

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147960.5 - phase: 0
(617 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
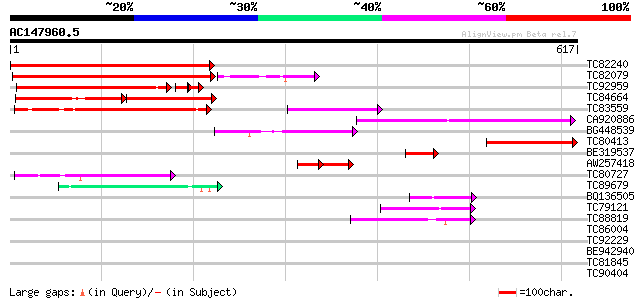
Score E
Sequences producing significant alignments: (bits) Value
TC82240 weakly similar to GP|11994399|dbj|BAB02358. contains sim... 451 e-127
TC82079 weakly similar to GP|11994399|dbj|BAB02358. contains sim... 221 2e-64
TC92959 weakly similar to GP|11994399|dbj|BAB02358. contains sim... 222 2e-62
TC84664 weakly similar to GP|15027837|gb|AAK76449.1 putative pro... 143 1e-61
TC83559 similar to GP|11994399|dbj|BAB02358. contains similarity... 204 8e-53
CA920886 similar to GP|11994399|db contains similarity to recept... 182 3e-46
BG448539 similar to GP|11994399|dbj contains similarity to recep... 93 3e-19
TC80413 similar to GP|11994399|dbj|BAB02358. contains similarity... 85 7e-17
BE319537 similar to GP|11994399|dbj contains similarity to recep... 80 2e-15
AW257418 similar to GP|11994399|dbj contains similarity to recep... 70 2e-12
TC80727 similar to GP|15982870|gb|AAL09782.1 At1g21880/T26F17_5 ... 64 1e-10
TC89679 weakly similar to PIR|C84747|C84747 probable protein kin... 50 2e-06
BQ136505 similar to PIR|T05754|T05 S-receptor kinase (EC 2.7.1.-... 46 5e-05
TC79121 similar to GP|9651971|gb|AAF91337.1| Pti1 kinase-like pr... 44 2e-04
TC88819 similar to GP|9758951|dbj|BAB09338.1 serine/threonine-sp... 42 5e-04
TC86004 weakly similar to GP|22758281|gb|AAN05509.1 Unknown prot... 41 0.002
TC92229 similar to PIR|B86369|B86369 hypothetical protein AAC980... 40 0.003
BE942940 weakly similar to GP|13506749|gb receptor-like protein ... 38 0.013
TC81845 similar to PIR|T01617|T01617 probable protein kinase [im... 37 0.017
TC90404 similar to GP|21593619|gb|AAM65586.1 receptor protein ki... 35 0.11
>TC82240 weakly similar to GP|11994399|dbj|BAB02358. contains similarity to
receptor protein kinase~gene_id:MIL23.20 {Arabidopsis
thaliana}, partial (21%)
Length = 856
Score = 451 bits (1160), Expect = e-127
Identities = 220/222 (99%), Positives = 220/222 (99%)
Frame = +1
Query: 1 MKPIKFRLSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKP 60
MKPIKFRLSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKP
Sbjct: 34 MKPIKFRLSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKP 213
Query: 61 EDIVSYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYS 120
EDIVSYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYS
Sbjct: 214 EDIVSYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYS 393
Query: 121 NLTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELIS 180
NLTTSEWLQNFNSYPSNDIPDTGTLNVTVN SCGNSDVSKDYGLFITYPLRPEDSLELIS
Sbjct: 394 NLTTSEWLQNFNSYPSNDIPDTGTLNVTVNWSCGNSDVSKDYGLFITYPLRPEDSLELIS 573
Query: 181 NKTEIDAELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHT 222
NKTEIDAELLQKYNPGV FSQGSGLVYIPGKDQNRNYVPFHT
Sbjct: 574 NKTEIDAELLQKYNPGVKFSQGSGLVYIPGKDQNRNYVPFHT 699
>TC82079 weakly similar to GP|11994399|dbj|BAB02358. contains similarity to
receptor protein kinase~gene_id:MIL23.20 {Arabidopsis
thaliana}, partial (23%)
Length = 1057
Score = 221 bits (562), Expect(2) = 2e-64
Identities = 109/223 (48%), Positives = 148/223 (65%), Gaps = 2/223 (0%)
Frame = +2
Query: 4 IKFRLSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLV-TKPED 62
+K L + L F ESKC K C++ALASYY+ L +SN MQS +V T D
Sbjct: 35 LKNGLLLFILFLDCVFFKVESKCVKGCDVALASYYIIPSIQLRNISNFMQSKIVLTNSFD 214
Query: 63 IV-SYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSN 121
++ SYN D + +K + S+TR+NVPFPC+CI EFLGH+F+Y D Y +A+ Y++
Sbjct: 215 VIMSYNRDVVFDKSGLISYTRINVPFPCECIGGEFLGHVFEYTTKEGDDYDLIANTYYAS 394
Query: 122 LTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELISN 181
LTT E L+ FNSY N IP +NVTV CSCGNS +SKDYGLF+TYPLR +D+L I+
Sbjct: 395 LTTVELLKKFNSYDPNHIPVKAKINVTVICSCGNSQISKDYGLFVTYPLRSDDTLAKIAT 574
Query: 182 KTEIDAELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHTRS 224
K +D L+Q +N NFS GSG+V+IPG+DQN ++ P ++R+
Sbjct: 575 KAGLDEGLIQNFNQDANFSIGSGIVFIPGRDQNGHFFPLYSRT 703
Score = 43.9 bits (102), Expect(2) = 2e-64
Identities = 45/120 (37%), Positives = 58/120 (47%), Gaps = 9/120 (7%)
Frame = +1
Query: 227 WSYNWNICWSSSRTNSIIILHLCYVLPKEEDKE--ARISFGRIQRDLRSS*E**SLR*CY 284
+SY NI W T+ I L++C +LPKE +E +F I L+*C
Sbjct: 730 YSYGRNI-W----TSIICYLYICQILPKEGRREN*TSTNF*GIFNSR-------CLK*CR 873
Query: 285 IWNFRFCESC*YDW-------N*GGKIRRVFI*RTSQCHK*LQHG*QNRSRWFR*SLLRR 337
I NFR +C W + GGK+ RV++ R SQ K* Q G* N SRW LL R
Sbjct: 874 I*NFRIQWAC--YW*CCRPYRHYGGKVDRVYVSRISQGDK*FQLG**NWSRWIWSCLLCR 1047
>TC92959 weakly similar to GP|11994399|dbj|BAB02358. contains similarity to
receptor protein kinase~gene_id:MIL23.20 {Arabidopsis
thaliana}, partial (11%)
Length = 705
Score = 222 bits (565), Expect(3) = 2e-62
Identities = 103/169 (60%), Positives = 133/169 (77%)
Frame = +2
Query: 8 LSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKPEDIVSYN 67
L + LAS FI ESKC+K C++ALA++Y+ +NLTY+S+IM+SN+ T+PEDIV Y+
Sbjct: 92 LPLFSIFLASIPFITESKCTKGCSLALANFYVSQGSNLTYISSIMRSNIQTRPEDIVEYS 271
Query: 68 TDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTTSEW 127
+ I +KD VQ+ R+NVPFPCDCI +FLGH F Y V T DTY +VA+NNY+NLT EW
Sbjct: 272 REIIPSKDSVQAGQRLNVPFPCDCIDGQFLGHKFSYDVETGDTYETVATNNYANLTNVEW 451
Query: 128 LQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSL 176
L+ FN+YP NDIPDTGTLNVTVNCSCG++DV +Y LF+TYPLRP ++L
Sbjct: 452 LRRFNTYPPNDIPDTGTLNVTVNCSCGDADVG-NYALFVTYPLRPGETL 595
Score = 30.8 bits (68), Expect(3) = 2e-62
Identities = 12/17 (70%), Positives = 15/17 (87%)
Frame = +1
Query: 195 PGVNFSQGSGLVYIPGK 211
PGVNF+QGSG+V+ GK
Sbjct: 652 PGVNFNQGSGIVFXXGK 702
Score = 26.2 bits (56), Expect(3) = 2e-62
Identities = 9/19 (47%), Positives = 14/19 (73%)
Frame = +3
Query: 181 NKTEIDAELLQKYNPGVNF 199
N +++D+ LLQ+YNP F
Sbjct: 609 NSSKVDSSLLQRYNPWCQF 665
>TC84664 weakly similar to GP|15027837|gb|AAK76449.1 putative protein kinase
{Arabidopsis thaliana}, partial (6%)
Length = 708
Score = 143 bits (360), Expect(2) = 1e-61
Identities = 65/98 (66%), Positives = 79/98 (80%)
Frame = +1
Query: 128 LQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELISNKTEIDA 187
+QNFN Y +IPD + VTVNCSCGN +VS DYGLFITYPLR ED+LE I+ EI+A
Sbjct: 391 MQNFNVYRPTNIPDFAMIKVTVNCSCGNKEVSMDYGLFITYPLRSEDTLESIAKGAEIEA 570
Query: 188 ELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHTRSI 225
ELLQ+YNPGVNFS+GSGLV+IPGKDQN +Y+P H ++
Sbjct: 571 ELLQRYNPGVNFSKGSGLVFIPGKDQNGSYLPLHPSTV 684
Score = 112 bits (280), Expect(2) = 1e-61
Identities = 57/121 (47%), Positives = 81/121 (66%)
Frame = +3
Query: 7 RLSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKPEDIVSY 66
++SF F++L ESKC++ C++ALASY L +NLTY+SNIM+SN+++KP+DI+
Sbjct: 57 KISFSFIVLILLIASTESKCNEGCSLALASYTLNHVSNLTYISNIMKSNVLSKPQDIIIN 236
Query: 67 NTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTTSE 126
N NK R NVPFPC+CI+ EFL + F Y++ +TY SVA ++SNLTT
Sbjct: 237 ND---KNK-------RANVPFPCNCINGEFLAYTFLYELQPGETYTSVAEESFSNLTTDV 386
Query: 127 W 127
W
Sbjct: 387 W 389
>TC83559 similar to GP|11994399|dbj|BAB02358. contains similarity to
receptor protein kinase~gene_id:MIL23.20 {Arabidopsis
thaliana}, partial (21%)
Length = 1217
Score = 204 bits (519), Expect = 8e-53
Identities = 105/215 (48%), Positives = 145/215 (66%), Gaps = 1/215 (0%)
Frame = +2
Query: 6 FRLSFL-FMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKPEDIV 64
F L+FL F+LL K+ C+ C++ALASYY+++ TNLTY+SN+ +I+
Sbjct: 17 FILNFLLFVLLIMKTV---RSCNSGCDLALASYYIEEGTNLTYISNLFNQ----PTSEIL 175
Query: 65 SYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTT 124
YN + I N D +QS TR+N+PF CDC+ FLGH F Y++ + Y +VA+ YSNLTT
Sbjct: 176 KYNPN-IQNPDTIQSHTRLNIPFTCDCLSGLFLGHTFSYKLKEGENYKAVANGYYSNLTT 352
Query: 125 SEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELISNKTE 184
++L NSYP+ DIP +NVTVNCSCG+ DVSKDYGLF+TYPLR DSL I+ ++
Sbjct: 353 IDFLIRVNSYPATDIPAGTVINVTVNCSCGDRDVSKDYGLFLTYPLRNGDSLPGIAMESG 532
Query: 185 IDAELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVP 219
+ EL+++YNP NF G LV++P KD+N N+ P
Sbjct: 533 VPVELVKRYNPASNFRAGE-LVFLPAKDENGNFPP 634
Score = 73.2 bits (178), Expect = 3e-13
Identities = 48/103 (46%), Positives = 60/103 (57%)
Frame = +2
Query: 303 KIRRVFI*RTSQCHK*LQHG*QNRSRWFR*SLLRRAEWRESCNKKDGHESNKRISC*VEG 362
++ V I* SQ + + HG* NRSRW +L RA ESCNKKDG+ S KRI * +G
Sbjct: 902 QVSGVLI*GVSQSY*RI*HG*YNRSRWLWIGILCRATK*ESCNKKDGYASIKRIPS*TKG 1081
Query: 363 VDACSSREPGTLDRILC*GIPFPCL*IH**WKLRPTSTQFRWR 405
D S E G + ILC*G+ L*+H* W+ + TS R R
Sbjct: 1082 SDTRPSLELGAANWILC*GLFIFGL*VH*EWQFKSTSAWRRLR 1210
>CA920886 similar to GP|11994399|db contains similarity to receptor protein
kinase~gene_id:MIL23.20 {Arabidopsis thaliana}, partial
(25%)
Length = 818
Score = 182 bits (462), Expect = 3e-46
Identities = 118/239 (49%), Positives = 141/239 (58%), Gaps = 1/239 (0%)
Frame = -3
Query: 378 LC*GIPFPCL*IH**WKLRPTSTQFRWRTFVMVYQG*NCSGFSQRS*IHSRTYRPYIYSS 437
+ * + FP +* +* WKL T +R+RT MV +CS FSQR IHS T+ +Y S
Sbjct: 801 MA*RVTFPRI*TY*QWKLGSIFTWYRYRTITMV**SADCSRFSQRPRIHS*THCACLYPS 622
Query: 438 RHKVRKYSIRQKLLCKGCRLRINQAD*CWNFISSHC*YGWHIWLHATRIC-IW*CFFKNR 496
R K+ KY RQK KGC ++Q * W +SH G +IW+HATRIC IW CF KNR
Sbjct: 621 RRKISKYIDRQKFAWKGC*FWLDQTY*SWKLDTSHSSCG-NIWIHATRICSIWRCFSKNR 445
Query: 497 CICFWSSSL*TNFC*SSCNNG*RFWC*FKGPCSFV*RSF*SATSYRRS*KTGGS*AWR*L 556
CICFW SL*T +C C C KG C+ V*RS S S+RRS K GGS*A R L
Sbjct: 444 CICFWRCSL*TYYCKECCPEDR*ICCRIKGSCTIV*RSTSSNGSFRRSSKIGGS*A*RKL 265
Query: 557 SN*SCIQDGTTCESMYK**SAATSKYEFCCSCSYHTNFNY*GLGYYFYIQKSKSCKSNV 615
S+* C QDG+T ESMY+ SA T K+E CSY T * L +I KS S KS V
Sbjct: 264 SH*FCSQDGSTWESMYERQSATTPKHEIYSCCSYDTFITN*RL***LFI*KSISHKSVV 88
>BG448539 similar to GP|11994399|dbj contains similarity to receptor protein
kinase~gene_id:MIL23.20 {Arabidopsis thaliana}, partial
(13%)
Length = 537
Score = 93.2 bits (230), Expect = 3e-19
Identities = 67/159 (42%), Positives = 86/159 (53%), Gaps = 3/159 (1%)
Frame = +1
Query: 223 RSIRWSYNWNICWSSSRTNSIIILHLCYVLPKEEDKE---ARISFGRIQRDLRSS*E**S 279
RS Y WNICWS+S +I+I + +L KEE+ E + FG Q ++ S+
Sbjct: 115 RSRNCGYYWNICWSTSSAFTIVIFCVY*ILSKEEE*EDMGEKFDFG*FQDEVSSN----- 279
Query: 280 LR*CYIWNFRFCESC*YDWN*GGKIRRVFI*RTSQCHK*LQHG*QNRSRWFR*SLLRRAE 339
W + + GGKIRR+FI RT CHK* QHG* N RWF SLL RAE
Sbjct: 280 ------WYQ-------HRKHYGGKIRRIFIQRTIDCHK*FQHG**NW*RWFWRSLLCRAE 420
Query: 340 WRESCNKKDGHESNKRISC*VEGVDACSSREPGTLDRIL 378
ES + +D E KRI *+E +++CSS E G D IL
Sbjct: 421 RTESRH*EDEDEGIKRILR*IESLNSCSSFELGRFDWIL 537
>TC80413 similar to GP|11994399|dbj|BAB02358. contains similarity to
receptor protein kinase~gene_id:MIL23.20 {Arabidopsis
thaliana}, partial (10%)
Length = 662
Score = 85.1 bits (209), Expect = 7e-17
Identities = 50/99 (50%), Positives = 62/99 (62%)
Frame = +2
Query: 519 RFWC*FKGPCSFV*RSF*SATSYRRS*KTGGS*AWR*LSN*SCIQDGTTCESMYK**SAA 578
R C FKG C FV*RS *SA SYRRS + GS AW *L QDG+ +S++ S+
Sbjct: 2 RICCRFKGTCWFV*RSX*SA*SYRRSSQNSGSKAWG*LPCRFSSQDGSISQSVHTGKSST 181
Query: 579 TSKYEFCCSCSYHTNFNY*GLGYYFYIQKSKSCKSNVWK 617
S+YE C CS+ T FNY LG +F ++KSK C+S V K
Sbjct: 182 ASEYEIHCGCSHDTFFNYGRLGRWFLLRKSKPCESYVRK 298
>BE319537 similar to GP|11994399|dbj contains similarity to receptor protein
kinase~gene_id:MIL23.20 {Arabidopsis thaliana}, partial
(5%)
Length = 110
Score = 80.5 bits (197), Expect = 2e-15
Identities = 36/36 (100%), Positives = 36/36 (100%)
Frame = +1
Query: 431 RPYIYSSRHKVRKYSIRQKLLCKGCRLRINQAD*CW 466
RPYIYSSRHKVRKYSIRQKLLCKGCRLRINQAD*CW
Sbjct: 1 RPYIYSSRHKVRKYSIRQKLLCKGCRLRINQAD*CW 108
>AW257418 similar to GP|11994399|dbj contains similarity to receptor protein
kinase~gene_id:MIL23.20 {Arabidopsis thaliana}, partial
(9%)
Length = 458
Score = 70.1 bits (170), Expect = 2e-12
Identities = 31/37 (83%), Positives = 34/37 (91%)
Frame = +3
Query: 338 AEWRESCNKKDGHESNKRISC*VEGVDACSSREPGTL 374
+ + ESCNKKDGHESNKRISC*VEGVDACSSREPG +
Sbjct: 285 SNYTESCNKKDGHESNKRISC*VEGVDACSSREPGNM 395
Score = 60.1 bits (144), Expect = 2e-09
Identities = 27/28 (96%), Positives = 28/28 (99%)
Frame = +2
Query: 314 QCHK*LQHG*QNRSRWFR*SLLRRAEWR 341
QCHK*LQHG*QNRSRWFR*SLLR+AEWR
Sbjct: 2 QCHK*LQHG*QNRSRWFR*SLLRKAEWR 85
>TC80727 similar to GP|15982870|gb|AAL09782.1 At1g21880/T26F17_5
{Arabidopsis thaliana}, partial (69%)
Length = 1019
Score = 64.3 bits (155), Expect = 1e-10
Identities = 50/180 (27%), Positives = 85/180 (46%), Gaps = 5/180 (2%)
Frame = +3
Query: 6 FRLSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKPEDIVS 65
F L L + SKS I S TCN +L Y L D ++ +S++ Q + P +++
Sbjct: 126 FLLFKLLITTTSKSTIEPCTTSDTCN-SLLGYTLYTDLKVSELSSLFQID----PISLLT 290
Query: 66 YNTDTITNKD----FVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSN 121
N+ I+ D + S + +P C CI Y++ DT S+A + Y
Sbjct: 291 ANSIDISYPDVEHHILPSKLYLKIPIQCSCIDGIRKSVSTNYKIRPSDTLSSIADSIYGG 470
Query: 122 LTTSEWLQNFNSYPSNDIPDTG-TLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELIS 180
L +S+ L+ NS ++ D G L V + C+C N + ++++Y ++P DSL I+
Sbjct: 471 LVSSDQLREANSVTDPNVLDVGQNLVVPLPCTCFNGSDNGLPAIYMSYVVQPLDSLNNIA 650
>TC89679 weakly similar to PIR|C84747|C84747 probable protein kinase
[imported] - Arabidopsis thaliana, partial (13%)
Length = 880
Score = 50.4 bits (119), Expect = 2e-06
Identities = 47/205 (22%), Positives = 83/205 (39%), Gaps = 27/205 (13%)
Frame = +3
Query: 54 SNLVTKPEDIVSYNTDTITNKDFVQSFTRVNVPFPCDCIHDE-FLGHIFQYQVA-TKDTY 111
S+L+ ++S ++ I+ + + T + VP C C ++ + H Y + T +TY
Sbjct: 210 SHLLNSSASLIS-QSNNISTVQTLPTDTIITVPINCTCSNNNTYYQHNTSYTIQNTGETY 386
Query: 112 LSVASNNYSNLTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKD-YGLFITYPL 170
+VA+N Y L+T + L N Y I L V + C+C S + + +TY +
Sbjct: 387 FTVANNTYQALSTCQALIAQNPYNERKIVRGNNLTVPLRCACPTKKQSDEGFKYLLTYLV 566
Query: 171 RPEDSLELISNKTEIDAELLQKYNPGVNFSQGSGLVY-----IPGKDQNRN--------- 216
+S+ I+ +D + + + N S S + Y IP K++ R
Sbjct: 567 SEGESVSSIAEIFNVDPQSINEAN---ELSSTSFIFYFTPLLIPLKNERRKK*SNRLHHR 737
Query: 217 ----------YVPFHTRSIRWSYNW 231
FH S +W W
Sbjct: 738 ILRLRRHRRXKTEFHPPSTKWVIRW 812
>BQ136505 similar to PIR|T05754|T05 S-receptor kinase (EC 2.7.1.-) M4I22.110
precursor - Arabidopsis thaliana, partial (23%)
Length = 618
Score = 45.8 bits (107), Expect = 5e-05
Identities = 29/75 (38%), Positives = 42/75 (55%), Gaps = 2/75 (2%)
Frame = +1
Query: 436 SSRHKVRKYSIRQKLLCKGCRLRINQAD*CWNFI-SSHC*YGWHIWLHATRIC-IW*CFF 493
S R KV++Y R+ K RLR + +D W S + GW+IWLHA+R+C W
Sbjct: 322 SQRFKVKQYLTRRPYESKNLRLRPS-SDILWRSS*SQNKEIGWNIWLHASRVCRAWSLLH 498
Query: 494 KNRCICFWSSSL*TN 508
+ RC+ F S+S+ N
Sbjct: 499 EIRCV*FRSNSIGDN 543
>TC79121 similar to GP|9651971|gb|AAF91337.1| Pti1 kinase-like protein
{Glycine max}, partial (96%)
Length = 1283
Score = 43.5 bits (101), Expect = 2e-04
Identities = 35/106 (33%), Positives = 50/106 (47%), Gaps = 2/106 (1%)
Frame = +2
Query: 404 WRTFVMVYQG*NCSGFSQRS*IHSRTYRPYIYSSRHKVRKYSIRQKLLCKGCRL-RINQA 462
W + M + *NC SQR+*I S R S H ++ ++ + CK C + +
Sbjct: 632 WSSSFMG*KS*NCCWSSQRT*ISS*KGRGSYSPSLH*IQ*HTPIRGRRCKDC*F*SVKSS 811
Query: 463 D*CWNFISSHC*YGWHIWLHATRIC-IW*CFFKNRCICFWSSSL*T 507
*C SS W+ WL +RIC W FK C+ FWS ++ T
Sbjct: 812 P*CCR-TSSFYPCSWNFWLSCSRICNDWKPLFKE*CLQFWSYTVGT 946
>TC88819 similar to GP|9758951|dbj|BAB09338.1 serine/threonine-specific
protein kinase-like protein {Arabidopsis thaliana},
partial (86%)
Length = 1231
Score = 42.4 bits (98), Expect = 5e-04
Identities = 45/142 (31%), Positives = 61/142 (42%), Gaps = 6/142 (4%)
Frame = +3
Query: 372 GTLDRILC*-GIPFPCL*IH**WKLRPTSTQFRWRTFVMVYQG*NCSGFSQRS*IHSRTY 430
G ILC G + CL +H * +L Q R + ++G S+R + S
Sbjct: 564 GQFGWILCRKGSTYACLCLHE*RQLGFPLVQ*REWESGLGFEGSYSFRCSKRLGVSS*WG 743
Query: 431 RPYIYSSRHKVRKYSIRQKLLCKGCRLRINQAD*CWNFISSH----C*YGWHIWLHATRI 486
+SSRH++ +YS+ GC * W F C Y WH+WL + I
Sbjct: 744 SSSCHSSRHQI*QYSLGSVHASPGC--------*FWTFKRRDGG*TCCYSWHLWLSGS*I 899
Query: 487 CIW*CFF-KNRCICFWSSSL*T 507
I K RCI FWS +L*T
Sbjct: 900 YIIRDIHQKKRCIQFWSVAL*T 965
>TC86004 weakly similar to GP|22758281|gb|AAN05509.1 Unknown protein {Oryza
sativa (japonica cultivar-group)}, partial (28%)
Length = 1373
Score = 40.8 bits (94), Expect = 0.002
Identities = 22/77 (28%), Positives = 39/77 (50%), Gaps = 2/77 (2%)
Frame = +2
Query: 83 VNVPFPCDCIH-DEFLGHIFQYQVATKDTYLSVASNNYSNLTTSEWLQNFNSYP-SNDIP 140
+ VPFPC C + H+ +Y++ DT ++A ++ L + +Q N P +N+I
Sbjct: 401 IKVPFPCKCSNRTGTSNHVPRYKIVPGDTLDAIARVRFAGLVKYQQIQTANKIPDANNIT 580
Query: 141 DTGTLNVTVNCSCGNSD 157
T+ + + CSC D
Sbjct: 581 AGATIWIPLPCSCDPVD 631
>TC92229 similar to PIR|B86369|B86369 hypothetical protein AAC98010.1
[imported] - Arabidopsis thaliana, partial (44%)
Length = 1347
Score = 39.7 bits (91), Expect = 0.003
Identities = 43/152 (28%), Positives = 68/152 (44%), Gaps = 2/152 (1%)
Frame = +1
Query: 365 ACSSREPGTLDRIL-C*GIPFPCL*IH**WKLRPTSTQFRWRTFVMVYQG*NCSGFSQRS 423
+CSS G +D +L * P L*+ *W+ T F + + + + + S
Sbjct: 343 SCSSSTFGIVDWLLYS*ATKSPHL*VCS*WES*STFA*KSMECFGLAQEDEDSNWCCKGS 522
Query: 424 *IHSRTYRPYIYSSRHKVRKYSIRQKLLCKGCRLRINQAD*CWNFISSHC*YGWHIWLHA 483
I +R +P YS R+ + +Y+ L CRL Q + + + H G+ IWL+
Sbjct: 523 SISARRLQPKDYSQRY*IFEYTFG*FL*GTSCRLWTCQTNRRYKYSCIH*GNGY-IWLYG 699
Query: 484 TRIC-IW*CFFKNRCICFWSSSL*TNFC*SSC 514
+R+C W RC WS S *T + +C
Sbjct: 700 SRVCNKWKTNG*IRCFLIWSCSS*TCYWKETC 795
>BE942940 weakly similar to GP|13506749|gb receptor-like protein kinase 6
{Arabidopsis thaliana}, partial (23%)
Length = 466
Score = 37.7 bits (86), Expect = 0.013
Identities = 15/27 (55%), Positives = 20/27 (73%), Gaps = 1/27 (3%)
Frame = +3
Query: 477 WHIWLHATRIC-IW*CFFKNRCICFWS 502
W+IWL+ +RIC +W F K RC+ FWS
Sbjct: 339 WNIWLYVSRICNVWTIF*KIRCL*FWS 419
>TC81845 similar to PIR|T01617|T01617 probable protein kinase [imported] -
Arabidopsis thaliana, partial (42%)
Length = 762
Score = 37.4 bits (85), Expect = 0.017
Identities = 19/46 (41%), Positives = 26/46 (56%), Gaps = 1/46 (2%)
Frame = +1
Query: 470 SSHC*YGWHIWLHATRICI-W*CFFKNRCICFWSSSL*TNFC*SSC 514
S +C +IW TRI + W C +NRCICFW ++F * +C
Sbjct: 181 SLNCSNRRNIWTFGTRILLAWSCG*ENRCICFWCVLARSHFW*ETC 318
>TC90404 similar to GP|21593619|gb|AAM65586.1 receptor protein kinase-like
protein {Arabidopsis thaliana}, partial (52%)
Length = 1252
Score = 34.7 bits (78), Expect = 0.11
Identities = 19/45 (42%), Positives = 26/45 (57%)
Frame = +2
Query: 303 KIRRVFI*RTSQCHK*LQHG*QNRSRWFR*SLLRRAEWRESCNKK 347
K+ VFI RT C+K*LQ + RWF L+R+ +R CN +
Sbjct: 38 KLEEVFIQRTPSCYK*LQQQKLSWKRWFW-KCLQRSSFRWHCNSR 169
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.351 0.153 0.577
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,935,304
Number of Sequences: 36976
Number of extensions: 454632
Number of successful extensions: 4733
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 2581
Number of HSP's successfully gapped in prelim test: 214
Number of HSP's that attempted gapping in prelim test: 1994
Number of HSP's gapped (non-prelim): 2982
length of query: 617
length of database: 9,014,727
effective HSP length: 102
effective length of query: 515
effective length of database: 5,243,175
effective search space: 2700235125
effective search space used: 2700235125
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147960.5


