

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
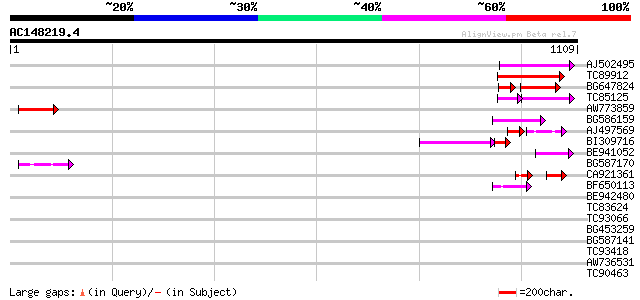
Query= AC148219.4 + phase: 0 /pseudo
(1109 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AJ502495 weakly similar to GP|18071369|g putative gag-pol polypr... 104 2e-22
TC89912 weakly similar to PIR|B84512|B84512 probable retroelemen... 94 4e-19
BG647824 weakly similar to PIR|G96722|G96 hypothetical protein F... 78 3e-18
TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-relate... 68 3e-17
AW773859 83 7e-16
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 68 2e-11
AJ497569 weakly similar to PIR|T04833|T04 hypothetical protein F... 46 1e-10
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 48 8e-09
BE941052 weakly similar to PIR|B85188|B85 retrotransposon like p... 57 4e-08
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 57 5e-08
CA921361 36 4e-06
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 47 3e-05
BE942480 42 0.001
TC83624 homologue to PIR|G84581|G84581 copia-like retroelement p... 41 0.002
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 40 0.004
BG453259 homologue to GP|21434|emb|CA ORF4 {Solanum tuberosum}, ... 40 0.005
BG587141 similar to PIR|H86461|H86 hypothetical protein AAF32440... 40 0.007
TC93418 35 0.21
AW736531 similar to PIR|D84481|D84 probable retroelement pol pol... 29 1.3
TC90463 32 1.8
>AJ502495 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (9%)
Length = 542
Score = 104 bits (259), Expect = 2e-22
Identities = 53/148 (35%), Positives = 85/148 (56%)
Frame = +2
Query: 958 DWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWLLYLL 1017
DWAG ++R+S G F LG ISW + KQ ++ S++EAEY A ++ + WL +L
Sbjct: 5 DWAGDTETRKSTSGYAFHLGTGAISWSSKKQPVVAFSTAEAEYIASTSCATQTVWLRRIL 184
Query: 1018 NDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLLVSSK 1077
+H +YCDN+SA+ + NP+FH +KHI+I H +R+ + + ++
Sbjct: 185 EVMHHEQNTPTKIYCDNKSAIALSKNPVFHGRSKHIDIQFHKIRELIAEKEVVIEYCPTE 364
Query: 1078 DQLADIFTNPLLPQPFSTLLSNLGMLNS 1105
+++ADIFT PL + F L LGM+ +
Sbjct: 365 EKIADIFTKPLKIESFYKLKKMLGMMKA 448
>TC89912 weakly similar to PIR|B84512|B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(10%)
Length = 814
Score = 93.6 bits (231), Expect = 4e-19
Identities = 45/132 (34%), Positives = 83/132 (62%)
Frame = +1
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
+ D D+AG +D+R+S+ G F L + ISW+ +Q ++ S+++AEY A + WL
Sbjct: 97 YVDADYAGNVDTRKSLSGFVFTLYGTTISWKANQQSVVTLSTTQAEYIAFVEGVKDAIWL 276
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
++ +L +T + + +CD+QSA+H+ + ++HE TKHI+I H +RD +++ I
Sbjct: 277 KGMIGELGITQEYVKI-HCDSQSAIHLANHQVYHERTKHIDIRLHFIRDMIESKEIVVEK 453
Query: 1074 VSSKDQLADIFT 1085
++S++ AD+FT
Sbjct: 454 MASEENPADVFT 489
>BG647824 weakly similar to PIR|G96722|G96 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (5%)
Length = 721
Score = 77.8 bits (190), Expect(2) = 3e-18
Identities = 40/78 (51%), Positives = 51/78 (65%), Gaps = 1/78 (1%)
Frame = -3
Query: 1000 YRALSAATYELQWLLYLLNDLHVTTVKLPVLYCDNQSAL-HIGTNPMFHEITKHIEIICH 1058
YR++ + E++WL YLLNDL T +K +LYCDNQSA HI N F E TKHIE+ CH
Sbjct: 374 YRSI*STICEIKWLTYLLNDLKFTFIKPAMLYCDNQSAARHIAANSSFLERTKHIELDCH 195
Query: 1059 LVRDKLQAGIIKFLLVSS 1076
+VR KLQ + L + S
Sbjct: 194 IVRVKLQLKLFHILHILS 141
Score = 33.5 bits (75), Expect(2) = 3e-18
Identities = 15/32 (46%), Positives = 23/32 (71%)
Frame = -2
Query: 957 VDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQ 988
+D + CLD+ +SI C FLG+SLI W++ K+
Sbjct: 489 LD*SSCLDT*KSISYFCIFLGDSLICWKS*KK 394
>TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
(EC 3.4.23.-);, partial (7%)
Length = 705
Score = 67.8 bits (164), Expect(2) = 3e-17
Identities = 38/104 (36%), Positives = 55/104 (52%)
Frame = +3
Query: 1002 ALSAATYELQWLLYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVR 1061
+L A E W+ L+ +L ++ V YCD+QSALHI NP FH TKHI I H VR
Sbjct: 228 SLPQACKEAIWMQRLMEELGHKQEQITV-YCDSQSALHIARNPAFHSRTKHIGIQYHFVR 404
Query: 1062 DKLQAGIIKFLLVSSKDQLADIFTNPLLPQPFSTLLSNLGMLNS 1105
+ ++ G + + + D LAD T + F S+ G+L +
Sbjct: 405 EVVEEGSVDMQKIHTNDNLADAMTKSINTDKFIWCRSSYGLLET 536
Score = 39.7 bits (91), Expect(2) = 3e-17
Identities = 19/49 (38%), Positives = 28/49 (56%)
Frame = +1
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRA 1002
+ D D+AG D R+S G F L +SW + Q ++ S++EAEY A
Sbjct: 82 YVDSDFAGDHDKRKSTTGYVFTLAGGAVSWLSKLQTVVALSTTEAEYMA 228
>AW773859
Length = 538
Score = 82.8 bits (203), Expect = 7e-16
Identities = 39/78 (50%), Positives = 52/78 (66%)
Frame = -3
Query: 17 AL*HFRLGHVSYERLAHMSQLYPF*LFFDSNATCDICHFARQKQLPFHLSSSVASNKFEL 76
AL HFRLGH+S +L + +PF + D N+ CDICH++R K+LPF LS++ AS +EL
Sbjct: 233 ALWHFRLGHLSNRKLLSLHSNFPF-ITIDQNSVCDICHYSRHKKLPFQLSTNRASKCYEL 57
Query: 77 LHFDI*GPLIVPFIHNHK 94
HFDI GP IHN +
Sbjct: 56 FHFDIWGPFSTQSIHNQR 3
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 68.2 bits (165), Expect = 2e-11
Identities = 34/103 (33%), Positives = 55/103 (53%)
Frame = +1
Query: 945 RNVLVKVFFFPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALS 1004
RN K+ + D D+AG LD R+S G F L + +SW + KQ ++ S+++AE+ A +
Sbjct: 343 RNGSEKLEAYTDSDYAGDLDDRKSTSGYVFMLSSGAVSWSSKKQPVVTLSTTKAEFIAAA 522
Query: 1005 AATYELQWLLYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFH 1047
+ W+ +L L T +YCDN S + + NP+ H
Sbjct: 523 FCACQSVWMRRVLEKLGYTQSGSITMYCDNNSTIKLSKNPVLH 651
>AJ497569 weakly similar to PIR|T04833|T04 hypothetical protein F21P8.50 -
Arabidopsis thaliana, partial (4%)
Length = 723
Score = 46.2 bits (108), Expect(2) = 1e-10
Identities = 29/77 (37%), Positives = 45/77 (57%)
Frame = +2
Query: 1012 WLLYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKF 1071
++ Y+++ H T V + YCDN SALHI N +FHE T H E ++V+ + +++
Sbjct: 242 FIFYMISAKHST*V---LQYCDNISALHIAANMVFHERT*HRETDPYIVQG---SRMLQL 403
Query: 1072 LLVSSKDQLADIFTNPL 1088
+ +SKDQ A T PL
Sbjct: 404 MPSASKDQPAYSLTKPL 454
Score = 38.9 bits (89), Expect(2) = 1e-10
Identities = 20/33 (60%), Positives = 25/33 (75%)
Frame = +3
Query: 975 FLGNSLISWRTMKQLTISRSSSEAEYRALSAAT 1007
FL +SLISW++ KQ +SRS SEA RAL+ AT
Sbjct: 126 FLSSSLISWKSKKQCVVSRSFSEA**RALANAT 224
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 48.1 bits (113), Expect(2) = 8e-09
Identities = 51/149 (34%), Positives = 74/149 (49%)
Frame = +3
Query: 802 FMLMMLLLLVIHLMNFNSSNIFYILHSKLKILVS*STFLV*SLLTLHKEYFCARESIVLI 861
+MLM L L + + N N+F ++ K K LV F V LL +K ++ +E+I+L
Sbjct: 180 YMLMTLF*LEMIYLKSNMLNVFLLIVLKSKTLVPYDIF*VLRLLEANKAFYLIKENILLN 359
Query: 862 FSLIQVTLHLNLFLHHMIHHANFIVIIVHLTLICLHTKD**GD*FT*PTLDLTLLLPLNN 921
F I V L N L MI N ++I T++ L+ +D * + F * L L L NN
Sbjct: 360 F*RIVVILL*NPLLLLMIFL*NSTILIRPFTMMKLNIEDS*ANLFI*LLLVLIYPLLFNN 539
Query: 922 LASSSLLLLRNTT*LPLES*DI*RNVLVK 950
A+ L + T L LE +I*+ L K
Sbjct: 540 *ANLFRNLSKFTIRLLLEFYNI*KLPLPK 626
Score = 30.8 bits (68), Expect(2) = 8e-09
Identities = 13/30 (43%), Positives = 19/30 (63%)
Frame = +2
Query: 949 VKVFFFPDVDWAGCLDSRRSIFGQCFFLGN 978
+K+ F D DWA C +R+S+ G FLG+
Sbjct: 653 LKLSSFADSDWATCPTTRKSVTGYWVFLGS 742
>BE941052 weakly similar to PIR|B85188|B85 retrotransposon like protein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 480
Score = 57.0 bits (136), Expect = 4e-08
Identities = 30/74 (40%), Positives = 41/74 (54%)
Frame = +2
Query: 1029 VLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLLVSSKDQLADIFTNPL 1088
+L CD SA ++ NP++H KHI I H VRD +Q G +K V + DQLAD T PL
Sbjct: 20 LLRCDYLSATYLTHNPVYHSRMKHISIDIHFVRDLVQQGKLKVQHVCTVDQLADCLTKPL 199
Query: 1089 LPQPFSTLLSNLGM 1102
L + +G+
Sbjct: 200 SKSRHQLLRNKIGV 241
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 56.6 bits (135), Expect = 5e-08
Identities = 37/111 (33%), Positives = 54/111 (48%), Gaps = 2/111 (1%)
Frame = -3
Query: 17 AL*HFRLGHVSYERLAHMSQLYPF*LFFDSNATCDICHFARQKQLPFHLSSSVASNKFEL 76
AL H RLGH L M F N C+ C + + F +S+V N F+L
Sbjct: 587 ALWHARLGHPHGRALNLMLPGVVF-----ENKNCEACILGKHCKNVFPRTSTVYENCFDL 423
Query: 77 LHFDI*GPLIVPFIH--NHKYFLTILDDSSRFVWIVLLKSKAEVSQHVKNF 125
++ D+ P + NHKYF+T +D+ S++ W+ L+ SK V KNF
Sbjct: 422 IYTDL---WTAPSLSRDNHKYFVTFIDEKSKYTWLTLIPSKDRVIDAFKNF 279
>CA921361
Length = 466
Score = 36.2 bits (82), Expect(2) = 4e-06
Identities = 18/34 (52%), Positives = 25/34 (72%)
Frame = -2
Query: 989 LTISRSSSEAEYRALSAATYELQWLLYLLNDLHV 1022
+TIS+SS + +YR +++ ELQWL YLLND V
Sbjct: 399 VTISKSS*D-KYRVMTSTICELQWLAYLLNDFKV 301
Score = 33.5 bits (75), Expect(2) = 4e-06
Identities = 19/39 (48%), Positives = 24/39 (60%)
Frame = -1
Query: 1050 TKHIEIICHLVRDKLQAGIIKFLLVSSKDQLADIFTNPL 1088
T+HIE+ C +V +KL + LLVSS LAD T PL
Sbjct: 238 TEHIELDCRIV*EKLPQNLFH-LLVSSSLHLADCVTKPL 125
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 47.4 bits (111), Expect = 3e-05
Identities = 27/76 (35%), Positives = 42/76 (54%)
Frame = +1
Query: 945 RNVLVKVFFFPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALS 1004
++ + ++ + D DW G RRS G F ++ ISW T KQ + SS EAEY A +
Sbjct: 262 KSEVYELICYSDSDWCG---DRRSTSGYVFKFNDAAISWCTKKQPITALSSYEAEYIAGT 432
Query: 1005 AATYELQWLLYLLNDL 1020
AT++ WL ++ +L
Sbjct: 433 FATFQALWLDSVIKEL 480
>BE942480
Length = 396
Score = 42.4 bits (98), Expect = 0.001
Identities = 18/43 (41%), Positives = 30/43 (68%)
Frame = -2
Query: 1000 YRALSAATYELQWLLYLLNDLHVTTVKLPVLYCDNQSALHIGT 1042
YRA+S+ E++WL Y+++ L V ++K + Y DNQ+A HI +
Sbjct: 317 YRAMSSIVCEIEWLTYIVDVLKVQSIKPTLPYYDNQAARHIAS 189
>TC83624 homologue to PIR|G84581|G84581 copia-like retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(1%)
Length = 831
Score = 41.2 bits (95), Expect = 0.002
Identities = 34/100 (34%), Positives = 47/100 (47%), Gaps = 8/100 (8%)
Frame = +1
Query: 1010 LQWLLYLL----NDLHVTTVKLPVLYCDNQSALHI---GTNPMFHEITKHIEIICHLVRD 1062
LQWLLY + +H TT L C + + +H G + + K ++I ++R
Sbjct: 421 LQWLLYFFTKPTSSMHTTTSHL---LCQSNNFIHCKKPGLS*KNQTLRKQLDIF--VLRK 585
Query: 1063 KLQAGIIKFLLVSSKDQLADIF-TNPLLPQPFSTLLSNLG 1101
+ GI + LADIF T LLP PF LLS LG
Sbjct: 586 SCKLGIASNFQYLPRTNLADIFFTKSLLP*PFHILLSKLG 705
Score = 40.8 bits (94), Expect = 0.003
Identities = 20/48 (41%), Positives = 28/48 (57%), Gaps = 5/48 (10%)
Frame = +2
Query: 1012 WLLY-----LLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIE 1054
W+LY L +L V +L ++YC NQ L+I N ++HE TKH E
Sbjct: 413 WILYNGCYTFLRNLQVQCTRLLLIYCVNQITLYIAKNQVYHERTKH*E 556
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 40.4 bits (93), Expect = 0.004
Identities = 19/63 (30%), Positives = 32/63 (50%)
Frame = +1
Query: 55 FARQKQLPFHLSSSVASNKFELLHFDI*GPLIVPFIHNHKYFLTILDDSSRFVWIVLLKS 114
F +K++ F ++ + +H D+ GP V +Y +TI+DD R VW+ L+
Sbjct: 40 FGNRKKVSFSTATHRTKGILDYIHSDLWGPSKVTSYGGRRYMMTIIDDFPRKVWVYFLRY 219
Query: 115 KAE 117
K E
Sbjct: 220 KNE 228
>BG453259 homologue to GP|21434|emb|CA ORF4 {Solanum tuberosum}, partial (5%)
Length = 657
Score = 40.0 bits (92), Expect = 0.005
Identities = 33/112 (29%), Positives = 51/112 (45%), Gaps = 2/112 (1%)
Frame = -2
Query: 960 AGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYEL--QWLLYLL 1017
AG + R S G FLG +++ KQ ++R + S + + L L
Sbjct: 473 AGWIVDRGSTSGY*MFLGGNMVE*---KQNVVAR*VQRHNFELCSQGL*RVMDEELKIKL 303
Query: 1018 NDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGII 1069
+DL + L+ +N I NP+ H TKHIEI H + +KL +G+I
Sbjct: 302 DDLIINYKDPMTLF*NNNFVSRIAHNPVQHYRTKHIEIDQHFIIEKLYSGLI 147
>BG587141 similar to PIR|H86461|H86 hypothetical protein AAF32440.1 [imported]
- Arabidopsis thaliana, partial (20%)
Length = 731
Score = 39.7 bits (91), Expect = 0.007
Identities = 27/99 (27%), Positives = 47/99 (47%)
Frame = +3
Query: 1006 ATYELQWLLYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQ 1065
A + WL LL+++ + V+ DNQS + + NP+FH HI H +R+ ++
Sbjct: 126 AARQAMWLQDLLSEVTWEPCEEVVIRIDNQSVIALTRNPVFHGRGNHIHKRYHFIRECVE 305
Query: 1066 AGIIKFLLVSSKDQLADIFTNPLLPQPFSTLLSNLGMLN 1104
G ++ V + A I T L F + +GM++
Sbjct: 306 NGQVEVEHVPGEKHRAYI*TKALGRIIFREIRYYIGMID 422
>TC93418
Length = 533
Score = 34.7 bits (78), Expect = 0.21
Identities = 14/22 (63%), Positives = 18/22 (81%)
Frame = +3
Query: 1033 DNQSALHIGTNPMFHEITKHIE 1054
DNQSALH+ +N +FHE T HI+
Sbjct: 225 DNQSALHVTSNLIFHEWTNHID 290
>AW736531 similar to PIR|D84481|D84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (1%)
Length = 635
Score = 28.9 bits (63), Expect(2) = 1.3
Identities = 12/18 (66%), Positives = 15/18 (82%)
Frame = +3
Query: 1038 LHIGTNPMFHEITKHIEI 1055
L+I +NP+FH TKHIEI
Sbjct: 453 LYIASNPVFH*QTKHIEI 506
Score = 21.6 bits (44), Expect(2) = 1.3
Identities = 9/17 (52%), Positives = 12/17 (69%)
Frame = +2
Query: 1072 LLVSSKDQLADIFTNPL 1088
L ++ DQLAD+FT L
Sbjct: 518 LSINPNDQLADMFTKVL 568
>TC90463
Length = 1175
Score = 31.6 bits (70), Expect = 1.8
Identities = 14/29 (48%), Positives = 17/29 (58%)
Frame = -2
Query: 1080 LADIFTNPLLPQPFSTLLSNLGMLNSYHS 1108
L D T L P F + +S LGMLN YH+
Sbjct: 322 LPDFLTKALPPPKFHSFISKLGMLNIYHA 236
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.362 0.160 0.603
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 46,646,226
Number of Sequences: 36976
Number of extensions: 889695
Number of successful extensions: 12535
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 3983
Number of HSP's successfully gapped in prelim test: 749
Number of HSP's that attempted gapping in prelim test: 8074
Number of HSP's gapped (non-prelim): 5672
length of query: 1109
length of database: 9,014,727
effective HSP length: 106
effective length of query: 1003
effective length of database: 5,095,271
effective search space: 5110556813
effective search space used: 5110556813
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (22.0 bits)
S2: 64 (29.3 bits)
Medicago: description of AC148219.4


