

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147538.3 - phase: 0
(123 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
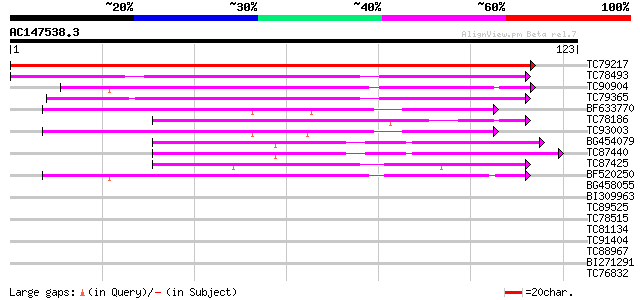
Score E
Sequences producing significant alignments: (bits) Value
TC79217 weakly similar to GP|21553541|gb|AAM62634.1 lipid transf... 243 1e-65
TC78493 weakly similar to GP|15010698|gb|AAK74008.1 AT3g22600/F1... 61 9e-11
TC90904 similar to GP|3128176|gb|AAC16080.1| unknown protein {Ar... 61 1e-10
TC79365 weakly similar to GP|15010698|gb|AAK74008.1 AT3g22600/F1... 59 5e-10
BF633770 48 1e-06
TC78186 weakly similar to GP|3128176|gb|AAC16080.1| unknown prot... 45 5e-06
TC93003 45 7e-06
BG454079 similar to GP|10176956|dbj contains similarity to nonsp... 43 3e-05
TC87440 similar to GP|17979494|gb|AAL50083.1 AT5g64080/MHJ24_6 {... 43 3e-05
TC87425 weakly similar to GP|20161478|dbj|BAB90402. contains EST... 41 1e-04
BF520250 weakly similar to GP|3128176|gb| unknown protein {Arabi... 39 4e-04
BG458055 weakly similar to GP|16904376|gb| lipid transfer protei... 38 0.001
BI309963 37 0.002
TC89525 37 0.002
TC78515 36 0.004
TC81134 weakly similar to GP|14334664|gb|AAK59510.1 unknown prot... 36 0.004
TC91404 35 0.006
TC88967 weakly similar to GP|18477856|emb|CAC86258. lipid transf... 35 0.006
BI271291 similar to GP|6782436|gb|A lipid-transfer protein {Nico... 35 0.007
TC76832 weakly similar to GP|21592534|gb|AAM64483.1 unknown {Ara... 35 0.007
>TC79217 weakly similar to GP|21553541|gb|AAM62634.1 lipid transfer protein
putative {Arabidopsis thaliana}, partial (43%)
Length = 992
Score = 243 bits (621), Expect = 1e-65
Identities = 114/114 (100%), Positives = 114/114 (100%)
Frame = +1
Query: 1 MKNQQHQMFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDA 60
MKNQQHQMFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDA
Sbjct: 178 MKNQQHQMFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDA 357
Query: 61 ANSIKATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCP 114
ANSIKATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCP
Sbjct: 358 ANSIKATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCP 519
>TC78493 weakly similar to GP|15010698|gb|AAK74008.1 AT3g22600/F16J14_17
{Arabidopsis thaliana}, partial (57%)
Length = 797
Score = 61.2 bits (147), Expect = 9e-11
Identities = 33/113 (29%), Positives = 53/113 (46%)
Frame = +2
Query: 1 MKNQQHQMFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDA 60
M + + M + L V+A++ G S C +V+ + PCL++ TG + TP CC
Sbjct: 71 MAHSKMNMNLVLVVIAMMCAGATAQ----SSCTNVLVSLSPCLNYITGNSSTPSSGCCSN 238
Query: 61 ANSIKATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDC 113
S+ ++ P CLC ++ G S + I + + L LP C V S C
Sbjct: 239 LASVVSSQPLCLCQVL----GGGASSLGISINQTQALALPGACKVQTPPTSQC 385
>TC90904 similar to GP|3128176|gb|AAC16080.1| unknown protein {Arabidopsis
thaliana}, partial (28%)
Length = 672
Score = 60.8 bits (146), Expect = 1e-10
Identities = 33/104 (31%), Positives = 55/104 (52%), Gaps = 1/104 (0%)
Frame = +3
Query: 12 LCVLALIIG-GCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPE 70
L + ++I+G G + + +C + + + CL + G+ TP +CCD ++ T+ +
Sbjct: 96 LLLASMIVGIGMSDSSKEKQECTAQLTGLASCLPYVEGEGKTPAPDCCDGLKTLLKTNKK 275
Query: 71 CLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCP 114
CLC II+ + P+ + I L LPTVC+ ANIS CP
Sbjct: 276 CLCVIIKD--RNDPDLGGIVINVTLALNLPTVCNA-PANISKCP 398
>TC79365 weakly similar to GP|15010698|gb|AAK74008.1 AT3g22600/F16J14_17
{Arabidopsis thaliana}, partial (51%)
Length = 783
Score = 58.9 bits (141), Expect = 5e-10
Identities = 29/105 (27%), Positives = 48/105 (45%)
Frame = +2
Query: 9 FMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATD 68
++ V+ L+ C E S C S + + PCL++ G + P CC +S+ +
Sbjct: 68 YILFLVMVLVANMCTQNE-AQSGCTSTITSLSPCLNYIMGSSSNPSSSCCSQLSSVVQSS 244
Query: 69 PECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDC 113
P+CLC ++ G S + I + L LP+ C V +S C
Sbjct: 245 PQCLCSLL----NGGGSSFGITINQTLALSLPSACKVQTPPVSQC 367
>BF633770
Length = 510
Score = 47.8 bits (112), Expect = 1e-06
Identities = 26/106 (24%), Positives = 49/106 (45%), Gaps = 7/106 (6%)
Frame = +1
Query: 8 MFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAP-TPKKECCDAANSI-- 64
+F+ L VL++++ ++ C + ++PCL F G P TP +CC A ++
Sbjct: 67 LFVPLMVLSMLMTTLYASQINDITCSEAISSLLPCLPFLEGSLPATPSTDCCAGATNLFH 246
Query: 65 ----KATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
+ +C+ ++ T S G+ + QLP +CH+N
Sbjct: 247 KVDTTKIQRKNICHCLKNT------STKFGVNSKRSKQLPQLCHIN 366
>TC78186 weakly similar to GP|3128176|gb|AAC16080.1| unknown protein
{Arabidopsis thaliana}, partial (32%)
Length = 766
Score = 45.4 bits (106), Expect = 5e-06
Identities = 23/85 (27%), Positives = 41/85 (48%), Gaps = 3/85 (3%)
Frame = +2
Query: 32 CGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHK---GSPESKS 88
C + + CL + G A TP +CC + +C+C +I+ ++ G P + +
Sbjct: 95 CADKLVTLASCLPYVGGSANTPTIDCCTNLKQVLNNTKKCICILIKDSNDPKLGFPMNAT 274
Query: 89 MGIQEDKLLQLPTVCHVNGANISDC 113
+ + QLP CH+ +NIS+C
Sbjct: 275 LAV------QLPNACHI-PSNISEC 328
>TC93003
Length = 691
Score = 45.1 bits (105), Expect = 7e-06
Identities = 25/106 (23%), Positives = 47/106 (43%), Gaps = 7/106 (6%)
Frame = +1
Query: 8 MFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAP-TPKKECCDAANS--- 63
+F+ L L++++ ++ C + ++PCL F G P TP +CC A +
Sbjct: 67 LFVPLMALSMLMTTLYASQINDITCSEAISSLLPCLPFLEGSLPATPSTDCCAGATN*FH 246
Query: 64 ---IKATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
+ +C+ ++ T S G+ + QLP +CH+N
Sbjct: 247 KVDTTKIQRKNICHCLKNT------STKFGVNSKRSKQLPQLCHIN 366
>BG454079 similar to GP|10176956|dbj contains similarity to nonspecific
lipid-transfer protein~gene_id:MHJ24.6 {Arabidopsis
thaliana}, partial (50%)
Length = 418
Score = 43.1 bits (100), Expect = 3e-05
Identities = 26/87 (29%), Positives = 38/87 (42%), Gaps = 2/87 (2%)
Frame = +1
Query: 32 CGSVVQKVIPCLDFATGKAPTPKKE--CCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
C ++V + CL F T + T K E CC S+ T P CLC + K S + +
Sbjct: 121 CTNLVLTMADCLSFVTNGSTTTKPEGTCCSGLKSVLKTAPSCLC----EAFKSSAQF-GV 285
Query: 90 GIQEDKLLQLPTVCHVNGANISDCPSK 116
+ K LP C V+ + + C K
Sbjct: 286 VLNVTKATSLPAACKVSAPSATKCGCK 366
>TC87440 similar to GP|17979494|gb|AAL50083.1 AT5g64080/MHJ24_6 {Arabidopsis
thaliana}, partial (48%)
Length = 818
Score = 42.7 bits (99), Expect = 3e-05
Identities = 26/91 (28%), Positives = 39/91 (42%), Gaps = 2/91 (2%)
Frame = +1
Query: 32 CGSVVQKVIPCLDFATGKAPTPKKE--CCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
C ++V + CL F T + T K E CC S+ T P CLC + K S + +
Sbjct: 133 CTNLVLTMADCLSFVTNGSTTTKPEGTCCSGLKSVLKTAPSCLC----EAFKSSAQF-GV 297
Query: 90 GIQEDKLLQLPTVCHVNGANISDCPSKSKTD 120
+ K LP C V+ + + C T+
Sbjct: 298 VLNVTKATSLPAACKVSAPSATKCGLSEVTE 390
>TC87425 weakly similar to GP|20161478|dbj|BAB90402. contains ESTs
C74501(E31745) AU094804(E31745)~unknown protein, partial
(24%)
Length = 929
Score = 40.8 bits (94), Expect = 1e-04
Identities = 27/86 (31%), Positives = 39/86 (44%), Gaps = 4/86 (4%)
Frame = +1
Query: 32 CGSVVQKVIPCLDFAT--GKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
C + + CL F K P K CC + +P CLC ++ GS + S
Sbjct: 208 CLMALTNMSDCLTFVEDGSKLTKPDKGCCPELAGLIDGNPICLCKLL-----GSNTADSF 372
Query: 90 GIQ--EDKLLQLPTVCHVNGANISDC 113
GI+ +K L+LPT+C V +S C
Sbjct: 373 GIKINVNKALKLPTICGVTTPPVSAC 450
>BF520250 weakly similar to GP|3128176|gb| unknown protein {Arabidopsis
thaliana}, partial (69%)
Length = 541
Score = 39.3 bits (90), Expect = 4e-04
Identities = 26/107 (24%), Positives = 46/107 (42%), Gaps = 1/107 (0%)
Frame = +3
Query: 8 MFMCLCVLALIIG-GCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKA 66
+F L L L+ G + ++C + + + CL F T +A +P +CC +
Sbjct: 51 LFSTLICLVLMFGLVTSDINQDKAECTNKLLTLAGCLPFVTNQAKSPTIDCCTGVKEVAD 230
Query: 67 TDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDC 113
CLC +I+ + + I L+LP C+ + NI+ C
Sbjct: 231 KSKRCLCILIKD---HDDPNLGLTINVTLALKLPNDCN-SPTNITQC 359
>BG458055 weakly similar to GP|16904376|gb| lipid transfer protein {Setaria
italica}, partial (61%)
Length = 621
Score = 37.7 bits (86), Expect = 0.001
Identities = 27/101 (26%), Positives = 44/101 (42%), Gaps = 5/101 (4%)
Frame = +2
Query: 11 CLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIK----- 65
CL ++ ++ G D S C V K++PC+ + TG + CCD +I
Sbjct: 53 CLTMMMCMVLGLPQTLDALS-CLQVETKLMPCVPYVTGNGGYVPQPCCDGVKAINNQAVT 229
Query: 66 ATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
+D + C I+ + S G+ D L LP+ C V+
Sbjct: 230 KSDRQAACRCIK-----TATSAIHGLNMDILAGLPSKCGVH 337
>BI309963
Length = 529
Score = 37.0 bits (84), Expect = 0.002
Identities = 27/105 (25%), Positives = 42/105 (39%), Gaps = 5/105 (4%)
Frame = +1
Query: 11 CLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDA-----ANSIK 65
CL + L++G + C ++ CL + P+P + CC A +I
Sbjct: 28 CLATICLVLG--IPLANAVITCRDAAITLMGCLPYVAHPTPSPPQNCCAAVLDVTGQAIT 201
Query: 66 ATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANI 110
D + +C ++ G P G+ L LP VC GANI
Sbjct: 202 REDRQAVCSCLKGLMNGIP-----GLDLTALASLPKVC---GANI 312
>TC89525
Length = 655
Score = 37.0 bits (84), Expect = 0.002
Identities = 17/55 (30%), Positives = 31/55 (55%), Gaps = 1/55 (1%)
Frame = +2
Query: 11 CLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGK-APTPKKECCDAANSI 64
CL ++ L++G + A+ C + Q + PCL++ T +P P + CC+A +I
Sbjct: 68 CLAMICLVLG--IPLANAAASCPEIEQTLTPCLEYVTHPGSPPPPEPCCNAVKAI 226
>TC78515
Length = 639
Score = 35.8 bits (81), Expect = 0.004
Identities = 17/55 (30%), Positives = 28/55 (50%), Gaps = 1/55 (1%)
Frame = +2
Query: 11 CLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDF-ATGKAPTPKKECCDAANSI 64
CL V+ L++ D KC +V + +IPC+++ T A P CC+ S+
Sbjct: 83 CLSVMCLLLAIPLANADPDPKCKNVAESIIPCVEYIMTPDASNPPAPCCNGMTSL 247
>TC81134 weakly similar to GP|14334664|gb|AAK59510.1 unknown protein
{Arabidopsis thaliana}, partial (42%)
Length = 900
Score = 35.8 bits (81), Expect = 0.004
Identities = 20/77 (25%), Positives = 36/77 (45%), Gaps = 7/77 (9%)
Frame = +3
Query: 39 VIPCLDFAT---GKAPTPKKECCDAANSIKATDPECLCYIIQ-QTHKGSPESKSMGIQED 94
+ PCL F T G +P +CC+A ++ + +C+C I P ++++ I
Sbjct: 177 ISPCLSFLTNSSGNGTSPTADCCNAIKTLTSGSKDCMCLIATGNVPFALPINRTLAISLP 356
Query: 95 KLLQLPTV---CHVNGA 108
+ LP V C +G+
Sbjct: 357 RACNLPGVPLQCKTSGS 407
>TC91404
Length = 595
Score = 35.4 bits (80), Expect = 0.006
Identities = 28/105 (26%), Positives = 42/105 (39%), Gaps = 5/105 (4%)
Frame = +3
Query: 11 CLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKA---- 66
CL ++ L++G + C + CL + P P + CCDA + A
Sbjct: 87 CLAMICLVLG--IPLANAVITCPDTDITLTACLSYVAYPNPPPPQPCCDAVLDVTAQART 260
Query: 67 -TDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANI 110
D + +C ++ G P G+ L LP VC GANI
Sbjct: 261 REDRQAVCSCLKGLMIGIP-----GLDLTALAALPKVC---GANI 371
>TC88967 weakly similar to GP|18477856|emb|CAC86258. lipid transfer protein
{Fragaria x ananassa}, partial (48%)
Length = 743
Score = 35.4 bits (80), Expect = 0.006
Identities = 26/103 (25%), Positives = 43/103 (41%), Gaps = 6/103 (5%)
Frame = +1
Query: 10 MCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDF-ATGKAPTPKKECCDAANSIK--- 65
+CL ++ L+ G AE + CG VV + PC+ + G P +CC+ ++
Sbjct: 76 VCLAIVCLVTFGPK-AEAAVTSCGPVVTSLYPCVSYIMNGGNTVPAAQCCNGIRNLNTMA 252
Query: 66 --ATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
D +C I+ S S + + + LP C VN
Sbjct: 253 QTTNDRRAVCTCIKNAVSQSGFSYT-NLNLNLAAGLPRKCGVN 378
>BI271291 similar to GP|6782436|gb|A lipid-transfer protein {Nicotiana
glauca}, partial (23%)
Length = 575
Score = 35.0 bits (79), Expect = 0.007
Identities = 26/96 (27%), Positives = 39/96 (40%), Gaps = 5/96 (5%)
Frame = +3
Query: 16 ALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSI-----KATDPE 70
AL+ G DL+ CG V+ + PCL + + CC+ + +D +
Sbjct: 90 ALVFGIPLANADLS--CGQVLSTLYPCLGYIRNPGASVPAPCCNGIRIVNDEAKNTSDRQ 263
Query: 71 CLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
+C ++ T GI D L LPT C VN
Sbjct: 264 SVCRCLKST------IVLPGINLDALANLPTNCGVN 353
>TC76832 weakly similar to GP|21592534|gb|AAM64483.1 unknown {Arabidopsis
thaliana}, partial (71%)
Length = 601
Score = 35.0 bits (79), Expect = 0.007
Identities = 26/111 (23%), Positives = 48/111 (42%), Gaps = 6/111 (5%)
Frame = +1
Query: 10 MCLCVLALIIGGCNGAEDLASKCG------SVVQKVIPCLDFATGKAPTPKKECCDAANS 63
+C+ +A+++ G + E A +CG + K+IPC A + + + CC
Sbjct: 76 LCIVGIAVLVAGTHRVES-AGECGRGTTPDNEAFKLIPCASAAKDENASVSQSCCAQVKK 252
Query: 64 IKATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCP 114
+ +P CLC ++ S +K G + +P C N++D P
Sbjct: 253 L-GQNPSCLCAVML-----SNVAKMSGANPQIAVTIPKRC-----NLADRP 372
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.134 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,608,449
Number of Sequences: 36976
Number of extensions: 61823
Number of successful extensions: 337
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 337
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 337
length of query: 123
length of database: 9,014,727
effective HSP length: 84
effective length of query: 39
effective length of database: 5,908,743
effective search space: 230440977
effective search space used: 230440977
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC147538.3


