1/287 (0%)
Frame = +3
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRKPDKRD 66
P + M F S +A+ ++ Y + GF VR + K++ +CC + KR
Sbjct: 213 PALAMEFESYDDAYSYYICYAKEVGFCVRVKNSWFKRNSKEKYGAVLCCSS---QGFKRT 383
Query: 67 YKTKNPRLETRTNCEARLGLKNVDG-KLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIH 125
N R ETRT C A + +K V+ + + + +HNH L + I
Sbjct: 384 KDVNNLRKETRTGCPAMIRMKLVESQRWRICEVTLEHNHVL----------GAKIHKSIK 533
Query: 126 CQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQ 185
+ +D+ G + K + L + G NL D + + + + +L +G+ +
Sbjct: 534 KNSLPSSDAEG-KTIKVYHALVIDTEGNDNLNSNARDDRAFSKYSNKLNLRKGDTQAIYN 710
Query: 186 YFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPL 245
+ R + NP F++ + E + N W DA+ YF DV+ D+TY N PL
Sbjct: 711 FLCRMQLTNPNFFYLMDFNDEGHLRNALWVDAKSRAACGYFSDVIYFDNTYLVNKYEIPL 890
Query: 246 AIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTV 292
+ G NHH +V+ G LL E ESY+WLF T+++ + PQT+
Sbjct: 891 VALVGINHHGQSVLLGCGLLAGEIIESYKWLFRTWIKCIPRCSPQTI 1031
>AW687028 weakly similar to GP|9909176|dbj| hypothetical protein~similar to
Arabidopsis thaliana F20D10.300, partial (11%)
Length = 662
Score = 116 bits (290), Expect = 2e-26
Identities = 59/130 (45%), Positives = 78/130 (59%)
Frame = +1
Query: 228 DVVSLDSTYCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQK 287
DV++ D+TY N + PL I SG NHH +IFG ALL DET ESY+W+ FLE + K
Sbjct: 1 DVLAFDTTYKKNKYNYPLCIFSGCNHHSQTIIFGVALLEDETIESYKWVLNRFLECMENK 180
Query: 288 MPQTVFTDQAKSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMY 347
P+ V TD SM +A+ +V P+A H LC WHL +N +++ ++ FL F K MY
Sbjct: 181 FPKAVVTDGDGSMREAIKQVFPDASHRLCAWHLHKNAQENI-----KKTPFLEGFRKAMY 345
Query: 348 GYEDVEQFED 357
EQFED
Sbjct: 346 SNFTPEQFED 375
>CB892328 GP|15075873|em PROBABLE GLYCOSYL HYDROLASE PROTEIN {Sinorhizobium
meliloti}, partial (1%)
Length = 828
Score = 102 bits (253), Expect = 3e-22
Identities = 65/214 (30%), Positives = 106/214 (49%), Gaps = 5/214 (2%)
Frame = +1
Query: 15 SLKNAWEFWNTYXGKXGFGVRKD----YTHRKKDGSVSSCRFVCCKEGMRKPDKRDYKTK 70
S A++ + + K GF VRK Y + KK + F C K+G K ++ D +
Sbjct: 178 SKDEAYDLYQEHAFKTGFSVRKGKELYYDNEKKKTRLKD--FYCSKQGF-KNNEPDGEVA 348
Query: 71 NPRLETRTNCEARLGLK-NVDGKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIHCQQI 129
R ++RTNC A + +G V + DHNHE + + ++L S R +S + +I
Sbjct: 349 YKRADSRTNCLAMVRFNVTKEGVWKVTKLILDHNHEFVPLQQRYLLRSMRNMSNLKEDRI 528
Query: 130 ELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQYFQR 189
+ + ++ L KEVGG G + D NY+ + + + G+A LL + Q
Sbjct: 529 KSLVNDAIRVTNVGGYLGKEVGGVDKRGVMLKDMHNYVSTGKLKFIEAGDAQSLLNHLQS 708
Query: 190 KTVENPTFYHAYQLDIEDQITNVFWADARMLVDY 223
K ++ FY++ QLD E ++ NVFW D + +VDY
Sbjct: 709 KQAQDSMFYYSVQLDHESRLNNVFWRDGKSVVDY 810
>BG449141 similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidopsis
thaliana}, partial (33%)
Length = 690
Score = 93.2 bits (230), Expect = 1e-19
Identities = 53/166 (31%), Positives = 81/166 (47%)
Frame = +3
Query: 156 LGFIRLDQKNYLRKKRERSLVQGEAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWA 215
L F D +N L+ R+ + E LL+ + ++P F Y LD +++ N+ W+
Sbjct: 165 LPFTEKDVRNLLQSFRKLD-PEEETLDLLRMCRNIKDKDPNFKFEYTLDANNRLENIAWS 341
Query: 216 DARMLVDYSYFGDVVSLDSTYCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEW 275
A + Y FGD V D+T+ + PL + G N++ FG LL DET S+ W
Sbjct: 342 YASSIQLYDIFGDAVVFDTTHRLTAFDMPLGLWVGINNYGMPCFFGCVLLRDETVRSFSW 521
Query: 276 LFETFLEAHKQKMPQTVFTDQAKSMAKALAEVMPEAYHGLCTWHLM 321
+ FL K PQT+ TDQ + +AL+ MP H C W ++
Sbjct: 522 AIKAFLGFMNGKAPQTILTDQNICLKEALSAEMPMTKHAFCIWMIV 659
>TC90562 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20
{Arabidopsis thaliana}, partial (66%)
Length = 1169
Score = 65.1 bits (157), Expect = 4e-11
Identities = 47/148 (31%), Positives = 71/148 (47%), Gaps = 8/148 (5%)
Frame = +2
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHR-KKDGSVSSCRFVCCKEGMRKPDKR 65
P + F S A F+N+Y + GF +R R ++DGSV VC KEG R PDKR
Sbjct: 323 PYIGQEFVSEAEAHAFYNSYATRVGFVIRVSKLSRSRRDGSVIGRALVCNKEGFRMPDKR 502
Query: 66 DYKTKNPRLETRTNCEARLGLKNVD-GKLMVHDFVEDHNHELLLPETTHML------SSQ 118
+ K R ETR C A + ++ ++ G + FV++H H L ++ S
Sbjct: 503 E-KIVRQRAETRVGCRAMIMVRKLNSGLWSITKFVKEHTHPLTPGKSRRDFVYEQYPSGH 679
Query: 119 RKVSEIHCQQIELADSVGVQQKKSFDLL 146
+V E+ QQ+ + K++ DLL
Sbjct: 680 HRVREL-TQQLAIEKKRAETYKRNLDLL 760
>TC81450 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20
{Arabidopsis thaliana}, partial (59%)
Length = 832
Score = 57.0 bits (136), Expect = 1e-08
Identities = 37/108 (34%), Positives = 59/108 (54%), Gaps = 7/108 (6%)
Frame = +2
Query: 42 KKDGSVSSCRFVCCKEGMRKPDKRDYKTKNPRLETRTNCEARLGLKNVD-GKLMVHDFVE 100
+KDG+ VC KEG R PDKR+ K R ETR C A + ++ ++ GK ++ FV+
Sbjct: 146 RKDGTAIGRALVCNKEGYRMPDKRE-KIVRQRAETRVGCRAMIMMRKINSGKWVITKFVK 322
Query: 101 DHNHELLLPETTHMLSSQRKVSEIHCQQ------IELADSVGVQQKKS 142
+H H L + SS+R + EI+ Q EL+ + +++K+S
Sbjct: 323 EHTHPL------NPGSSRRDMFEIYPSQNEHDKIRELSQQLAIEKKRS 448
>CB893278 weakly similar to PIR|T05645|T056 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (5%)
Length = 821
Score = 54.3 bits (129), Expect = 7e-08
Identities = 44/137 (32%), Positives = 66/137 (48%), Gaps = 12/137 (8%)
Frame = +3
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKK-DGSVSSCRFVCCKEGMR--KPD 63
P + M F S + A F++ Y + GF VR + R + + + FVC KEG R K
Sbjct: 315 PFIGMQFNSREEARGFYDGYGRRIGFTVRIHHNRRSRVNNELIGQDFVCSKEGFRAKKYV 494
Query: 64 KRDYKTKNPRLETRTNCEA--RLGLKNVDGKLMVHDFVEDHNHELLLP-------ETTHM 114
R + P TR C+A RL L++ +GK +V FV++H H+L+ P H+
Sbjct: 495 HRKDRVLPPPPATREGCQAMIRLALRD-EGKWVVTKFVKEHTHKLMSPGEVPWRGSGKHL 671
Query: 115 LSSQRKVSEIHCQQIEL 131
+S K I +EL
Sbjct: 672 VSEDEKDRRIRELSLEL 722
>TC92547 similar to PIR|A56235|A56235 transcription activator MafB - chicken,
partial (6%)
Length = 1158
Score = 50.4 bits (119), Expect = 1e-06
Identities = 28/68 (41%), Positives = 38/68 (55%), Gaps = 2/68 (2%)
Frame = +3
Query: 19 AWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMR--KPDKRDYKTKNPRLET 76
A+EF+ Y GF +RK R G ++ FVC K G+R K R+ + ++ R T
Sbjct: 954 AYEFYYRYGKCKGFSIRKGDVRRNSSGIITMREFVCNKNGLRDKKHLSRNDRKRDHRRLT 1133
Query: 77 RTNCEARL 84
RTNCEARL
Sbjct: 1134 RTNCEARL 1157
>CA917189 similar to PIR|T05645|T05 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (12%)
Length = 312
Score = 43.5 bits (101), Expect = 1e-04
Identities = 19/76 (25%), Positives = 38/76 (50%)
Frame = +1
Query: 281 LEAHKQKMPQTVFTDQAKSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLS 340
L A + P ++ TD + A+ +V P+ H C WH+ + + L ++ F +
Sbjct: 1 LTAMSGRPPLSITTDHDSVIQSAIMQVFPDTRHRFCKWHIFKQCQEKLSHIFLQFPNFEA 180
Query: 341 DFEKCMYGYEDVEQFE 356
+F KC+ + +++FE
Sbjct: 181 EFHKCVNLTDSIDEFE 228
>BQ147857 weakly similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidopsis
thaliana}, partial (26%)
Length = 560
Score = 42.7 bits (99), Expect = 2e-04
Identities = 24/70 (34%), Positives = 36/70 (51%), Gaps = 2/70 (2%)
Frame = +1
Query: 289 PQTVFTDQAKSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESY--FLSDFEKCM 346
PQT+ TD + +A+A MPE+ H C WH++ L+ G Y + +DF + +
Sbjct: 4 PQTLLTDHNTWLKEAIAVEMPESKHAFCIWHILSK-FSDWXYLLLGSQYDEWKADFHR-L 177
Query: 347 YGYEDVEQFE 356
Y E E FE
Sbjct: 178 YNLEMXEDFE 207
>BG451959 homologue to GP|6175165|gb| Mutator-like transposase {Arabidopsis
thaliana}, partial (28%)
Length = 658
Score = 39.3 bits (90), Expect = 0.002
Identities = 22/99 (22%), Positives = 44/99 (44%), Gaps = 4/99 (4%)
Frame = +2
Query: 229 VVSLDSTYCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFE---TFLEAHK 285
++ LD Y + L + +GF+ ++ +E +++ W LE +
Sbjct: 350 LLGLDRIYLKSKYLGTLLLATGFDGDGALFPLAFGVVDEENDDNWMWFLSKLHNLLEINT 529
Query: 286 QKMPQ-TVFTDQAKSMAKALAEVMPEAYHGLCTWHLMQN 323
+ MP+ T+ +D+ + + + P A+HG C HL N
Sbjct: 530 ENMPRLTILSDRQQGIVDGVEANFPTAFHGFCMRHLSDN 646
>TC89448 similar to PIR|C84864|C84864 hypothetical protein At2g43280
[imported] - Arabidopsis thaliana, partial (80%)
Length = 1177
Score = 35.4 bits (80), Expect(2) = 0.003
Identities = 30/95 (31%), Positives = 43/95 (44%), Gaps = 3/95 (3%)
Frame = +1
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRK-KDGSVSSCRFVCCKEG-MRKPDK 64
P M F S A F++ Y GF +R R +DG + + RF C KEG
Sbjct: 301 PYEGMEFVSEDAAKIFYDEYARHVGFVMRVMSCRRSXRDGRILARRFGCNKEGHCVSIQG 480
Query: 65 RDYKTKNPRLETRTNCEARLGLK-NVDGKLMVHDF 98
+ + PR TR C+A + +K + GK M+ F
Sbjct: 481 KLGPVRKPRASTREGCKAMIHIKYDQSGKWMITXF 585
Score = 22.7 bits (47), Expect(2) = 0.003
Identities = 11/34 (32%), Positives = 17/34 (49%)
Frame = +2
Query: 98 FVEDHNHELLLPETTHMLSSQRKVSEIHCQQIEL 131
FV+DHNH L++ + K +I +EL
Sbjct: 584 FVKDHNHPLVVSPREARQTMDEKDKKIQELTVEL 685
>TC82992 homologue to PIR|T05645|T05645 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (7%)
Length = 354
Score = 37.0 bits (84), Expect = 0.012
Identities = 20/56 (35%), Positives = 30/56 (52%), Gaps = 1/56 (1%)
Frame = +1
Query: 4 DWNPKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHR-KKDGSVSSCRFVCCKEG 58
D P M F S + A F+N+Y + GF R + R ++DG++ + VC KEG
Sbjct: 184 DLEPSEGMEFESEEAAKAFYNSYARRVGFSTRVSSSRRSRRDGAIIQRQXVCAKEG 351
>TC84106 similar to GP|9502168|gb|AAF88018.1| contains simlarity to
Arabidopsis thaliana far-red impaired response protein
(GB:AAD51282.1), partial (25%)
Length = 673
Score = 35.0 bits (79), Expect = 0.045
Identities = 23/68 (33%), Positives = 33/68 (47%), Gaps = 4/68 (5%)
Frame = +2
Query: 73 RLETRTNCEARLGLKN--VDG--KLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIHCQQ 128
R R C+A++ L VDG + V F HNHELL + +L + RK+ E ++
Sbjct: 401 RKSVRCGCDAKMYLSKDVVDGVSQWTVLQFSNVHNHELLEDDQVRLLPAYRKIHEADQER 580
Query: 129 IELADSVG 136
I L G
Sbjct: 581 ILLLSKAG 604
>CB891859 similar to GP|6175165|gb| Mutator-like transposase {Arabidopsis
thaliana}, partial (27%)
Length = 625
Score = 34.3 bits (77), Expect = 0.077
Identities = 16/53 (30%), Positives = 27/53 (50%), Gaps = 1/53 (1%)
Frame = +1
Query: 281 LEAHKQKMPQ-TVFTDQAKSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLM 332
LE + + MP+ T+ +D+ + + + P A+HG C HL + K N M
Sbjct: 4 LEVNTENMPRLTILSDRQQGIVDGVEANFPTAFHGFCMRHLSDSFRKEFNNTM 162
>TC87780 similar to GP|6403499|gb|AAF07839.1| hypothetical protein
{Arabidopsis thaliana}, partial (79%)
Length = 1188
Score = 28.9 bits (63), Expect = 3.2
Identities = 15/51 (29%), Positives = 21/51 (40%), Gaps = 2/51 (3%)
Frame = +3
Query: 12 FFCSLKNAWEFWNT--YXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMR 60
F+CS W W T + G + + + KD SC F CCK +
Sbjct: 66 FYCSTWRHWRRWQTSIFVGC*RSSLHRSWCIGVKDLYGQSCDFCCCKSSCK 218
>TC84790 similar to GP|13676624|gb|AAK38197.1 phosphate transporter 2
{Lupinus albus}, partial (37%)
Length = 639
Score = 28.9 bits (63), Expect = 3.2
Identities = 15/27 (55%), Positives = 17/27 (62%)
Frame = +2
Query: 123 EIHCQQIELADSVGVQQKKSFDLLSKE 149
EI +Q E D +GVQ K SF L SKE
Sbjct: 89 EIEAEQ-EKVDKIGVQDKNSFGLFSKE 166
>BQ150549
Length = 350
Score = 28.5 bits (62), Expect = 4.2
Identities = 14/44 (31%), Positives = 22/44 (49%)
Frame = -1
Query: 202 QLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPL 245
+L+ +TN+ W D + Y Y GD+ + ST C+ H L
Sbjct: 311 RLECNVMLTNIIWYDQSSVDIYRYLGDIFCV-STLCSLRVHLSL 183
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,270,647
Number of Sequences: 36976
Number of extensions: 172983
Number of successful extensions: 869
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 862
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 866
length of query: 362
length of database: 9,014,727
effective HSP length: 97
effective length of query: 265
effective length of database: 5,428,055
effective search space: 1438434575
effective search space used: 1438434575
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC148236.2


