

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147714.16 - phase: 0
(322 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
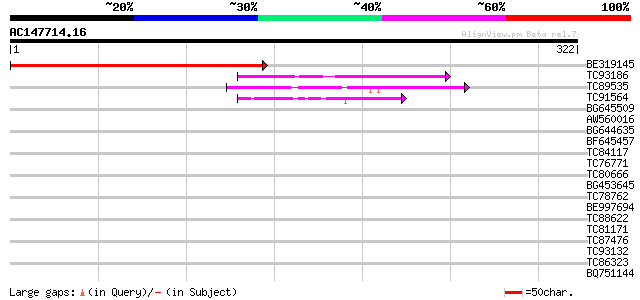
Score E
Sequences producing significant alignments: (bits) Value
BE319145 302 1e-82
TC93186 similar to PIR|A86193|A86193 hypothetical protein [impor... 94 9e-20
TC89535 weakly similar to PIR|A96808|A96808 hypothetical protein... 70 8e-13
TC91564 weakly similar to GP|6573754|gb|AAF17674.1| F28K19.1 {Ar... 64 1e-10
BG645509 similar to PIR|T47966|T47 hypothetical protein F15G16.1... 40 0.001
AW560016 similar to GP|8778713|gb T25N20.3 {Arabidopsis thaliana... 39 0.004
BG644635 weakly similar to GP|8843772|db contains similarity to ... 37 0.008
BF645457 similar to PIR|T52301|T5 GYMNOS/PICKLE protein [importe... 35 0.039
TC84117 weakly similar to GP|439289|emb|CAA81388.1| verprolin {S... 35 0.051
TC76771 homologue to SP|Q42961|PGKH_TOBAC Phosphoglycerate kinas... 34 0.066
TC80666 similar to PIR|E85112|E85112 probable methyltransferase ... 34 0.066
BG453645 34 0.066
TC78762 similar to GP|9294670|dbj|BAB03019.1 gene_id:F14O13.20~u... 34 0.087
BE997694 similar to GP|13646986|db DNA-binding protein DF1 {Pisu... 34 0.087
TC88622 weakly similar to GP|3336892|emb|CAA12389.1 Hsp20.0 prot... 32 0.25
TC81171 similar to GP|9795609|gb|AAF98427.1| Unknown protein {Ar... 32 0.25
TC87476 homologue to PIR|T11622|T11622 extensin class 1 precurso... 32 0.33
TC93132 32 0.33
TC86323 weakly similar to GP|15450363|gb|AAK96475.1 At1g16840/F1... 32 0.43
BQ751144 weakly similar to PIR|PQ0479|PQ04 pistil extensin-like ... 32 0.43
>BE319145
Length = 455
Score = 302 bits (774), Expect = 1e-82
Identities = 145/146 (99%), Positives = 145/146 (99%)
Frame = +1
Query: 1 MDFTKNITDLPPNKRLRFIYQQQQQQEEQDLSHCSLLPTKKRKESRNSSLFHTPPSSPTP 60
MDFTKNITDLPPNKRLRFIYQQQQQQEEQDLSHCSLLPTKKRKESRNSSLFHTPPSSPTP
Sbjct: 16 MDFTKNITDLPPNKRLRFIYQQQQQQEEQDLSHCSLLPTKKRKESRNSSLFHTPPSSPTP 195
Query: 61 PPSTYSLPTKKRITALQPHLHHHNNIPNDAVPLIDLNVEYSPSLPSATPIEKQSQKQDIE 120
PPSTYSLPTKKRITALQPHLHHHNNIPNDAVPLIDLNVEYSPSLPSATPIEKQSQKQDIE
Sbjct: 196 PPSTYSLPTKKRITALQPHLHHHNNIPNDAVPLIDLNVEYSPSLPSATPIEKQSQKQDIE 375
Query: 121 FEVDDDEILCCVCHSTDANAEDPIVF 146
FEVDDDEILC VCHSTDANAEDPIVF
Sbjct: 376 FEVDDDEILCWVCHSTDANAEDPIVF 453
>TC93186 similar to PIR|A86193|A86193 hypothetical protein [imported] -
Arabidopsis thaliana, partial (11%)
Length = 639
Score = 93.6 bits (231), Expect = 9e-20
Identities = 48/122 (39%), Positives = 62/122 (50%), Gaps = 1/122 (0%)
Frame = +3
Query: 130 CCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGP 189
C VCH + + + CD C +MVHA CYG + + W C+ CR + P
Sbjct: 297 CNVCHMDEEYENNLFLQCDKCRMMVHARCYGEH--EPVNGVLWLCNLCR------SGAPP 452
Query: 190 IRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGID-CSKIPKKRWLEKCY 248
C LCP GAMK TTDG+W HL CA+ +PE D + E ID +I K RW C
Sbjct: 453 PPCCLCPLIGGAMKPTTDGRWAHLACAMWIPETCLADVKRMEPIDGLRRISKDRWKLLCS 632
Query: 249 VC 250
+C
Sbjct: 633 IC 638
>TC89535 weakly similar to PIR|A96808|A96808 hypothetical protein T32E8.13
[imported] - Arabidopsis thaliana, partial (9%)
Length = 1729
Score = 70.5 bits (171), Expect = 8e-13
Identities = 47/148 (31%), Positives = 68/148 (45%), Gaps = 10/148 (6%)
Frame = +3
Query: 124 DDDEILCCVCHSTDANAE-DPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQC-RFKN 181
D D+ C C D++ + + +V C C + VH CYG V+ D W C C + K
Sbjct: 1113 DGDQPYCHYCGRGDSDTDSNRVVVCASCKVAVHRKCYG---VQDDVDDSWLCSWCSKQKG 1283
Query: 182 DIDTDTGPIRCSLCPTKEGAMK---QTTDG----KWVHLVCALLVPEVFFVDPEGREGI- 233
D+D P C LC K GA+K DG +VHL C L +PEV+ D + E +
Sbjct: 1284 DVDDSVNP--CVLCSKKGGALKPVYSAVDGVGSSPFVHLYCCLWMPEVYIEDLKKMEPVM 1457
Query: 234 DCSKIPKKRWLEKCYVCGCFDGCALVCS 261
+ I + R C +C G + C+
Sbjct: 1458 NVGGIKENRRKLMCNICKLRCGACVQCT 1541
>TC91564 weakly similar to GP|6573754|gb|AAF17674.1| F28K19.1 {Arabidopsis
thaliana}, partial (22%)
Length = 875
Score = 63.5 bits (153), Expect = 1e-10
Identities = 34/108 (31%), Positives = 56/108 (51%), Gaps = 12/108 (11%)
Frame = +1
Query: 130 CCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGP 189
C +C + N +PI+ C GC + VH+ CY + VK+ G W+C+ C ++ + +GP
Sbjct: 358 CDICRRFE-NVLNPILVCSGCKVAVHSVCYRS--VKETT-GPWYCELC--EDLLSRSSGP 519
Query: 190 ------------IRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFV 225
C+LC GA ++++DG+WVH CA + FF+
Sbjct: 520 SAINSWEKPYFVAECALCGGTTGAFRKSSDGQWVHAFCAEVTLSFFFL 663
>BG645509 similar to PIR|T47966|T47 hypothetical protein F15G16.130 -
Arabidopsis thaliana, partial (15%)
Length = 631
Score = 40.4 bits (93), Expect = 0.001
Identities = 22/64 (34%), Positives = 32/64 (49%), Gaps = 6/64 (9%)
Frame = +3
Query: 130 CCVC----HSTDANAEDPIVFCDGCNLMVHASC--YGNPLVKQIPDGDWFCDQCRFKNDI 183
C +C H +DA V CDGCN+ VHA C + K + + D++C C+ K+D
Sbjct: 441 CGICKKIWHHSDAG---DWVCCDGCNVWVHAECDKISSKRFKDLENIDYYCPDCKGKSDC 611
Query: 184 DTDT 187
T
Sbjct: 612 KLST 623
>AW560016 similar to GP|8778713|gb T25N20.3 {Arabidopsis thaliana}, partial
(13%)
Length = 657
Score = 38.5 bits (88), Expect = 0.004
Identities = 24/89 (26%), Positives = 33/89 (36%), Gaps = 24/89 (26%)
Frame = +1
Query: 134 HSTDANAEDP-------------IVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFK 180
HS D + DP ++ CDGC H SC ++ +P G+W C C K
Sbjct: 295 HSVDVDGNDPNDDTCGICGDGGDLICCDGCPSTFHQSCLD---IQMLPPGEWRCPNCTCK 465
Query: 181 ----------NDIDTDTGPIR-CSLCPTK 198
+ D +R C LC K
Sbjct: 466 FCGLASATTDKEDDATVNALRTCDLCEKK 552
>BG644635 weakly similar to GP|8843772|db contains similarity to zinc finger
protein~gene_id:MYN8.4 {Arabidopsis thaliana}, partial
(15%)
Length = 784
Score = 37.4 bits (85), Expect = 0.008
Identities = 19/58 (32%), Positives = 28/58 (47%), Gaps = 3/58 (5%)
Frame = +2
Query: 130 CCVCHSTDANAEDPI-VFCDGCNLMVHASC--YGNPLVKQIPDGDWFCDQCRFKNDID 184
C +C +++ V CDGC + VHA C + K + D+FC CR K D +
Sbjct: 248 CGICKKVSNHSDSGSWVRCDGCKVWVHAECDKISSNHFKDLETTDYFCPTCRGKFDFE 421
>BF645457 similar to PIR|T52301|T5 GYMNOS/PICKLE protein [imported] -
Arabidopsis thaliana, partial (9%)
Length = 560
Score = 35.0 bits (79), Expect = 0.039
Identities = 31/115 (26%), Positives = 48/115 (40%)
Frame = +2
Query: 86 IPNDAVPLIDLNVEYSPSLPSATPIEKQSQKQDIEFEVDDDEILCCVCHSTDANAEDPIV 145
+ +D P+ +L+ P Q + + I+ D E LC C + ++
Sbjct: 125 VRSDRKPVYNLDESDDEDFLLKKPGASQEKFERID-RSDAKEDLCQACGESG-----DLL 286
Query: 146 FCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEG 200
C CN H+SC PL PD +W C +C ID D + C + PT +G
Sbjct: 287 SCATCNYAYHSSCLLPPLKGPAPD-NWRCPEC-VTPLIDIDK-LLDCEMRPTVQG 442
>TC84117 weakly similar to GP|439289|emb|CAA81388.1| verprolin
{Saccharomyces cerevisiae}, partial (3%)
Length = 674
Score = 34.7 bits (78), Expect = 0.051
Identities = 28/88 (31%), Positives = 39/88 (43%), Gaps = 9/88 (10%)
Frame = +2
Query: 32 SHCSLLPTKKRKESRNSSLF------HTPPSSPTPPP---STYSLPTKKRITALQPHLHH 82
S S +P K SR+S H PP PTPPP S Y+LP+ + QP ++
Sbjct: 353 SFSSSVPMPDSKYSRHSISSPSGPSRHAPPLPPTPPPYASSPYNLPSSTNTSVSQPAPYN 532
Query: 83 HNNIPNDAVPLIDLNVEYSPSLPSATPI 110
I N L + +S + SA P+
Sbjct: 533 QAGIGN--TELSXAXIAHSGARLSAYPL 610
>TC76771 homologue to SP|Q42961|PGKH_TOBAC Phosphoglycerate kinase
chloroplast precursor (EC 2.7.2.3). [Common tobacco]
{Nicotiana tabacum}, partial (87%)
Length = 1708
Score = 34.3 bits (77), Expect = 0.066
Identities = 17/47 (36%), Positives = 25/47 (53%), Gaps = 5/47 (10%)
Frame = -1
Query: 78 PHLHHHNNIPNDAVPLIDL-----NVEYSPSLPSATPIEKQSQKQDI 119
P + HH+N+P+ + PL L + SPS P A P E Q Q+ +
Sbjct: 316 PSIQHHSNLPHSSSPLTPLLLLLSRLLSSPSTPPAPPTEGQLQRTQV 176
>TC80666 similar to PIR|E85112|E85112 probable methyltransferase [imported]
- Arabidopsis thaliana, partial (39%)
Length = 1616
Score = 34.3 bits (77), Expect = 0.066
Identities = 20/56 (35%), Positives = 29/56 (51%), Gaps = 4/56 (7%)
Frame = +3
Query: 48 SSLFHTP--PSSPTPPPSTYSLPTKKRITAL--QPHLHHHNNIPNDAVPLIDLNVE 99
SS H P P+ P PPP T P+K + L P +++NN+ N + L+ N E
Sbjct: 348 SSFNHRPFAPTPPLPPPLTNFNPSKSSLQQLPQNPFNNNNNNLQNPKISLVTKNPE 515
>BG453645
Length = 622
Score = 34.3 bits (77), Expect = 0.066
Identities = 25/75 (33%), Positives = 35/75 (46%), Gaps = 10/75 (13%)
Frame = -2
Query: 32 SHCSLLPTKKRKESRNSSLFHTPPSSPTPPPST-YSLPTKKRITALQP---------HLH 81
+ C + PT K + S L H PP + P S +S PTK + P H H
Sbjct: 366 TEC*ISPTSISKTTP*SLLSHPPPHNLAP*KSNHFS*PTKPK**YATPTNGLLLPYLHSH 187
Query: 82 HHNNIPNDAVPLIDL 96
H+NIPN+ + L +L
Sbjct: 186 LHSNIPNEQINLEEL 142
>TC78762 similar to GP|9294670|dbj|BAB03019.1 gene_id:F14O13.20~unknown
protein {Arabidopsis thaliana}, partial (87%)
Length = 976
Score = 33.9 bits (76), Expect = 0.087
Identities = 16/60 (26%), Positives = 27/60 (44%)
Frame = +1
Query: 118 DIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQC 177
D++ VD +E C C+ +V CD + + +G +K+ P G W+C C
Sbjct: 592 DLDLPVDPNEPTYCFCNQVSYGE---MVACDNPDCKIEWFHFGCVGLKEQPKGKWYCSSC 762
>BE997694 similar to GP|13646986|db DNA-binding protein DF1 {Pisum sativum},
partial (19%)
Length = 609
Score = 33.9 bits (76), Expect = 0.087
Identities = 28/132 (21%), Positives = 63/132 (47%), Gaps = 6/132 (4%)
Frame = +3
Query: 16 LRFIYQQQQQQEEQDL-----SHCSLLPTKKRKESRNSSLFHTPPSSPTPPPSTYSLPTK 70
+ F+ + +QQE+Q+L ++ +++P ++++ + + TP +PTP P+ LP
Sbjct: 219 MAFLQKIAEQQEQQNLVPPVLNNSTIVP-QQQQAPQETIPTPTPKPTPTPTPTPVPLPAA 395
Query: 71 KRITALQ-PHLHHHNNIPNDAVPLIDLNVEYSPSLPSATPIEKQSQKQDIEFEVDDDEIL 129
+ P + + N V + + + +P A+P+ Q Q+Q + +V +++
Sbjct: 396 AAPLPIPIPAIPTPQQVQNPTVTV----QQQTSVIPQASPL-PQHQQQQQQVQVQQQQVM 560
Query: 130 CCVCHSTDANAE 141
+D N E
Sbjct: 561 NMEVAKSDNNGE 596
>TC88622 weakly similar to GP|3336892|emb|CAA12389.1 Hsp20.0 protein
{Lycopersicon peruvianum}, partial (64%)
Length = 1032
Score = 32.3 bits (72), Expect = 0.25
Identities = 25/103 (24%), Positives = 42/103 (40%), Gaps = 14/103 (13%)
Frame = +2
Query: 48 SSLFHTPPSSPTPPPSTYSLPTKKR--ITALQPHLHHHNNIP---NDAVPLIDLNVE--- 99
+S+F S P T+ P+ + Q + HH P N+ P+I+ +E
Sbjct: 65 NSIFGRRRSEPKDHHQTWHHPSYQNHGYGISQTNTPHHITPPPFHNEPSPIINTQIEWKE 244
Query: 100 ------YSPSLPSATPIEKQSQKQDIEFEVDDDEILCCVCHST 136
Y LP ++ D+ EVD+D +LC +C +
Sbjct: 245 THEAHIYKAHLPGL-------KRSDVRVEVDEDRVLCIICEKS 352
>TC81171 similar to GP|9795609|gb|AAF98427.1| Unknown protein {Arabidopsis
thaliana}, partial (40%)
Length = 932
Score = 32.3 bits (72), Expect = 0.25
Identities = 31/114 (27%), Positives = 43/114 (37%)
Frame = +3
Query: 26 QEEQDLSHCSLLPTKKRKESRNSSLFHTPPSSPTPPPSTYSLPTKKRITALQPHLHHHNN 85
QEEQ S SL K + N S + P P SLP+ K T H H+HN+
Sbjct: 36 QEEQKSSLTSLPSPKTQSNGHNHS------HNQIPSPRPISLPSPKTQTQSNGHNHNHNH 197
Query: 86 IPNDAVPLIDLNVEYSPSLPSATPIEKQSQKQDIEFEVDDDEILCCVCHSTDAN 139
+ I + +P + + S KQ ++ E ST AN
Sbjct: 198 NQIPSPRPITRSEPGNPYPTTFVQADTTSFKQVVQMLTGSSETAKQASTSTKAN 359
>TC87476 homologue to PIR|T11622|T11622 extensin class 1 precursor - cowpea,
partial (48%)
Length = 939
Score = 32.0 bits (71), Expect = 0.33
Identities = 20/62 (32%), Positives = 28/62 (44%)
Frame = +3
Query: 48 SSLFHTPPSSPTPPPSTYSLPTKKRITALQPHLHHHNNIPNDAVPLIDLNVEYSPSLPSA 107
S+ H PPS PPP Y P + P+++ P+ + P V SP PSA
Sbjct: 351 STSHHRPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPPSPSPP--PPYVYKSPPPPSA 524
Query: 108 TP 109
+P
Sbjct: 525 SP 530
>TC93132
Length = 638
Score = 32.0 bits (71), Expect = 0.33
Identities = 24/91 (26%), Positives = 37/91 (40%), Gaps = 10/91 (10%)
Frame = +3
Query: 24 QQQEEQDLSHCSLLPTKKRKESRNSSLFHTPPSSPT----------PPPSTYSLPTKKRI 73
Q EE+++S S L +K + +HTPP+SP+ PP L R
Sbjct: 156 QHTEEKEVS--SDLERRKEVQEEEHYYYHTPPTSPSKNSFDLVCPPPPKKRQRLAVTTRR 329
Query: 74 TALQPHLHHHNNIPNDAVPLIDLNVEYSPSL 104
T+ Q +P+D + L + S L
Sbjct: 330 TSTQSQERKFFQVPDDLTSIFLLRTKPSHQL 422
>TC86323 weakly similar to GP|15450363|gb|AAK96475.1 At1g16840/F17F16.27
{Arabidopsis thaliana}, partial (28%)
Length = 1069
Score = 31.6 bits (70), Expect = 0.43
Identities = 22/63 (34%), Positives = 29/63 (45%), Gaps = 8/63 (12%)
Frame = +1
Query: 31 LSHCSLLPTKKRKESRNS---SLFHTPPSSP---TPPPST--YSLPTKKRITALQPHLHH 82
LSH PT K + +R S + PSSP T PPS + P + + H HH
Sbjct: 82 LSHTIFSPTTKTQPNRQPWPVSSRNPKPSSPIHQTSPPSVTVHHAPQENSTKSTHHHHHH 261
Query: 83 HNN 85
HN+
Sbjct: 262 HNH 270
>BQ751144 weakly similar to PIR|PQ0479|PQ04 pistil extensin-like protein
(clone pMG14) - common tobacco (fragment), partial (11%)
Length = 632
Score = 31.6 bits (70), Expect = 0.43
Identities = 25/69 (36%), Positives = 30/69 (43%), Gaps = 4/69 (5%)
Frame = +1
Query: 45 SRNSSLFHTPPSSPT----PPPSTYSLPTKKRITALQPHLHHHNNIPNDAVPLIDLNVEY 100
+R SSLF PP SP+ PPPST S P +PL+ L E
Sbjct: 295 TRPSSLFSPPPPSPSRSRPPPPSTPS--------------------PLLPLPLLPLATEP 414
Query: 101 SPSLPSATP 109
SP PS+ P
Sbjct: 415 SPFPPSSPP 441
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.137 0.445
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,298,139
Number of Sequences: 36976
Number of extensions: 288655
Number of successful extensions: 3449
Number of sequences better than 10.0: 141
Number of HSP's better than 10.0 without gapping: 2929
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3297
length of query: 322
length of database: 9,014,727
effective HSP length: 96
effective length of query: 226
effective length of database: 5,465,031
effective search space: 1235097006
effective search space used: 1235097006
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC147714.16


