

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147712.6 + phase: 0
(80 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
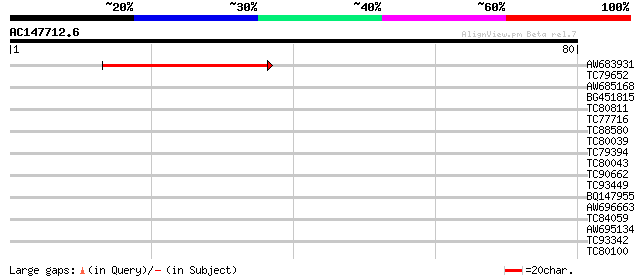
Score E
Sequences producing significant alignments: (bits) Value
AW683931 weakly similar to GP|21553880|gb| unknown {Arabidopsis ... 42 4e-05
TC79652 weakly similar to GP|19698951|gb|AAL91211.1 unknown prot... 33 0.020
AW685168 weakly similar to PIR|T01757|T0 hypothetical protein A_... 33 0.020
BG451815 similar to GP|21537091|gb unknown {Arabidopsis thaliana... 33 0.026
TC80811 similar to GP|21537091|gb|AAM61432.1 unknown {Arabidopsi... 32 0.058
TC77716 similar to GP|20465587|gb|AAM20276.1 unknown protein {Ar... 29 0.38
TC88580 similar to GP|9758308|dbj|BAB08782.1 gene_id:MUA2.4~unkn... 28 0.84
TC80039 homologue to GP|6437558|gb|AAF08585.1| putative ATPase (... 26 2.4
TC79394 weakly similar to PIR|T49019|T49019 probable RNA binding... 26 3.2
TC80043 similar to GP|9759561|dbj|BAB11163.1 MtN21 nodulin prote... 25 5.4
TC90662 similar to GP|22136654|gb|AAM91646.1 unknown protein {Ar... 25 5.4
TC93449 similar to GP|21328382|gb|AAM48550.1 Hypothetical protei... 25 7.1
BQ147955 weakly similar to GP|15290125|dbj putative kinase {Oryz... 24 9.3
AW696663 weakly similar to GP|17827193|dbj ORF1 {TT virus}, part... 24 9.3
TC84059 weakly similar to PIR|T02508|T02508 hypothetical protein... 24 9.3
AW695134 24 9.3
TC93342 homologue to GP|20339364|gb|AAM19355.1 UOS1 {Pisum sativ... 24 9.3
TC80100 similar to GP|21618014|gb|AAM67064.1 unknown {Arabidopsi... 24 9.3
>AW683931 weakly similar to GP|21553880|gb| unknown {Arabidopsis thaliana},
partial (6%)
Length = 152
Score = 42.0 bits (97), Expect = 4e-05
Identities = 20/24 (83%), Positives = 22/24 (91%)
Frame = +2
Query: 14 KRSSQKKLEEELMNVNNQIAELNS 37
+RSS+KKLEEELM VNNQI ELNS
Sbjct: 8 ERSSRKKLEEELMEVNNQITELNS 79
>TC79652 weakly similar to GP|19698951|gb|AAL91211.1 unknown protein
{Arabidopsis thaliana}, partial (20%)
Length = 1590
Score = 33.1 bits (74), Expect = 0.020
Identities = 16/63 (25%), Positives = 38/63 (59%)
Frame = +2
Query: 10 DLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVFSDRP 69
DL+ +R ++K++E++L + L ++ E + + ++E+ ++++KC++ DR
Sbjct: 602 DLEKERFAKKRVEKDLEVARRNFSHLKAQD-EDSSETDKLQQELGEYRDIVKCSICRDRT 778
Query: 70 KEV 72
KEV
Sbjct: 779 KEV 787
>AW685168 weakly similar to PIR|T01757|T0 hypothetical protein A_IG002P16.20
- Arabidopsis thaliana, partial (17%)
Length = 607
Score = 33.1 bits (74), Expect = 0.020
Identities = 14/34 (41%), Positives = 23/34 (67%)
Frame = +2
Query: 8 EKDLDSKRSSQKKLEEELMNVNNQIAELNSETGE 41
+KDL S+ SSQK+L +E+ + I E +++GE
Sbjct: 446 QKDLKSRESSQKQLRDEVFRIERDIMEALAKSGE 547
>BG451815 similar to GP|21537091|gb unknown {Arabidopsis thaliana}, partial
(36%)
Length = 664
Score = 32.7 bits (73), Expect = 0.026
Identities = 17/63 (26%), Positives = 35/63 (54%)
Frame = +2
Query: 17 SQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVFSDRPKEVKTGQ 76
+QK L ++ + + + N + VV++L+E++E R C VF D+P++++ Q
Sbjct: 383 NQKALAQQHFFI*DDFIDANIQLVLAVVKKLQEKKETRAC------VVFPDKPEKLRASQ 544
Query: 77 LIQ 79
L +
Sbjct: 545 LFK 553
>TC80811 similar to GP|21537091|gb|AAM61432.1 unknown {Arabidopsis
thaliana}, partial (39%)
Length = 655
Score = 31.6 bits (70), Expect = 0.058
Identities = 20/69 (28%), Positives = 36/69 (51%)
Frame = +2
Query: 11 LDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVFSDRPK 70
L S SS K ++ ++ N Q+ VV++L+E++E R C VF D+P+
Sbjct: 338 LPSNISSYKGSSDDFIDANIQLVL-------AVVKKLQEKKETRAC------VVFPDKPE 478
Query: 71 EVKTGQLIQ 79
+++ QL +
Sbjct: 479 KLRASQLFK 505
>TC77716 similar to GP|20465587|gb|AAM20276.1 unknown protein {Arabidopsis
thaliana}, partial (63%)
Length = 2392
Score = 28.9 bits (63), Expect = 0.38
Identities = 16/53 (30%), Positives = 25/53 (46%)
Frame = +3
Query: 19 KKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVFSDRPKE 71
++LE L+ NN + + E VQ +E + + C N I+ D PKE
Sbjct: 660 QELERALLGDNNDECDDDEENMFRTVQSMEIDPDFAECANPIQNKSHHDSPKE 818
>TC88580 similar to GP|9758308|dbj|BAB08782.1 gene_id:MUA2.4~unknown protein
{Arabidopsis thaliana}, partial (51%)
Length = 1388
Score = 27.7 bits (60), Expect = 0.84
Identities = 16/73 (21%), Positives = 41/73 (55%), Gaps = 5/73 (6%)
Frame = +1
Query: 8 EKDLDSKRSSQKKLEEELMNVNNQI-----AELNSETGETVVQQLEEEEEIRVCKNMIKC 62
+ +D+ +++ +L +E + +++ A + SE E + QQ +++EE+ V K+
Sbjct: 898 QASIDAFANARFELPQETLEAGDEVVASLAAPVTSEQNEEI-QQKQKQEEVEVEKDPFAA 1074
Query: 63 TVFSDRPKEVKTG 75
+ ++P+E+ +G
Sbjct: 1075SDAINKPQELVSG 1113
>TC80039 homologue to GP|6437558|gb|AAF08585.1| putative ATPase (ISW2-like)
{Arabidopsis thaliana}, partial (47%)
Length = 1840
Score = 26.2 bits (56), Expect = 2.4
Identities = 13/32 (40%), Positives = 19/32 (58%)
Frame = +2
Query: 7 SEKDLDSKRSSQKKLEEELMNVNNQIAELNSE 38
S+ + S +S EEE+ NVN+QI E + E
Sbjct: 89 SKSHVSSASNSSSSEEEEVENVNDQIEEEDEE 184
>TC79394 weakly similar to PIR|T49019|T49019 probable RNA binding protein -
Arabidopsis thaliana, partial (74%)
Length = 1909
Score = 25.8 bits (55), Expect = 3.2
Identities = 17/56 (30%), Positives = 30/56 (53%), Gaps = 8/56 (14%)
Frame = +2
Query: 5 SASEKDL----DSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQ----LEEEEE 52
S + DL D + SS++++E E + V ++ E E E V++ L+EE+E
Sbjct: 224 SGEQVDLDGDNDQEESSEEEVEYEEVEVEEEVEEEEEEEEEEEVEEESKPLDEEDE 391
>TC80043 similar to GP|9759561|dbj|BAB11163.1 MtN21 nodulin protein-like
{Arabidopsis thaliana}, partial (28%)
Length = 1421
Score = 25.0 bits (53), Expect = 5.4
Identities = 13/44 (29%), Positives = 22/44 (49%)
Frame = +2
Query: 14 KRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCK 57
K K++EEE + + N ETV++ +EE +I + K
Sbjct: 431 KSKENKEIEEETITEGMKCCVENGVILETVIEGVEETNDIEMQK 562
>TC90662 similar to GP|22136654|gb|AAM91646.1 unknown protein {Arabidopsis
thaliana}, partial (89%)
Length = 909
Score = 25.0 bits (53), Expect = 5.4
Identities = 15/48 (31%), Positives = 22/48 (45%)
Frame = -2
Query: 6 ASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEI 53
A+ +D D++ + E M NNQ LNS E V+ E+I
Sbjct: 392 AAYQDQDTESE*HQNDREPRMRSNNQQKALNSHQNEVVLLHCNHIEQI 249
>TC93449 similar to GP|21328382|gb|AAM48550.1 Hypothetical protein W02D3.12
{Caenorhabditis elegans}, partial (33%)
Length = 631
Score = 24.6 bits (52), Expect = 7.1
Identities = 13/30 (43%), Positives = 20/30 (66%)
Frame = +3
Query: 12 DSKRSSQKKLEEELMNVNNQIAELNSETGE 41
D + SQK+ EEE + ++ I ++N ETGE
Sbjct: 318 DDDKQSQKEEEEEEEDNDDDI-DMNKETGE 404
>BQ147955 weakly similar to GP|15290125|dbj putative kinase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 592
Score = 24.3 bits (51), Expect = 9.3
Identities = 15/39 (38%), Positives = 21/39 (53%)
Frame = +3
Query: 2 SVASASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETG 40
SVA + ++ +K SSQK LEEE V +A +G
Sbjct: 375 SVAEIAPLNISTKISSQKVLEEEKGMVLQNLATERPASG 491
>AW696663 weakly similar to GP|17827193|dbj ORF1 {TT virus}, partial (2%)
Length = 660
Score = 24.3 bits (51), Expect = 9.3
Identities = 13/29 (44%), Positives = 18/29 (61%)
Frame = -1
Query: 24 ELMNVNNQIAELNSETGETVVQQLEEEEE 52
E+ N N+ +++ET QQLEEEEE
Sbjct: 231 EM*NSNHPTLCVHAETSYHQQQQLEEEEE 145
>TC84059 weakly similar to PIR|T02508|T02508 hypothetical protein At2g38370
[imported] - Arabidopsis thaliana, partial (20%)
Length = 905
Score = 24.3 bits (51), Expect = 9.3
Identities = 12/55 (21%), Positives = 29/55 (51%)
Frame = -1
Query: 7 SEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIK 61
+ K +++ S+K++ + ++ V+ E NS + + E EE++ CK ++
Sbjct: 836 TRKAREAEEQSKKRVADAMLEVD----EANSSQMDVFKRVEEATEEVQTCKKALE 684
>AW695134
Length = 587
Score = 24.3 bits (51), Expect = 9.3
Identities = 13/33 (39%), Positives = 18/33 (54%), Gaps = 2/33 (6%)
Frame = +1
Query: 23 EELMNVNNQIAELNSET--GETVVQQLEEEEEI 53
E + NNQI EL SET + V+ L E ++
Sbjct: 256 EAIKRTNNQIPELKSETKLDQPVIPSLINETQV 354
>TC93342 homologue to GP|20339364|gb|AAM19355.1 UOS1 {Pisum sativum},
partial (33%)
Length = 857
Score = 24.3 bits (51), Expect = 9.3
Identities = 11/42 (26%), Positives = 22/42 (52%)
Frame = +3
Query: 33 AELNSETGETVVQQLEEEEEIRVCKNMIKCTVFSDRPKEVKT 74
A+L + G+ + ++ EE R+C ++ D+ EVK+
Sbjct: 369 ADLIFDQGDNITGKISREEVARMCVAALESPYACDKTFEVKS 494
>TC80100 similar to GP|21618014|gb|AAM67064.1 unknown {Arabidopsis
thaliana}, partial (59%)
Length = 740
Score = 24.3 bits (51), Expect = 9.3
Identities = 12/60 (20%), Positives = 27/60 (45%), Gaps = 2/60 (3%)
Frame = +2
Query: 4 ASASEKDLDSKRSSQKKLEEELMNVNNQIAELNS--ETGETVVQQLEEEEEIRVCKNMIK 61
A+ + D ++ E +N ++ S + ++QQ+ E ++ R+ NM+K
Sbjct: 167 ATPTNNDYSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVK 346
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.304 0.122 0.303
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,513,809
Number of Sequences: 36976
Number of extensions: 11132
Number of successful extensions: 81
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 81
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 81
length of query: 80
length of database: 9,014,727
effective HSP length: 56
effective length of query: 24
effective length of database: 6,944,071
effective search space: 166657704
effective search space used: 166657704
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.9 bits)
S2: 51 (24.3 bits)
Medicago: description of AC147712.6


