

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148098.5 + phase: 0
(189 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
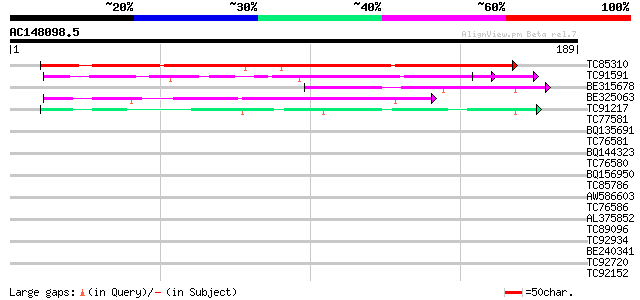
TC85310 homologue to GP|21537011|gb|AAM61352.1 unknown {Arabidop... 181 1e-46
TC91591 homologue to GP|10177252|dbj|BAB10720. gb|AAF34833.1~gen... 96 2e-21
BE315678 54 5e-08
BE325063 similar to PIR|G84718|G847 hypothetical protein At2g312... 47 6e-06
TC91217 similar to GP|5306274|gb|AAD42006.1| expressed protein {... 40 5e-04
TC77581 similar to PIR|T10861|T10861 phaseolin G-box binding pro... 32 0.19
BQ135691 similar to PIR|E96636|E966 hypothetical protein T7P1.21... 30 0.73
TC76581 homologue to PIR|S40240|S40240 ubiquitin/ribosomal prote... 29 0.96
BQ144323 similar to GP|10177252|dbj gb|AAF34833.1~gene_id:K19P17... 27 1.2
TC76580 homologue to PIR|S40240|S40240 ubiquitin/ribosomal prote... 29 1.2
BQ156950 homologue to PIR|S40240|S402 ubiquitin/ribosomal protei... 29 1.2
TC85786 homologue to SP|Q05046|CH62_CUCMA Chaperonin CPN60-2 mi... 28 2.1
AW586603 similar to GP|5306274|gb|A expressed protein {Arabidops... 28 2.8
TC76586 homologue to PIR|S40240|S40240 ubiquitin/ribosomal prote... 27 3.6
AL375852 similar to PIR|T47877|T47 hypothetical protein T4C21.70... 27 4.7
TC89096 similar to GP|15983801|gb|AAL10497.1 At3g07700/F17A17.4 ... 27 6.2
TC92934 weakly similar to GP|5918254|emb|CAB56299.1 NBS-LRR prot... 27 6.2
BE240341 similar to GP|16549089|db respiratory burst oxidase hom... 27 6.2
TC92720 27 6.2
TC92152 weakly similar to PIR|G86266|G86266 hypothetical protein... 27 6.2
>TC85310 homologue to GP|21537011|gb|AAM61352.1 unknown {Arabidopsis
thaliana}, partial (44%)
Length = 1071
Score = 181 bits (460), Expect = 1e-46
Identities = 90/176 (51%), Positives = 118/176 (66%), Gaps = 17/176 (9%)
Frame = +3
Query: 11 LLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVW 70
LLQHTLR+LC +S+WVYAVFWR++PR +PPP+WE G + DRS+GN+RNWI+VW
Sbjct: 528 LLQHTLRSLCIHE----NSQWVYAVFWRILPRNYPPPKWE-GQGAYDRSRGNRRNWILVW 692
Query: 71 EDGFCDF----------NECEQRKSGCLN-------ERFGADVFFKMSHEVYSYGEGLVG 113
EDGFC+F +C S N + ++FFKMSHE+Y+YGEGL+G
Sbjct: 693 EDGFCNFAASAAPEINTGDCPSSSSVYGNCELIQPYQGLQPELFFKMSHEIYNYGEGLIG 872
Query: 114 KVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKE 169
KVAAD+ HKW+Y + N E N++ W+ S D PR WE Q SGI+TIA+IAV+E
Sbjct: 873 KVAADHSHKWIYKE-PNDQEINFLSAWHNSADSHPRTWEAQFLSGIKTIALIAVRE 1037
>TC91591 homologue to GP|10177252|dbj|BAB10720.
gb|AAF34833.1~gene_id:K19P17.6~similar to unknown
protein {Arabidopsis thaliana}, partial (48%)
Length = 603
Score = 95.5 bits (236), Expect(2) = 2e-21
Identities = 62/156 (39%), Positives = 82/156 (51%), Gaps = 5/156 (3%)
Frame = +1
Query: 12 LQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFG-GTSLDRSKGNKRNWIIVW 70
L LR +C +S W Y+VFW + PR PR G G + G+ +++W
Sbjct: 64 LHEALRTVC------LNSDWTYSVFWTIRPR----PRVRGGNGCKVGDDNGSL---MLMW 204
Query: 71 EDGFCDFNECEQRKSGCLNERFGAD----VFFKMSHEVYSYGEGLVGKVAADNGHKWVYS 126
EDGFC + +SG + E G D F KMS ++Y+YGEGL+GKVA+D HKWV+
Sbjct: 205 EDGFC------RGRSG-VEEIDGEDPVRKAFSKMSIQLYNYGEGLMGKVASDKCHKWVFK 363
Query: 127 DTQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTI 162
+ CE N W +S D P W Q SGIQTI
Sbjct: 364 EPTE-CEPNISNYWQSSFDALPPEWTDQFESGIQTI 468
Score = 23.1 bits (48), Expect(2) = 2e-21
Identities = 11/22 (50%), Positives = 13/22 (59%)
Frame = +2
Query: 155 LNSGIQTIAVIAVKEGLVQLGS 176
LN + AVI GL+QLGS
Sbjct: 446 LNQAFRPYAVIQAGHGLLQLGS 511
>BE315678
Length = 528
Score = 53.5 bits (127), Expect = 5e-08
Identities = 33/84 (39%), Positives = 45/84 (53%), Gaps = 2/84 (2%)
Frame = +3
Query: 99 KMSHEVYSYGEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNAS-IDHQPRAWEFQLNS 157
+MS YS GEG VGK+A H WV CE + G ++ S I P W Q S
Sbjct: 150 EMSRRKYSLGEGAVGKLALAKDHFWV------SCEDIFTGKFDTSLIPECPDEWLLQFAS 311
Query: 158 GIQTIAVIAV-KEGLVQLGSFDKV 180
GI+TI ++ V +G++Q SF+ V
Sbjct: 312 GIKTIVLVPVLPQGVLQFASFNSV 383
>BE325063 similar to PIR|G84718|G847 hypothetical protein At2g31280
[imported] - Arabidopsis thaliana, partial (5%)
Length = 610
Score = 46.6 bits (109), Expect = 6e-06
Identities = 34/138 (24%), Positives = 61/138 (43%), Gaps = 7/138 (5%)
Frame = +3
Query: 12 LQHTLRNLCSFPTSSTSSKWVYAVFWRV---VPRIFPPPRWEFGGTSLDRSKGNKRNWII 68
L LR+LC +++W YA+FW++ P I +L+ + + ++
Sbjct: 87 LHQLLRSLCF------NTQWNYAIFWKLKHSAPLIL----------TLEDAYYDNSDYFD 218
Query: 69 VWEDGFCDFNECEQRKSGCLNERFGADVFFKMSHEVYSYGEGLVGKVAADNGHKWVYSD- 127
E+ +C EQ + G + KM + Y G+G+VG+VA H+W+ +D
Sbjct: 219 ASENKYCQ-KTLEQIEGGKFSHDALGLAVAKMRYNAYPLGQGIVGQVAVTGKHQWICADN 395
Query: 128 ---TQNGCEANYVGPWNA 142
T +G +V W +
Sbjct: 396 NQVTSSGLSFEFVDGWQS 449
>TC91217 similar to GP|5306274|gb|AAD42006.1| expressed protein {Arabidopsis
thaliana}, partial (23%)
Length = 1076
Score = 40.0 bits (92), Expect = 5e-04
Identities = 45/196 (22%), Positives = 76/196 (37%), Gaps = 29/196 (14%)
Frame = +1
Query: 11 LLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVW 70
LL+ L+ LC+ ++W YAVFW++ G + +++W
Sbjct: 241 LLKEALKTLCA------RNQWSYAVFWKI---------------------GCNNSKLLIW 339
Query: 71 EDGFCD-------------FNECEQRKSGCLNERFGADVFFKMSHE-------------- 103
ED + + N Q + GC F +D ++ +
Sbjct: 340 EDCYYEPLPSTFLPQNVGTSNLPYQDREGCW---FSSDSQLRIQEDDRVCSLINKMMVNN 510
Query: 104 -VYSYGEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTI 162
V G+G++G+ A H+W+ N + P + H Q ++G+QT+
Sbjct: 511 SVNVAGQGILGRAAFTGNHQWI---LLNNFIKDVYPPEVLNEVH------CQFSAGMQTV 663
Query: 163 AVIAV-KEGLVQLGSF 177
VI V G+VQLGSF
Sbjct: 664 VVIPVLPHGVVQLGSF 711
>TC77581 similar to PIR|T10861|T10861 phaseolin G-box binding protein PG1 -
kidney bean, partial (27%)
Length = 1033
Score = 31.6 bits (70), Expect = 0.19
Identities = 41/190 (21%), Positives = 70/190 (36%), Gaps = 22/190 (11%)
Frame = +2
Query: 13 QHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVWED 72
Q TL+ W YA+FW+ P +++ G+SL + W D
Sbjct: 242 QDTLQQRLQALIEGVKEIWTYAIFWQ--------PSYDYSGSSL-----------LGWGD 364
Query: 73 GFCDFNECEQR-KSGCLNERFGADVFFKMSHEVYSY-------GEGLVGKVAADNGHKWV 124
G+ E + + K + + K+ E+YS E V + D ++
Sbjct: 365 GYYKGEEDKTKVKKSIVTSPAEQEHRRKVLRELYSLISGNPVTEESPVDEEVTDMEWFFL 544
Query: 125 YSDTQ--------------NGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKEG 170
S TQ N VG N + H RA + Q G++T+ + G
Sbjct: 545 VSMTQSFVNDGGLPGQAYFNSTPVWLVGGENLVLSHCERARQGQ-EHGLETLVCVPSANG 721
Query: 171 LVQLGSFDKV 180
+++LGS + +
Sbjct: 722 VLELGSTELI 751
>BQ135691 similar to PIR|E96636|E966 hypothetical protein T7P1.21 [imported]
- Arabidopsis thaliana, partial (4%)
Length = 1215
Score = 29.6 bits (65), Expect = 0.73
Identities = 15/38 (39%), Positives = 21/38 (54%)
Frame = +1
Query: 83 RKSGCLNERFGADVFFKMSHEVYSYGEGLVGKVAADNG 120
R +GC+ R+GA V+ E+ +GEG G A NG
Sbjct: 700 RGTGCV*VRYGARVYGVRVDEMGQWGEGSAGMARAVNG 813
>TC76581 homologue to PIR|S40240|S40240 ubiquitin/ribosomal protein S27a
fusion protein - white lupine, complete
Length = 665
Score = 29.3 bits (64), Expect = 0.96
Identities = 12/23 (52%), Positives = 16/23 (69%)
Frame = -2
Query: 5 LPLLHSLLQHTLRNLCSFPTSST 27
+P HS L H+ +LC+FP SST
Sbjct: 427 VPAPHSALGHSFLSLCTFPESST 359
>BQ144323 similar to GP|10177252|dbj gb|AAF34833.1~gene_id:K19P17.6~similar
to unknown protein {Arabidopsis thaliana}, partial (15%)
Length = 817
Score = 26.6 bits (57), Expect(2) = 1.2
Identities = 11/31 (35%), Positives = 16/31 (51%)
Frame = +1
Query: 12 LQHTLRNLCSFPTSSTSSKWVYAVFWRVVPR 42
L L+ +C +S W Y+VFW + PR
Sbjct: 157 LHEALKTVC------LNSDWTYSVFWTIRPR 231
Score = 20.8 bits (42), Expect(2) = 1.2
Identities = 5/9 (55%), Positives = 9/9 (99%)
Frame = +2
Query: 67 IIVWEDGFC 75
+++WE+GFC
Sbjct: 287 MLMWENGFC 313
>TC76580 homologue to PIR|S40240|S40240 ubiquitin/ribosomal protein S27a
fusion protein - white lupine, complete
Length = 881
Score = 28.9 bits (63), Expect = 1.2
Identities = 12/23 (52%), Positives = 15/23 (65%)
Frame = -3
Query: 5 LPLLHSLLQHTLRNLCSFPTSST 27
+P HS L H+ NLC+ P SST
Sbjct: 552 VPAPHSALGHSFLNLCTLPESST 484
>BQ156950 homologue to PIR|S40240|S402 ubiquitin/ribosomal protein S27a
fusion protein - white lupine, partial (56%)
Length = 678
Score = 28.9 bits (63), Expect = 1.2
Identities = 12/23 (52%), Positives = 15/23 (65%)
Frame = -1
Query: 5 LPLLHSLLQHTLRNLCSFPTSST 27
+P HS L H+ NLC+ P SST
Sbjct: 516 VPAPHSALGHSFLNLCTLPESST 448
>TC85786 homologue to SP|Q05046|CH62_CUCMA Chaperonin CPN60-2 mitochondrial
precursor (HSP60-2). [Pumpkin Winter squash] {Cucurbita
maxima}, partial (58%)
Length = 1259
Score = 28.1 bits (61), Expect = 2.1
Identities = 12/33 (36%), Positives = 20/33 (60%), Gaps = 1/33 (3%)
Frame = -3
Query: 59 SKGNKRNWIIVWEDGFCDFNECEQR-KSGCLNE 90
S+ ++ +I WE G F+ C QR +GC+N+
Sbjct: 987 SRHSRTIFIFTWEIGHNSFSRCHQR*YTGCINQ 889
>AW586603 similar to GP|5306274|gb|A expressed protein {Arabidopsis
thaliana}, partial (4%)
Length = 534
Score = 27.7 bits (60), Expect = 2.8
Identities = 11/29 (37%), Positives = 18/29 (61%)
Frame = +1
Query: 11 LLQHTLRNLCSFPTSSTSSKWVYAVFWRV 39
LL+ L+ LC +++W YAVFW++
Sbjct: 262 LLKEALKTLCG-----RNNQWSYAVFWKI 333
>TC76586 homologue to PIR|S40240|S40240 ubiquitin/ribosomal protein S27a
fusion protein - white lupine, partial (47%)
Length = 439
Score = 27.3 bits (59), Expect = 3.6
Identities = 11/19 (57%), Positives = 13/19 (67%)
Frame = -1
Query: 9 HSLLQHTLRNLCSFPTSST 27
HS L H+ NLC+ P SST
Sbjct: 184 HSALGHSFLNLCTLPESST 128
>AL375852 similar to PIR|T47877|T47 hypothetical protein T4C21.70 -
Arabidopsis thaliana, partial (38%)
Length = 545
Score = 26.9 bits (58), Expect = 4.7
Identities = 13/34 (38%), Positives = 17/34 (49%)
Frame = +1
Query: 33 YAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNW 66
Y + R + P+ E G SL +S GNKR W
Sbjct: 43 YGLLCRSKCSTYRSPQKEVGRKSLGKSSGNKRYW 144
>TC89096 similar to GP|15983801|gb|AAL10497.1 At3g07700/F17A17.4
{Arabidopsis thaliana}, partial (46%)
Length = 1149
Score = 26.6 bits (57), Expect = 6.2
Identities = 12/34 (35%), Positives = 21/34 (61%)
Frame = +1
Query: 155 LNSGIQTIAVIAVKEGLVQLGSFDKVLLFFPSKY 188
+ +GI I + + + +L F+K+L FFP+KY
Sbjct: 907 VGAGILFILFLRSMQRVQKLDKFEKML*FFPNKY 1008
>TC92934 weakly similar to GP|5918254|emb|CAB56299.1 NBS-LRR protein
{Solanum acaule}, partial (5%)
Length = 625
Score = 26.6 bits (57), Expect = 6.2
Identities = 17/56 (30%), Positives = 27/56 (47%)
Frame = +2
Query: 30 KWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVWEDGFCDFNECEQRKS 85
K+++AVFW +PR P R +F R W+ +GF CE+R++
Sbjct: 419 KFMHAVFWVYIPRTTPSLRRDF-----------TRQWM---AEGFV---RCEERRN 535
>BE240341 similar to GP|16549089|db respiratory burst oxidase homolog
{Solanum tuberosum}, partial (16%)
Length = 569
Score = 26.6 bits (57), Expect = 6.2
Identities = 13/58 (22%), Positives = 25/58 (42%)
Frame = -2
Query: 99 KMSHEVYSYGEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLN 156
K+ + +GEG + N + ++ N CEA + P+++ + H Q N
Sbjct: 238 KLHPHHFWHGEGTEAL*SKPNERRLLHKHLYNNCEAQLLLPYHSPLPHSSHLSTIQTN 65
>TC92720
Length = 808
Score = 26.6 bits (57), Expect = 6.2
Identities = 10/21 (47%), Positives = 13/21 (61%)
Frame = +2
Query: 78 NECEQRKSGCLNERFGADVFF 98
+EC+ +KSGC R G FF
Sbjct: 698 SECQAKKSGCYAHRLGQGRFF 760
>TC92152 weakly similar to PIR|G86266|G86266 hypothetical protein AAD31075.1
[imported] - Arabidopsis thaliana, partial (5%)
Length = 645
Score = 26.6 bits (57), Expect = 6.2
Identities = 13/40 (32%), Positives = 24/40 (59%)
Frame = +1
Query: 5 LPLLHSLLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIF 44
LPL S L+++ N+ + ++ + SKW +VF + +IF
Sbjct: 277 LPLPTSPLENSKDNVIASGSNHSGSKWPVSVFRKCTSKIF 396
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.138 0.457
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,045,219
Number of Sequences: 36976
Number of extensions: 139190
Number of successful extensions: 712
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 702
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 704
length of query: 189
length of database: 9,014,727
effective HSP length: 91
effective length of query: 98
effective length of database: 5,649,911
effective search space: 553691278
effective search space used: 553691278
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC148098.5


