

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147959.10 + phase: 1 /pseudo
(378 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
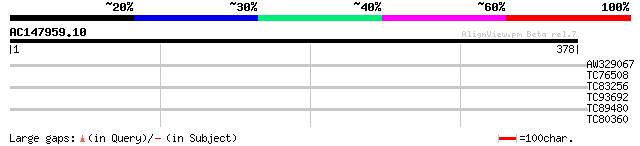
AW329067 similar to GP|9955519|emb| putative protein {Arabidopsi... 39 0.003
TC76508 GP|3293555|gb|AAC25775.1| chlorophyll a/b binding protei... 29 3.4
TC83256 similar to GP|21537173|gb|AAM61514.1 unknown {Arabidopsi... 28 5.8
TC93692 similar to GP|20161344|dbj|BAB90268. contains ESTs AU086... 28 7.6
TC89480 similar to GP|20260684|gb|AAM13240.1 unknown protein {Ar... 28 7.6
TC80360 PIR|T01970|T01970 probable wound-induced protein T9A4.6 ... 27 9.9
>AW329067 similar to GP|9955519|emb| putative protein {Arabidopsis thaliana},
partial (6%)
Length = 672
Score = 39.3 bits (90), Expect = 0.003
Identities = 19/43 (44%), Positives = 26/43 (60%)
Frame = +2
Query: 297 YPYMEAESKDLAMEDVDILLNHYKDLVAKYTIICKAINYLSMS 339
+PYM A DL + DV+ LL +YK LV+KY + K + S S
Sbjct: 197 FPYMFASVGDLTVSDVEDLLKNYKRLVSKYVCLSKGLGVSSSS 325
>TC76508 GP|3293555|gb|AAC25775.1| chlorophyll a/b binding protein {Medicago
sativa}, complete
Length = 1190
Score = 28.9 bits (63), Expect = 3.4
Identities = 24/96 (25%), Positives = 43/96 (44%)
Frame = +3
Query: 69 LLQTFLKPEHLDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRVINNLLLNA 128
+L T H +PP L E+ W L L I + + I+ CC + LN
Sbjct: 813 ILPTQSTTTHGPMPPTLFPESEWSLC--FLLCILSCNVLPAGM*IIIMCC-----MQLNV 971
Query: 129 AMSEYVPAGADDFIPVLIYVTIKARLALLVQTSNSL 164
++ YV + IP+ +++TI A L+++ ++
Sbjct: 972 SLFFYVNLLSQSHIPIFMFLTIYATKIELIKSLKNI 1079
>TC83256 similar to GP|21537173|gb|AAM61514.1 unknown {Arabidopsis
thaliana}, partial (17%)
Length = 531
Score = 28.1 bits (61), Expect = 5.8
Identities = 16/53 (30%), Positives = 23/53 (43%), Gaps = 8/53 (15%)
Frame = -1
Query: 317 NHYKDLVAKYTIIC--------KAINYLSMSEKEPLLHQLEMQGEGSMLSECH 361
NH K + K T+ C ++ NY + LHQL+ GE + CH
Sbjct: 339 NHNKPYIIKQTLTCDSRVPAIPRS*NYYQKTLFTCNLHQLQQHGEACFYACCH 181
>TC93692 similar to GP|20161344|dbj|BAB90268. contains ESTs AU086067(S4642)
D41819(S4642)~similar to Arabidopsis thaliana chromosome
1 At1g49040~, partial (7%)
Length = 626
Score = 27.7 bits (60), Expect = 7.6
Identities = 12/35 (34%), Positives = 18/35 (51%)
Frame = -2
Query: 198 PSQLMKSNLRSACKQPNWPRKSQVNYTLHVKLNKK 232
P QL A QP+W R+ Q NY + + ++K
Sbjct: 214 P*QLFSVGFCKAANQPSWTRRCQKNYQVSLGADEK 110
>TC89480 similar to GP|20260684|gb|AAM13240.1 unknown protein {Arabidopsis
thaliana}, partial (43%)
Length = 1068
Score = 27.7 bits (60), Expect = 7.6
Identities = 17/47 (36%), Positives = 27/47 (57%), Gaps = 1/47 (2%)
Frame = +1
Query: 300 MEAESKDLAMEDVDILLNHYKDLVAKYTIICKAINYLSMSEK-EPLL 345
M E + +E+ + LLN +DL+A YT K++ L SE EP++
Sbjct: 520 MTVEGQKKKLEESEQLLNQRRDLIANYT---KSVEELVRSEP*EPVI 651
>TC80360 PIR|T01970|T01970 probable wound-induced protein T9A4.6 -
Arabidopsis thaliana, partial (21%)
Length = 679
Score = 27.3 bits (59), Expect = 9.9
Identities = 15/46 (32%), Positives = 25/46 (53%)
Frame = +3
Query: 37 LEKYIMTKLFSRTFAASPEDAKIDHEISEKISLLQTFLKPEHLDIP 82
L+++I K+FS+ F +K+DH+ EK L+ T E+ P
Sbjct: 129 LKRFIRAKIFSKLFQTWRRQSKMDHK--EKTLLVLTKAPQEYNTFP 260
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.328 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,536,308
Number of Sequences: 36976
Number of extensions: 152402
Number of successful extensions: 956
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 950
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 955
length of query: 378
length of database: 9,014,727
effective HSP length: 98
effective length of query: 280
effective length of database: 5,391,079
effective search space: 1509502120
effective search space used: 1509502120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC147959.10


