

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147714.2 - phase: 0 /pseudo
(971 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
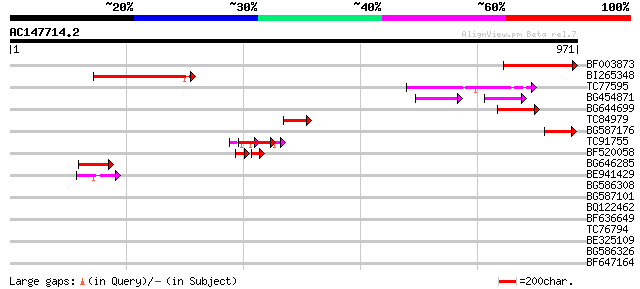
Score E
Sequences producing significant alignments: (bits) Value
BF003873 similar to GP|14715222|em putative polyprotein {Cicer a... 228 7e-60
BI265348 196 3e-50
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 96 7e-20
BG454871 weakly similar to GP|10140673|g putative gag-pol polypr... 72 9e-17
BG644699 similar to PIR|T07863|T078 probable polyprotein - pinea... 65 1e-10
TC84979 63 6e-10
BG587176 weakly similar to PIR|G84493|G84 probable retroelement ... 55 1e-07
TC91755 weakly similar to PIR|G75077|G75077 hypothetical protein... 55 1e-07
BF520058 40 6e-05
BG646285 46 8e-05
BE941429 weakly similar to GP|12323718|gb| hypothetical protein;... 43 7e-04
BG586308 weakly similar to PIR|F84528|F8 probable retroelement p... 34 0.24
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 33 0.53
BQ122462 33 0.53
BF636649 similar to GP|21628724|emb OSJNBa0033H08.7 {Oryza sativ... 32 0.91
TC76794 similar to GP|7416846|dbj|BAA94084.1 NAD-dependent sorbi... 30 4.5
BE325109 similar to PIR|T04011|T040 hypothetical protein T5L19.2... 30 5.9
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 30 5.9
BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis t... 29 7.7
>BF003873 similar to GP|14715222|em putative polyprotein {Cicer arietinum},
partial (82%)
Length = 559
Score = 228 bits (582), Expect = 7e-60
Identities = 114/126 (90%), Positives = 117/126 (92%)
Frame = +2
Query: 846 GVXRALKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHV 905
GV RALKSKKLT +FIGPYQI ERVGTVAYRVGLPPHL NLH+VFHVSQLRKYVPDPSHV
Sbjct: 2 GVGRALKSKKLTVRFIGPYQISERVGTVAYRVGLPPHLLNLHDVFHVSQLRKYVPDPSHV 181
Query: 906 IQSDDVQVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRVVWTGATSESLTWELESKMLES 965
IQSDDVQVRDNLTVETLPVRIDDRKVK LRGKEIPLVRVVW A ESLTWELESKM+ES
Sbjct: 182 IQSDDVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWDRANGESLTWELESKMVES 361
Query: 966 YPELFA 971
YPELFA
Sbjct: 362 YPELFA 379
>BI265348
Length = 556
Score = 196 bits (499), Expect = 3e-50
Identities = 121/181 (66%), Positives = 132/181 (72%), Gaps = 7/181 (3%)
Frame = +1
Query: 144 IQLTFGND*RRR--EL*VWGFQTNEN*YNK-RSLGIVGFI*ITSPSQTKLKAYNVMLDLI 200
IQ T ND* + + VW F N K +SLGI+G * + YN+MLDLI
Sbjct: 22 IQSTLDND*TKGL*DRKVWVF*LKVNNKRKDKSLGIIGPS**QVLHKPNY*TYNLMLDLI 201
Query: 201 SQDSLSYVPLATKPISLT*LLLDFGSFNIQAYFADLIVISSLEDRNYMSQRLQRLFRCLC 260
SQDS S+VPLATKPISLT*LLLDFGS N QAYFADLI+ISSLEDRNY+SQRL RLFRCL
Sbjct: 202 SQDSFSFVPLATKPISLT*LLLDFGSSNNQAYFADLIIISSLEDRNYISQRL*RLFRCLY 381
Query: 261 NPCLNSLV*LSKLHFRFMN*YLE*WINNYQTLHFRFHS----LIMVDPC*SGEFR*EIRV 316
N CLNSLV +SKLHFRFMN*YL *WINNYQTLHFRFHS LI V +GE R EIRV
Sbjct: 382 NLCLNSLVRISKLHFRFMN*YLR*WINNYQTLHFRFHSFDNGLIHVK---AGELRLEIRV 552
Query: 317 T 317
T
Sbjct: 553 T 555
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial
(14%)
Length = 1708
Score = 95.9 bits (237), Expect = 7e-20
Identities = 71/229 (31%), Positives = 108/229 (47%), Gaps = 6/229 (2%)
Frame = +2
Query: 680 GVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVL 739
G+P SIVSDR + RFW+ G LS++YHPQTDG +ER Q ++ +LR V
Sbjct: 563 GMPQSIVSDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQTDGGTERWNQEIQAVLRAYVC 742
Query: 740 EQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERVVLGPE---- 795
W LP ++ N ++SSIG PF +G P+ E VV E
Sbjct: 743 WSQDNWGDLLPTVQLALRNRHNSSIGATPFFVEHGYHV-DPIPTVEDTGGVVSEGEAAAQ 919
Query: 796 -IVQQTTEKVQMIQEKMKASQSRQKSYHDKRRKDLE-FQEGDHVFLRVTPMTGVXRALKS 853
+V++ + IQ ++ A+Q R ++ +KRR + +Q GD V+L V+ + K
Sbjct: 920 LLVKRMKDVTGFIQAEIVAAQQRSEASANKRRCPADRYQVGDKVWLNVSNYKSPRPSKKL 1099
Query: 854 KKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDP 902
L K Y++ V + +P ++ FHV LR+ DP
Sbjct: 1100DWLHHK----YEVTRFVTPHVVELNVP---GTVYPKFHVDLLRRAASDP 1225
>BG454871 weakly similar to GP|10140673|g putative gag-pol polyprotein {Oryza
sativa (japonica cultivar-group)}, partial (7%)
Length = 674
Score = 72.4 bits (176), Expect(2) = 9e-17
Identities = 37/81 (45%), Positives = 46/81 (56%)
Frame = +2
Query: 695 SRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEF 754
S FWK L + G+ L +SSAYHP +DGQSE + E LR + W P E+
Sbjct: 32 SNFWKQLFKLHGTILTMSSAYHP*SDGQSEALNKGXEMYLRCLMFTDPLKWSKAFPWAEY 211
Query: 755 TYNNSYHSSIGMAPFEALYGR 775
YN SY+ S M PF+ALYGR
Sbjct: 212 WYNTSYNISAAMTPFKALYGR 274
Score = 33.5 bits (75), Expect(2) = 9e-17
Identities = 22/73 (30%), Positives = 34/73 (46%), Gaps = 2/73 (2%)
Frame = +1
Query: 814 SQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGVXRALK--SKKLTPKFIGPYQILERVG 871
+Q K DK+R+ EFQ G+HV +++ P AL+ K +P F +
Sbjct: 388 AQQTMKHQADKKRRHFEFQLGEHVLVKLQPYQQSSVALRKYQKFGSPNFGSLLTVCSL*V 567
Query: 872 TVAYRVGLPPHLS 884
A+ PP+LS
Sbjct: 568 ESAFHCKSPPYLS 606
>BG644699 similar to PIR|T07863|T078 probable polyprotein - pineapple
retrotransposon dea1 (fragment), partial (5%)
Length = 231
Score = 65.1 bits (157), Expect = 1e-10
Identities = 31/74 (41%), Positives = 50/74 (66%), Gaps = 1/74 (1%)
Frame = +2
Query: 835 DHVFLRVTPMT-GVXRALKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVS 893
+ V L+V P G R K KL+ ++IGP+++++R+G VAY + LPP LS +H VFHVS
Sbjct: 2 EQVLLKVLPTERGDCRFGKRGKLSLRYIGPFEVIKRIGEVAYELALPPGLSGVHPVFHVS 181
Query: 894 QLRKYVPDPSHVIQ 907
++Y D +++I+
Sbjct: 182 MFKRYHGDGNYIIR 223
>TC84979
Length = 641
Score = 62.8 bits (151), Expect = 6e-10
Identities = 38/49 (77%), Positives = 40/49 (81%)
Frame = -1
Query: 469 PSNIHS*NKIKCNL**TILKRK*Y*YIIK*NLNKTKLKSI*K*SISDSS 517
PSNIHS*N IKCNL**TI KRK*Y*Y I NL K KL +I*K* I+DSS
Sbjct: 641 PSNIHS*NIIKCNL**TISKRK*Y*YKINLNLIKIKLNNI*K*IINDSS 495
>BG587176 weakly similar to PIR|G84493|G84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(1%)
Length = 729
Score = 55.5 bits (132), Expect = 1e-07
Identities = 26/54 (48%), Positives = 35/54 (64%)
Frame = -1
Query: 917 LTVETLPVRIDDRKVKMLRGKEIPLVRVVWTGATSESLTWELESKMLESYPELF 970
L +ET PVRI DR K +R K I +V++VW + E +TWE E++M YPE F
Sbjct: 717 LDLETRPVRILDRMEKAMRKKPIQMVKIVWDCSGREEITWETEARMKADYPEWF 556
>TC91755 weakly similar to PIR|G75077|G75077 hypothetical protein PAB1697 -
Pyrococcus abyssi (strain Orsay), partial (4%)
Length = 746
Score = 55.5 bits (132), Expect = 1e-07
Identities = 36/71 (50%), Positives = 45/71 (62%), Gaps = 7/71 (9%)
Frame = +3
Query: 392 DGWLLYL*HTLDLKSKQLH-------LG*K*SSLKSKQKAAKGTLEGGTAVPRLARAVPS 444
DG LLYL*+T+D KSKQ++ + + S+ K++K LEGGTAVPRLARAVPS
Sbjct: 48 DG*LLYL*YTID*KSKQVYS*NIPR*IVRQNRIALSQSKSSKRHLEGGTAVPRLARAVPS 227
Query: 445 F*LWGDIFLHP 455
F G L P
Sbjct: 228 FLTCGRYILFP 260
Score = 40.8 bits (94), Expect(2) = 1e-06
Identities = 30/59 (50%), Positives = 32/59 (53%), Gaps = 7/59 (11%)
Frame = +1
Query: 421 KQKAAKGTLEGGTAVPRLARAVPSF*LWGDIF-------LHPIGSKSLSFAPRLNPSNI 472
K KAAKG +F*L GDIF L PI SKSLSFAPRL+PSNI
Sbjct: 157 KAKAAKGI*RVARPCQG*HGPCQAF*LVGDIFCFPIFMFLCPICSKSLSFAPRLSPSNI 333
Score = 30.4 bits (67), Expect(2) = 1e-06
Identities = 24/61 (39%), Positives = 34/61 (55%), Gaps = 9/61 (14%)
Frame = +2
Query: 377 KQLSLIVFQVKKNICDGWL---------LYL*HTLDLKSKQLHLG*K*SSLKSKQKAAKG 427
KQ+SLIVFQVK N+ WL L +* ++ L+ + K*+S +SKQK K
Sbjct: 2 KQMSLIVFQVKMNV-**WLATLFIVHN*LKI*ASIFLEYSSVDCAPK*NSFESKQKQQKA 178
Query: 428 T 428
+
Sbjct: 179 S 181
>BF520058
Length = 616
Score = 40.0 bits (92), Expect(2) = 6e-05
Identities = 19/24 (79%), Positives = 20/24 (83%)
Frame = -1
Query: 387 KKNICDGWLLYL*HTLDLKSKQLH 410
K NICDG LLYL* T DL+SKQLH
Sbjct: 556 KMNICDG*LLYL*FTFDLESKQLH 485
Score = 25.4 bits (54), Expect(2) = 6e-05
Identities = 13/22 (59%), Positives = 15/22 (68%)
Frame = -3
Query: 414 K*SSLKSKQKAAKGTLEGGTAV 435
K*SSLKSKQK KG + T +
Sbjct: 458 K*SSLKSKQKQQKGIMGRATFI 393
>BG646285
Length = 640
Score = 45.8 bits (107), Expect = 8e-05
Identities = 32/61 (52%), Positives = 38/61 (61%), Gaps = 1/61 (1%)
Frame = -1
Query: 118 LMR*NRKCTVLPK*YKKIIIPTGSSLIQLTFGND*RRREL*VWGFQ-TNEN*YNKRSLGI 176
LM *N KCTVL K* K I+PTGS L QLT +D*+R V F EN KR+LG+
Sbjct: 295 LMG*NYKCTVLSK**K*SIVPTGSCLSQLTLVDD*KRLSCKVGVFNWLKENIIIKRALGL 116
Query: 177 V 177
+
Sbjct: 115 L 113
Score = 36.2 bits (82), Expect = 0.063
Identities = 26/53 (49%), Positives = 31/53 (58%), Gaps = 5/53 (9%)
Frame = -3
Query: 160 WGFQTNEN*YN-KRSLGIVGFI*ITSPSQTKLKAYNVMLDLISQD----SLSY 207
WGFQ E YN K+S GI+G * + K N+MLDLIS+D SLSY
Sbjct: 170 WGFQLVERKYNNKKSFGIIGSA**QILHKPKP*T*NLMLDLISKDRFLFSLSY 12
>BE941429 weakly similar to GP|12323718|gb| hypothetical protein; 28267-27009
{Arabidopsis thaliana}, partial (7%)
Length = 493
Score = 42.7 bits (99), Expect = 7e-04
Identities = 37/84 (44%), Positives = 45/84 (53%), Gaps = 8/84 (9%)
Frame = -2
Query: 115 ICPLMR*NRKCTVLPK*YKKIIIPTGS-------SLIQLTFGND*RRREL*VWGFQTNEN 167
+C LMR*NRKCTVL K* I+PTGS +I+L F ++ +GF
Sbjct: 306 LCALMR*NRKCTVLSK**N*SIVPTGSYEFNQL*IMIRLKF------YKIESFGFLIESK 145
Query: 168 *YNKR-SLGIVGFI*ITSPSQTKL 190
* KR FI*+TS SQTKL
Sbjct: 144 **KKR*KPWDYWFI*LTSSSQTKL 73
Score = 38.5 bits (88), Expect = 0.013
Identities = 21/42 (50%), Positives = 27/42 (64%)
Frame = -3
Query: 172 RSLGIVGFI*ITSPSQTKLKAYNVMLDLISQDSLSYVPLATK 213
+SLGI+G + YN+MLDL S+DS S+VPLATK
Sbjct: 128 KSLGIIGSSN*QVLHKPNY*TYNLMLDLTSKDSFSFVPLATK 3
>BG586308 weakly similar to PIR|F84528|F8 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 686
Score = 34.3 bits (77), Expect = 0.24
Identities = 20/70 (28%), Positives = 36/70 (50%)
Frame = -2
Query: 680 GVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVL 739
G+P IV+D F S ++ E +L +S +PQ++GQ+E + + + D L+ +
Sbjct: 682 GLPYEIVTDNGSHFISNKFREFCERWRIRLNTASPRYPQSNGQAEASNKIIIDGLKKRLD 503
Query: 740 EQGGTWDSHL 749
+ G W L
Sbjct: 502 LKKGCWADEL 473
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 33.1 bits (74), Expect = 0.53
Identities = 14/45 (31%), Positives = 28/45 (62%)
Frame = +2
Query: 680 GVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSE 724
GVP ++SD F ++ ++ L + G + ++++AYHPQ +S+
Sbjct: 482 GVPRVVISDGGSHFINKVFEKLLKKNGVRHKVATAYHPQKAERSK 616
>BQ122462
Length = 757
Score = 33.1 bits (74), Expect = 0.53
Identities = 27/45 (60%), Positives = 27/45 (60%), Gaps = 2/45 (4%)
Frame = -2
Query: 463 FAPRLNPSNI--HS*NKIKCNL**TILKRK*Y*YIIK*NLNKTKL 505
FAPRLNPSNI *N L** KRK Y Y *NLNKT L
Sbjct: 756 FAPRLNPSNIILEI*N---MQL**YNSKRKYYKYKYN*NLNKTIL 631
>BF636649 similar to GP|21628724|emb OSJNBa0033H08.7 {Oryza sativa}, partial
(4%)
Length = 653
Score = 32.3 bits (72), Expect = 0.91
Identities = 35/166 (21%), Positives = 66/166 (39%), Gaps = 4/166 (2%)
Frame = +1
Query: 719 TDGQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCR 778
+D Q+ +LE L EQ G + L E YN ++H + G PF+ +Y
Sbjct: 7 SDEQAGLLNHTLETHLLYFTSEQQGV*NFFLTWAECLYNTNFHRTAGCTPFKVVY---VV 177
Query: 779 TPLCWFESGERVVLGPEIVQQTTEKVQMIQEKMKASQSRQKSY----HDKRRKDLEFQEG 834
L F ++ E + +++ + + +Y H +R + +
Sbjct: 178 AHLQKFVVARDLIYRNEGLHKSST*TSFGRGTRAYEALSRPAYETC*HPRRPLSIVYTRD 357
Query: 835 DHVFLRVTPMTGVXRALKSKKLTPKFIGPYQILERVGTVAYRVGLP 880
+V P K GPYQ+++++G+VA+++ LP
Sbjct: 358 RTYEWQVLP-----------KYVA*CYGPYQVIKQIGSVAFKL*LP 462
>TC76794 similar to GP|7416846|dbj|BAA94084.1 NAD-dependent sorbitol
dehydrogenase {Prunus persica}, partial (93%)
Length = 1691
Score = 30.0 bits (66), Expect = 4.5
Identities = 12/21 (57%), Positives = 17/21 (80%)
Frame = -2
Query: 884 SNLHNVFHVSQLRKYVPDPSH 904
SNL N+FHVS + YVP+P++
Sbjct: 1690 SNLINIFHVS*FKVYVPNPTN 1628
>BE325109 similar to PIR|T04011|T040 hypothetical protein T5L19.200 -
Arabidopsis thaliana, partial (9%)
Length = 430
Score = 29.6 bits (65), Expect = 5.9
Identities = 14/43 (32%), Positives = 25/43 (57%)
Frame = +1
Query: 699 KSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQ 741
KSLQ G++++L + P+ D ERT+Q D +I + ++
Sbjct: 79 KSLQTKTGARIQLIPQHLPEGDDSKERTVQVTGDKRQIEIAQE 207
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 29.6 bits (65), Expect = 5.9
Identities = 26/64 (40%), Positives = 30/64 (46%)
Frame = +3
Query: 23 QDSSEFMRRITRRMI*SWRLWSLY*RYGDIACMVRGLRCLVITRA*SICSIRKN*I*GIE 82
Q S E MR T MI* W *R+G CMV R + + *SI +* *G
Sbjct: 126 QGS*ENMRETTPPMI*KWLR*YSP*RFGAHTCMVPRFRYIRTIKV*SIFLPSLS*T*GRG 305
Query: 83 GGSN 86
GG N
Sbjct: 306 GGWN 317
>BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana},
partial (6%)
Length = 469
Score = 29.3 bits (64), Expect = 7.7
Identities = 19/58 (32%), Positives = 29/58 (49%)
Frame = +1
Query: 809 EKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGVXRALKSKKLTPKFIGPYQI 866
E K + R K +HD+R EF+EG+ V L + + L KL + GP+Q+
Sbjct: 142 ENAKIYKERTKKWHDRRIIRREFREGELVLLFNSRL-----KLFPGKLRSHWSGPFQV 300
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.350 0.155 0.524
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,667,229
Number of Sequences: 36976
Number of extensions: 364592
Number of successful extensions: 3071
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 1258
Number of HSP's successfully gapped in prelim test: 129
Number of HSP's that attempted gapping in prelim test: 1775
Number of HSP's gapped (non-prelim): 1455
length of query: 971
length of database: 9,014,727
effective HSP length: 105
effective length of query: 866
effective length of database: 5,132,247
effective search space: 4444525902
effective search space used: 4444525902
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC147714.2


