

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147712.16 + phase: 0
(236 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
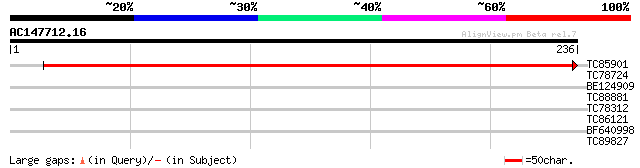
TC85901 similar to GP|20160828|dbj|BAB89768. contains ESTs C9802... 437 e-123
TC78724 similar to GP|4322475|gb|AAD16052.1| putative MADS box t... 28 3.1
BE124909 similar to GP|4322475|gb| putative MADS box transcripti... 28 3.1
TC88881 homologue to GP|15005009|dbj|BAB62076. APG1 {Arabidopsis... 28 4.0
TC78312 similar to GP|19698885|gb|AAL91178.1 putative protein {A... 27 6.8
TC86121 similar to GP|15027971|gb|AAK76516.1 unknown protein {Ar... 27 8.9
BF640998 similar to GP|22136786|gb unknown protein {Arabidopsis ... 27 8.9
TC89827 similar to GP|22654962|gb|AAM98074.1 AT5g60420/muf9_70 {... 27 8.9
>TC85901 similar to GP|20160828|dbj|BAB89768. contains ESTs C98021(C0384)
D15269(C0384)~similar to Arabidopsis thaliana chromosome
3 At3g13230~, partial (88%)
Length = 1484
Score = 437 bits (1124), Expect = e-123
Identities = 220/222 (99%), Positives = 221/222 (99%)
Frame = +1
Query: 15 SLNSCTMASSSMEVDNVLSEQKVLPPKPKFEPLKPHEMPGAAVQFRKVSVPPHRYTPLKK 74
S +SCTMASSSMEVDNVLSEQKVLPPKPKFEPLKPHEMPGAAVQFRKVSVPPHRYTPLKK
Sbjct: 541 SPSSCTMASSSMEVDNVLSEQKVLPPKPKFEPLKPHEMPGAAVQFRKVSVPPHRYTPLKK 720
Query: 75 VWMDIYTPVFEQMKIDIRMNLKGRKVELKTRHDTPDISNLQKCADFVHAFMLGFDVIDAI 134
VWMDIYTPVFEQMKIDIRMNLKGRKVELKTRHDTPDISNLQKCADFVHAFMLGFDVIDAI
Sbjct: 721 VWMDIYTPVFEQMKIDIRMNLKGRKVELKTRHDTPDISNLQKCADFVHAFMLGFDVIDAI 900
Query: 135 AILRLDELYVESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKFTIENASKTRIVIADTKIH 194
AILRLDELYVESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKFTIENASKTRIVIADTKIH
Sbjct: 901 AILRLDELYVESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKFTIENASKTRIVIADTKIH 1080
Query: 195 ILGSFANIKIARDSLCYLIMGSPAGKVYSKLRAVTSRMAERF 236
ILGSFANIKIARDSLCYLIMGSPAGKVYSKLRAVTSRMAERF
Sbjct: 1081ILGSFANIKIARDSLCYLIMGSPAGKVYSKLRAVTSRMAERF 1206
>TC78724 similar to GP|4322475|gb|AAD16052.1| putative MADS box
transcription factor ETL {Eucalyptus globulus subsp.
globulus}, partial (71%)
Length = 1089
Score = 28.1 bits (61), Expect = 3.1
Identities = 18/66 (27%), Positives = 32/66 (48%), Gaps = 1/66 (1%)
Frame = +1
Query: 172 GKTKFT-IENASKTRIVIADTKIHILGSFANIKIARDSLCYLIMGSPAGKVYSKLRAVTS 230
GKT+ IEN S ++ + + +L + + D+ LI+ S GK+Y + S
Sbjct: 235 GKTQMKRIENESNRQVTFSKRRNGLLKKAFELSVLCDAEVALIVFSTTGKLYEFSSSSIS 414
Query: 231 RMAERF 236
+ ER+
Sbjct: 415 KTVERY 432
>BE124909 similar to GP|4322475|gb| putative MADS box transcription factor
ETL {Eucalyptus globulus subsp. globulus}, partial (60%)
Length = 608
Score = 28.1 bits (61), Expect = 3.1
Identities = 18/66 (27%), Positives = 32/66 (48%), Gaps = 1/66 (1%)
Frame = +1
Query: 172 GKTKFT-IENASKTRIVIADTKIHILGSFANIKIARDSLCYLIMGSPAGKVYSKLRAVTS 230
GKT+ IEN S ++ + + +L + + D+ LI+ S GK+Y + S
Sbjct: 157 GKTQMKRIENESNRQVTFSKRRNGLLKKAFELSVLCDAEVALIVFSTTGKLYEFSSSSIS 336
Query: 231 RMAERF 236
+ ER+
Sbjct: 337 KTVERY 354
>TC88881 homologue to GP|15005009|dbj|BAB62076. APG1 {Arabidopsis thaliana},
partial (87%)
Length = 1242
Score = 27.7 bits (60), Expect = 4.0
Identities = 16/35 (45%), Positives = 20/35 (56%)
Frame = -3
Query: 31 VLSEQKVLPPKPKFEPLKPHEMPGAAVQFRKVSVP 65
V E K+LP K EP KP+ P +QF+ VS P
Sbjct: 184 VFGEIKLLPLKS--EPDKPNPFPFWVLQFKNVSFP 86
>TC78312 similar to GP|19698885|gb|AAL91178.1 putative protein {Arabidopsis
thaliana}, partial (14%)
Length = 1317
Score = 26.9 bits (58), Expect = 6.8
Identities = 15/42 (35%), Positives = 21/42 (49%)
Frame = -1
Query: 32 LSEQKVLPPKPKFEPLKPHEMPGAAVQFRKVSVPPHRYTPLK 73
LS +LPP F P +PH +P +QF + P + LK
Sbjct: 1131 LSAAPLLPP---FSPQQPHFLPQPVIQFHSPN*PKNCSIQLK 1015
>TC86121 similar to GP|15027971|gb|AAK76516.1 unknown protein {Arabidopsis
thaliana}, partial (89%)
Length = 1882
Score = 26.6 bits (57), Expect = 8.9
Identities = 10/22 (45%), Positives = 14/22 (63%)
Frame = +3
Query: 60 RKVSVPPHRYTPLKKVWMDIYT 81
+KV VP +KVWMD+Y+
Sbjct: 1320 KKVKVPTFLVPATQKVWMDVYS 1385
>BF640998 similar to GP|22136786|gb unknown protein {Arabidopsis thaliana},
partial (61%)
Length = 496
Score = 26.6 bits (57), Expect = 8.9
Identities = 12/28 (42%), Positives = 15/28 (52%)
Frame = -2
Query: 95 LKGRKVELKTRHDTPDISNLQKCADFVH 122
++G KV L RHDTP C F+H
Sbjct: 318 IRGIKVNLVIRHDTPHQLLSAYCFHFIH 235
>TC89827 similar to GP|22654962|gb|AAM98074.1 AT5g60420/muf9_70 {Arabidopsis
thaliana}, partial (46%)
Length = 1378
Score = 26.6 bits (57), Expect = 8.9
Identities = 21/57 (36%), Positives = 27/57 (46%), Gaps = 1/57 (1%)
Frame = +1
Query: 102 LKTRHDTPDISNLQKCADFVHA-FMLGFDVIDAIAILRLDELYVESFEIKDVKTLRG 157
L RH I L + D V A F LGF +D +A ++ Y E+KDV T G
Sbjct: 58 LLRRHHVLIIRFLIESVDSVFAEFFLGFVEMDLVAGIKEKLTYFRIKELKDVLTQLG 228
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.138 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,145,053
Number of Sequences: 36976
Number of extensions: 98091
Number of successful extensions: 587
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 586
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 587
length of query: 236
length of database: 9,014,727
effective HSP length: 93
effective length of query: 143
effective length of database: 5,575,959
effective search space: 797362137
effective search space used: 797362137
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC147712.16


