

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
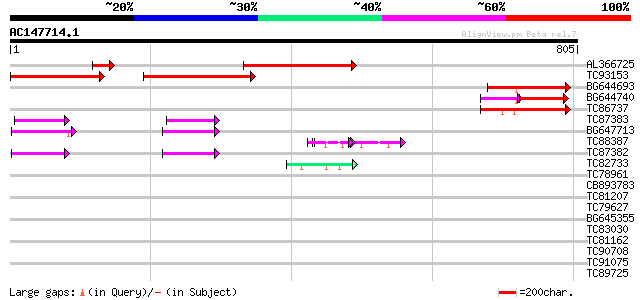
Query= AC147714.1 - phase: 0 /pseudo
(805 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AL366725 291 9e-79
TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Ci... 246 2e-65
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 122 5e-28
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 89 3e-25
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 97 2e-20
TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA PO... 56 6e-08
BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent vir... 55 8e-08
TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 53 4e-07
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 53 5e-07
TC82733 similar to GP|10177404|dbj|BAB10535. gene_id:K24M7.12~pi... 47 2e-05
TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Ar... 41 0.002
CB893783 weakly similar to GP|22830935|dbj hypothetical protein~... 40 0.005
TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-... 39 0.010
TC79627 37 0.030
BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.... 35 0.11
TC83030 weakly similar to GP|18855061|gb|AAL79753.1 putative RNA... 33 0.57
TC81162 weakly similar to PIR|T47673|T47673 hypothetical protein... 33 0.57
TC90708 weakly similar to PIR|S26703|S26703 dnaJ protein homolog... 32 0.74
TC91075 similar to PIR|T48025|T48025 hypothetical protein T12C14... 32 0.97
TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.... 32 1.3
>AL366725
Length = 485
Score = 291 bits (744), Expect = 9e-79
Identities = 139/161 (86%), Positives = 144/161 (89%), Gaps = 1/161 (0%)
Frame = +2
Query: 333 KFYPHYTAENAEFSRCIKFENGLRLDIKRAIGYQQLRVFQDLVNSCRIYEEDTKAHYKVV 392
KFYPHY AE AEFS+CIKFENGLR DIKRAIGYQQLRVF DLVN+CRIYEEDTKAH KVV
Sbjct: 2 KFYPHYAAETAEFSKCIKFENGLRPDIKRAIGYQQLRVFPDLVNTCRIYEEDTKAHDKVV 181
Query: 393 NERKGKGQQSRPKPYSAPADKGKQKMVDVRRPKKKDA-AEIVCFNCGEKGHKSNACPEEI 451
NERK KGQ SRPKPYSAPADKGKQ+MVD RRPKKKDA AEIVCFN GEKGHKSN CP+EI
Sbjct: 182 NERKTKGQ*SRPKPYSAPADKGKQRMVDDRRPKKKDAPAEIVCFNYGEKGHKSNVCPKEI 361
Query: 452 KKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPKR 492
KKCVRC KKGH+VADC R DIVCFNCN EGHI SQC QPKR
Sbjct: 362 KKCVRCDKKGHIVADCKRNDIVCFNCNEEGHIGSQCKQPKR 484
Score = 56.6 bits (135), Expect = 4e-08
Identities = 23/31 (74%), Positives = 26/31 (83%)
Frame = +2
Query: 118 KFYPHYTGENAEFSRCIKIQNGLRPDIKRRL 148
KFYPHY E AEFS+CIK +NGLRPDIKR +
Sbjct: 2 KFYPHYAAETAEFSKCIKFENGLRPDIKRAI 94
>TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Cicer
arietinum}, partial (8%)
Length = 516
Score = 246 bits (628), Expect = 2e-65
Identities = 120/160 (75%), Positives = 138/160 (86%)
Frame = +2
Query: 190 LQAVAQAVGQQPNVNAGANAEARMLETFMKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQ 249
L+AVAQAV Q P V+ G++ RMLETF++ +PPTFKGRY PDGA WLKEIERIFRVMQ
Sbjct: 38 LEAVAQAVQQLPKVDTGSDG-TRMLETFLRNHPPTFKGRYAPDGA*KWLKEIERIFRVMQ 214
Query: 250 YTEEQKVRFGTHQLAEEADDWWVALLPTLGQEGAVMTWAVFRREFLRRYFLEDVRGKKEI 309
E QKV+FGTH LAEEADDWW++LLP L Q+ AV+TWA+FR+EFL RYF EDVRGKKEI
Sbjct: 215 CFETQKVQFGTHMLAEEADDWWISLLPVLEQDDAVVTWAMFRKEFLGRYFPEDVRGKKEI 394
Query: 310 EFLELKQGNMSVTEYAAKFVELSKFYPHYTAENAEFSRCI 349
EFLELKQG+MSVTEYAAKFVEL+ FYPHY+AE AEFS+CI
Sbjct: 395 EFLELKQGDMSVTEYAAKFVELATFYPHYSAETAEFSKCI 514
Score = 227 bits (579), Expect = 1e-59
Identities = 105/134 (78%), Positives = 119/134 (88%)
Frame = +2
Query: 1 TFMKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQCTEEQKVRFGTHQLAEAADDWWVALL 60
TF++ +PPTFKGRY PDGA WLKEIERIFRVMQC E QKV+FGTH LAE ADDWW++LL
Sbjct: 113 TFLRNHPPTFKGRYAPDGA*KWLKEIERIFRVMQCFETQKVQFGTHMLAEEADDWWISLL 292
Query: 61 PTLGQEGAVVTWAVFRREFLRRYFPEDVRGKKEIEFLELKQGNMSVTEYAAQFVELSKFY 120
P L Q+ AVVTWA+FR+EFL RYFPEDVRGKKEIEFLELKQG+MSVTEYAA+FVEL+ FY
Sbjct: 293 PVLEQDDAVVTWAMFRKEFLGRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELATFY 472
Query: 121 PHYTGENAEFSRCI 134
PHY+ E AEFS+CI
Sbjct: 473 PHYSAETAEFSKCI 514
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 122 bits (306), Expect = 5e-28
Identities = 64/121 (52%), Positives = 84/121 (68%), Gaps = 3/121 (2%)
Frame = +2
Query: 679 DEIPDVPPEREVEFSIDLVPGTKPVSMAPYRISASELKK---QLEDLLEKKFVRPSVSPW 735
D + VPPE +++F IDL+P P+ + YRI+ +LK QL+DLLEK F++PS+ P
Sbjct: 2 DHLL*VPPEWKIDFGIDLLPNMNPI*IPSYRINPLKLKVLKLQLKDLLEKGFIQPSIYP* 181
Query: 736 GASVLLVKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGARVFSKIDLRSGYH 795
G VL +KKKDG +R+ IDY QLN V IK +YPLP ID+L D L G++ F KIDLR G H
Sbjct: 182 GVVVLFLKKKDGFLRMSIDYPQLNNVNIKIKYPLPLIDELFDNLQGSKWFFKIDLRLG*H 361
Query: 796 Q 796
Q
Sbjct: 362 Q 364
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 89.0 bits (219), Expect(2) = 3e-25
Identities = 42/69 (60%), Positives = 53/69 (75%)
Frame = -1
Query: 725 KKFVRPSVSPWGASVLLVKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGARV 784
K+F +PS+SP GA++L V+KKDG R+CIDYRQ NKVT KN+YPLPRID+L D++
Sbjct: 274 KRFQQPSISP*GAALLFVRKKDGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFDKIQEDCY 95
Query: 785 FSKIDLRSG 793
F IDLR G
Sbjct: 94 F*NIDLRLG 68
Score = 45.4 bits (106), Expect(2) = 3e-25
Identities = 28/62 (45%), Positives = 37/62 (59%), Gaps = 3/62 (4%)
Frame = -2
Query: 669 VVCDFPEVFPDEIPDVPPEREVEFSIDLVPGTKPVSMAPYRISASELKKQ---LEDLLEK 725
V+ F VFPD P +P ERE+ F IDL+ T+ +S P + +ELKK L+D LEK
Sbjct: 450 VLKGFS*VFPDNFPVIPLEREIFFCIDLLLDTQLISNPP*LMDRTELKKLKI*LKDSLEK 271
Query: 726 KF 727
F
Sbjct: 270 GF 265
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 97.4 bits (241), Expect = 2e-20
Identities = 57/138 (41%), Positives = 87/138 (62%), Gaps = 10/138 (7%)
Frame = +1
Query: 669 VVCDFPEVF-PDEIPDVPPEREV-EFSIDLVP----GTKPVSMAP-YRISASEL---KKQ 718
V+ +FP++F P++ VP R + + +I L+P P+ P Y +S EL KK
Sbjct: 343 VLEEFPDLFNPEKAYQVPASRGLLDHAIPLIPDKDGNDPPLPWGPLYGMSRQELLVLKKT 522
Query: 719 LEDLLEKKFVRPSVSPWGASVLLVKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQ 778
LEDLL+K F++ S S GA VL V+K G +R C+DYR LN +T K+RYPLP I + + +
Sbjct: 523 LEDLLDKGFIKASGSAAGAPVLFVRKPGGGIRFCVDYRALNAITKKDRYPLPLISETLRR 702
Query: 779 LVGARVFSKIDLRSGYHQ 796
+ GAR F+K+D+ + +H+
Sbjct: 703 VAGARWFTKLDVVAAFHK 756
>TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA POLYMERASE II
{Encephalitozoon cuniculi}, partial (0%)
Length = 1247
Score = 55.8 bits (133), Expect = 6e-08
Identities = 27/80 (33%), Positives = 42/80 (51%), Gaps = 2/80 (2%)
Frame = -2
Query: 8 PTFKGRYDPDGAQTWLKEIERIFRVMQCTEEQKVRFGTHQLAEAADDWWVALLPTLGQEG 67
P F+G PD WL+ +ER+F+ + EEQKV+ +L + A WW + +EG
Sbjct: 724 PDFEGNLQPDDLLDWLQIMERLFKYKEVLEEQKVKIVAAKLKKLASIWWENVKRRRKREG 545
Query: 68 --AVVTWAVFRREFLRRYFP 85
+ TW R++ R+Y P
Sbjct: 544 KSKIKTWEKMRQKLTRKYLP 485
Score = 52.8 bits (125), Expect = 5e-07
Identities = 26/78 (33%), Positives = 41/78 (52%), Gaps = 2/78 (2%)
Frame = -2
Query: 223 PTFKGRYDPDGAQTWLKEIERIFRVMQYTEEQKVRFGTHQLAEEADDWWVALLPTLGQEG 282
P F+G PD WL+ +ER+F+ + EEQKV+ +L + A WW + +EG
Sbjct: 724 PDFEGNLQPDDLLDWLQIMERLFKYKEVLEEQKVKIVAAKLKKLASIWWENVKRRRKREG 545
Query: 283 --AVMTWAVFRREFLRRY 298
+ TW R++ R+Y
Sbjct: 544 KSKIKTWEKMRQKLTRKY 491
>BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent virus}, partial
(1%)
Length = 726
Score = 55.5 bits (132), Expect = 8e-08
Identities = 29/83 (34%), Positives = 43/83 (50%), Gaps = 2/83 (2%)
Frame = +2
Query: 218 MKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQYTEEQKVRFGTHQLAEEADDWWVALLPT 277
+K + P F G+ PD WL+ IER+F+ + EEQKV+ +L + A WW L
Sbjct: 332 IKVDIPDF*GKLQPDEFVDWLQTIERVFKYKEVAEEQKVKIVAAKLKKHASIWWKNLKRK 511
Query: 278 LGQEG--AVMTWAVFRREFLRRY 298
EG + TW R++ R+Y
Sbjct: 512 RNCEGKSKIKTWDKMRQKLTRKY 580
Score = 54.7 bits (130), Expect = 1e-07
Identities = 32/100 (32%), Positives = 50/100 (50%), Gaps = 7/100 (7%)
Frame = +2
Query: 3 MKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQCTEEQKVRFGTHQLAEAADDWWVALLPT 62
+K + P F G+ PD WL+ IER+F+ + EEQKV+ +L + A WW L
Sbjct: 332 IKVDIPDF*GKLQPDEFVDWLQTIERVFKYKEVAEEQKVKIVAAKLKKHASIWWKNLKRK 511
Query: 63 LGQEG--AVVTWAVFRREFLRR-----YFPEDVRGKKEIE 95
EG + TW R++ R+ Y+ ++ KK I+
Sbjct: 512 RNCEGKSKIKTWDKMRQKLTRKYLHPHYYQDNFTQKKNIQ 631
>TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (96%)
Length = 1286
Score = 53.1 bits (126), Expect = 4e-07
Identities = 25/65 (38%), Positives = 35/65 (53%), Gaps = 6/65 (9%)
Frame = +1
Query: 433 VCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTD------IVCFNCNGEGHISSQ 486
+C+NC E GH +++CP E C CGK GH +C +C NC +GHI+ +
Sbjct: 535 LCWNCKEPGHMASSCPNE-GICHTCGKAGHRARECTVPQKPPGDLRLCNNCYKQGHIAVE 711
Query: 487 CTQPK 491
CT K
Sbjct: 712 CTNEK 726
Score = 51.6 bits (122), Expect = 1e-06
Identities = 37/141 (26%), Positives = 58/141 (40%), Gaps = 11/141 (7%)
Frame = +1
Query: 433 VCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPKR 492
+C NC GH CP + C C GH+ ++C+ T +C+NC GH++S C
Sbjct: 421 LCKNCKRPGHYVRECPN-VAVCHNCSLPGHIASECS-TKSLCWNCKEPGHMASSCPNEGI 594
Query: 493 APTTG----RVFALTGTQTENEDRLIRGTCYINNTPLVAIIDTGATHC-------FIAFD 541
T G R T Q D + CY +A+ T C +A D
Sbjct: 595 CHTCGKAGHRARECTVPQKPPGDLRLCNNCYKQGH--IAVECTNEKACNNCRKTGHLARD 768
Query: 542 CVSALGLDLSDMNGEMVVETP 562
C + +L +++G + + P
Sbjct: 769 CPNDPICNLCNISGHVARQCP 831
Score = 50.8 bits (120), Expect = 2e-06
Identities = 25/64 (39%), Positives = 34/64 (53%)
Frame = +1
Query: 424 PKKKDAAEIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHI 483
P+K +C NC ++GH + C E K C C K GH+ DC D +C CN GH+
Sbjct: 643 PQKPPGDLRLCNNCYKQGHIAVECTNE-KACNNCRKTGHLARDC-PNDPICNLCNISGHV 816
Query: 484 SSQC 487
+ QC
Sbjct: 817 ARQC 828
Score = 50.8 bits (120), Expect = 2e-06
Identities = 20/63 (31%), Positives = 33/63 (51%), Gaps = 6/63 (9%)
Frame = +1
Query: 431 EIVCFNCGEKGHKSNAC------PEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHIS 484
E +C CG+ GH++ C P +++ C C K+GH+ +C + C NC GH++
Sbjct: 586 EGICHTCGKAGHRARECTVPQKPPGDLRLCNNCYKQGHIAVECT-NEKACNNCRKTGHLA 762
Query: 485 SQC 487
C
Sbjct: 763 RDC 771
Score = 38.9 bits (89), Expect = 0.008
Identities = 22/79 (27%), Positives = 30/79 (37%), Gaps = 22/79 (27%)
Frame = +1
Query: 431 EIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRT-------------------- 470
E C NC + GH + CP + C C GHV C ++
Sbjct: 721 EKACNNCRKTGHLARDCPND-PICNLCNISGHVARQCPKSNVIGDRGGGGSLRGGYRDGG 897
Query: 471 --DIVCFNCNGEGHISSQC 487
D+VC +C GH+S C
Sbjct: 898 FRDVVCRSCQQFGHMSRDC 954
Score = 32.3 bits (72), Expect = 0.74
Identities = 19/78 (24%), Positives = 28/78 (35%), Gaps = 23/78 (29%)
Frame = +1
Query: 433 VCFNCGEKGHKSNACPEEIK----------------------KCVRCGKKGHVVADCNRT 470
+C C GH + CP+ C C + GH+ DC
Sbjct: 784 ICNLCNISGHVARQCPKSNVIGDRGGGGSLRGGYRDGGFRDVVCRSCQQFGHMSRDCMGG 963
Query: 471 DI-VCFNCNGEGHISSQC 487
+ +C NC G GH + +C
Sbjct: 964 PLMICQNCGGRGHQAYEC 1017
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 52.8 bits (125), Expect = 5e-07
Identities = 29/85 (34%), Positives = 43/85 (50%), Gaps = 2/85 (2%)
Frame = +2
Query: 3 MKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQCTEEQKVRFGTHQLAEAADDWWVALLPT 62
+K + P F+G D WL+ IER+F + EEQKV+ +L + A WW L
Sbjct: 614 IKVDIPDFEGNLQLDDFLDWLQTIERVFEYKEVPEEQKVKIVAAKLKKHALIWWENLKRR 793
Query: 63 LGQEG--AVVTWAVFRREFLRRYFP 85
+EG + TW R++ R+Y P
Sbjct: 794 RKREGKSKIKTWDKMRQKLTRKYLP 868
Score = 51.2 bits (121), Expect = 2e-06
Identities = 28/83 (33%), Positives = 42/83 (49%), Gaps = 2/83 (2%)
Frame = +2
Query: 218 MKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQYTEEQKVRFGTHQLAEEADDWWVALLPT 277
+K + P F+G D WL+ IER+F + EEQKV+ +L + A WW L
Sbjct: 614 IKVDIPDFEGNLQLDDFLDWLQTIERVFEYKEVPEEQKVKIVAAKLKKHALIWWENLKRR 793
Query: 278 LGQEG--AVMTWAVFRREFLRRY 298
+EG + TW R++ R+Y
Sbjct: 794 RKREGKSKIKTWDKMRQKLTRKY 862
>TC82733 similar to GP|10177404|dbj|BAB10535.
gene_id:K24M7.12~pir||S42136~similar to unknown protein
{Arabidopsis thaliana}, partial (57%)
Length = 710
Score = 47.4 bits (111), Expect = 2e-05
Identities = 32/118 (27%), Positives = 46/118 (38%), Gaps = 17/118 (14%)
Frame = +3
Query: 393 NERKGKGQQSRPKPYSAPAD-----KGKQKMVDVRRPKKKDAAEIVCFNCGEKGHKSNAC 447
N + G++ P P D KG + K + +C C +GH++ C
Sbjct: 219 NSKPRTGKRPLRVPGMKPGDSCFICKGLDHIAKFCTQKAEWEKNKICLRCRRRGHRAQNC 398
Query: 448 P-----EEIKKCVRCGKKGHVVADC-------NRTDIVCFNCNGEGHISSQCTQPKRA 493
P E+ K CG GH +A+C CF C +GH+S C PK A
Sbjct: 399 PDGGSKEDFKY*YNCGDNGHSLANCPHPLQEGGTMFAQCFVCKEQGHLSKNC--PKNA 566
>TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Arabidopsis
thaliana}, partial (71%)
Length = 974
Score = 40.8 bits (94), Expect = 0.002
Identities = 16/39 (41%), Positives = 23/39 (58%), Gaps = 2/39 (5%)
Frame = +2
Query: 434 CFNCGEKGHKSNACP--EEIKKCVRCGKKGHVVADCNRT 470
CFNCG GH + C + KC RCG++GH+ +C +
Sbjct: 359 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNS 475
Score = 35.0 bits (79), Expect = 0.11
Identities = 16/43 (37%), Positives = 19/43 (43%), Gaps = 3/43 (6%)
Frame = +2
Query: 453 KCVRCGKKGHVVADCNRTD--IVCFNCNGEGHISSQC-TQPKR 492
+C CG GH DC D C+ C GHI C PK+
Sbjct: 356 RCFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNSPKK 484
>CB893783 weakly similar to GP|22830935|dbj hypothetical protein~similar to
gag-pol polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 853
Score = 39.7 bits (91), Expect = 0.005
Identities = 42/195 (21%), Positives = 83/195 (42%), Gaps = 23/195 (11%)
Frame = +2
Query: 528 IIDTGATHCFIAFDCVSALGLDLSDMNGEMVVETPAKGSVTTSLVCLKCPLSMFGRDFEM 587
IID+G+ ++ V L L D ++ KG+ C S+ G+ ++
Sbjct: 269 IIDSGSCENVVSNYMVEKLELPTKDHPHRYKLQWLKKGNEVRVSKCCLVSFSI-GQKYKD 445
Query: 588 DLVC--LPLSGMDVILGMNWLEYNHVLINCFSKSVHF-------------SSVEEESGAE 632
++ C + + ++LG W H L + + + F + +E +
Sbjct: 446 NVWCDVISMDACHMLLGRPWQYDRHALYDGHANTYTFVKYGVKIKLVPLPPNAFDEGKKD 625
Query: 633 F-----LSTKQLKQLERDGILMFSLMATLSIENQAVIDRLP--VVCDFPEVFPDEIPD-V 684
F L +K+ ++ I SL+ + ++ I + ++ DF +V P EIP +
Sbjct: 626 FKPIVSLVSKEPFKVTTKDIQDMSLILLVKSNEESTIQKEVEHLLVDFTDVVPSEIPSGL 805
Query: 685 PPEREVEFSIDLVPG 699
PP R+++ +ID +PG
Sbjct: 806 PPMRDIQHAIDFIPG 850
>TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-1 {Triticum
aestivum}, partial (39%)
Length = 630
Score = 38.5 bits (88), Expect = 0.010
Identities = 31/136 (22%), Positives = 40/136 (28%), Gaps = 54/136 (39%)
Frame = +3
Query: 409 APADKGKQKMVDVRRPK-----------------------KKDAAEIVCFNCGEKGHKSN 445
A D GK K VDV PK ++ C+ CG+ GH +
Sbjct: 207 ADGDNGKTKAVDVTGPKGEPLQVRQDNHGGGGGGRGFRGGERRNGGGGCYTCGDTGHIAR 386
Query: 446 ACP-----------------EEIKKCVRCGKKGHVVADCNRTD--------------IVC 474
C + + C CG H DC R C
Sbjct: 387 DCDRSDRNDRNDRSGGGGGGDRDRACYTCGSFEHFARDCMRGGGNNNNGGGGYGGGGTSC 566
Query: 475 FNCNGEGHISSQCTQP 490
+ C G GHI+ C P
Sbjct: 567 YRCGGVGHIARDCATP 614
>TC79627
Length = 1522
Score = 37.0 bits (84), Expect = 0.030
Identities = 23/82 (28%), Positives = 40/82 (48%), Gaps = 7/82 (8%)
Frame = +2
Query: 583 RDFEMDLVCLPLSGMDVILGMNWLEYNHVLINCFS-------KSVHFSSVEEESGAEFLS 635
R++ +C GMD++L W+ YN+ L++C +V FS++E+ A L
Sbjct: 929 RNYPFVFICRTARGMDLMLV--WIAYNYSLVSCIHAGFCFLLPTVVFSTLEDNPAANLL* 1102
Query: 636 TKQLKQLERDGILMFSLMATLS 657
K L + + MF+L+ S
Sbjct: 1103 HKDSFNLVPEVVRMFNLLVNTS 1168
>BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.5 [imported]
- Arabidopsis thaliana, partial (5%)
Length = 627
Score = 35.0 bits (79), Expect = 0.11
Identities = 23/96 (23%), Positives = 31/96 (31%), Gaps = 35/96 (36%)
Frame = -1
Query: 434 CFNCGEKGHKSNAC------PEEIK----------------KCVRCGKKGHVVADCNRTD 471
C NCG H S C PE+ KC +C + GH ++C
Sbjct: 627 CSNCGGSDHSSAQCLHLRNPPEQTAGGAYVNTVSGSGGASGKCYKCQQPGHWASNCPSMS 448
Query: 472 IV-------------CFNCNGEGHISSQCTQPKRAP 494
C+ CN GH ++ C AP
Sbjct: 447 AANRVSGGSGGASGNCYKCNQPGHWANNCPNMSAAP 340
>TC83030 weakly similar to GP|18855061|gb|AAL79753.1 putative RNA helicase
{Oryza sativa}, partial (7%)
Length = 624
Score = 32.7 bits (73), Expect = 0.57
Identities = 19/60 (31%), Positives = 26/60 (42%), Gaps = 4/60 (6%)
Frame = +1
Query: 395 RKGKGQQSRPKPYSAPADK----GKQKMVDVRRPKKKDAAEIVCFNCGEKGHKSNACPEE 450
R K S KP + D G+Q P + A CF CGE GH+++ CP +
Sbjct: 43 RSYKSGNSWSKPERSSRDDWLIGGRQSSRSSSSPNRSFAG--TCFTCGESGHRASDCPNK 216
>TC81162 weakly similar to PIR|T47673|T47673 hypothetical protein T26I12.220
- Arabidopsis thaliana, partial (9%)
Length = 756
Score = 32.7 bits (73), Expect = 0.57
Identities = 28/109 (25%), Positives = 46/109 (41%), Gaps = 5/109 (4%)
Frame = +2
Query: 345 FSRCIKFENGLRLDIKRAIGYQQLRVFQDLVNSCRI-YEEDTKAHYKVVNERKGKGQQSR 403
F R ++ + L+ K G + + V + I E+ A V E+ G G++S
Sbjct: 155 FGRPVRISCAVPLNKKPVAGEKSVAVEKPGAGEKSIAVEKSVIAEKSVAGEKSGAGEKSF 334
Query: 404 PKPYSAPADKGKQKMVD----VRRPKKKDAAEIVCFNCGEKGHKSNACP 448
P GK V+ V K+K+ +C+ C +KGH + CP
Sbjct: 335 A--VEQPGAGGKPSSVEQPSSVASGKRKNR---MCYGCRQKGHNLSECP 466
Score = 29.6 bits (65), Expect = 4.8
Identities = 12/33 (36%), Positives = 19/33 (57%)
Frame = +2
Query: 464 VADCNRTDIVCFNCNGEGHISSQCTQPKRAPTT 496
VA R + +C+ C +GH S+C P+ A +T
Sbjct: 392 VASGKRKNRMCYGCRQKGHNLSECPNPQIAAST 490
>TC90708 weakly similar to PIR|S26703|S26703 dnaJ protein homolog YDJ1 -
yeast (Saccharomyces cerevisiae), partial (43%)
Length = 1098
Score = 32.3 bits (72), Expect = 0.74
Identities = 24/78 (30%), Positives = 32/78 (40%), Gaps = 9/78 (11%)
Frame = +2
Query: 415 KQKMVDVRRPKKKDAA---EIVCFNCGEKGHKSNACPEEIKKCVRCGKKG------HVVA 465
K + DV R K A I+C C +G K A +K+C C G +
Sbjct: 644 KVSLEDVYRGKISKLALQRSIICSKCEGRGGKEGA----VKRCTGCDGHGMKTMMRQMGP 811
Query: 466 DCNRTDIVCFNCNGEGHI 483
R VC +CNGEG +
Sbjct: 812 MIQRFQTVCPDCNGEGEM 865
>TC91075 similar to PIR|T48025|T48025 hypothetical protein T12C14.30 -
Arabidopsis thaliana, partial (29%)
Length = 686
Score = 32.0 bits (71), Expect = 0.97
Identities = 12/30 (40%), Positives = 17/30 (56%)
Frame = +2
Query: 422 RRPKKKDAAEIVCFNCGEKGHKSNACPEEI 451
R KK+ E++C CGE GH + CP +
Sbjct: 353 RNWKKRKEEEMICKLCGESGHFTQGCPSTL 442
>TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.150 -
Arabidopsis thaliana, partial (17%)
Length = 378
Score = 31.6 bits (70), Expect = 1.3
Identities = 15/54 (27%), Positives = 18/54 (32%), Gaps = 20/54 (37%)
Frame = +1
Query: 434 CFNCGEKGHKSNACPE--------------------EIKKCVRCGKKGHVVADC 467
C+NCGE GH + C C CG+ GH DC
Sbjct: 16 CYNCGESGHMARECTSGGGGGGGRYGGGGGGGGGGGGGGSCYSCGESGHFARDC 177
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.138 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,340,397
Number of Sequences: 36976
Number of extensions: 310834
Number of successful extensions: 1540
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 1333
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1503
length of query: 805
length of database: 9,014,727
effective HSP length: 104
effective length of query: 701
effective length of database: 5,169,223
effective search space: 3623625323
effective search space used: 3623625323
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC147714.1


