

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
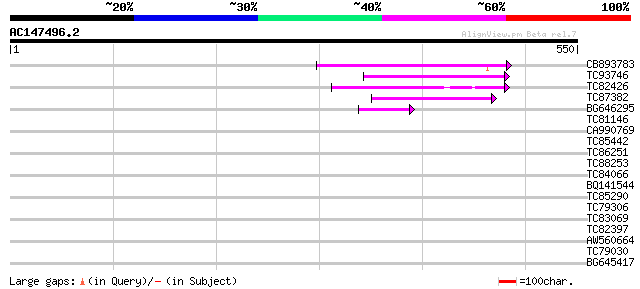
Query= AC147496.2 - phase: 0 /pseudo
(550 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CB893783 weakly similar to GP|22830935|dbj hypothetical protein~... 150 1e-36
TC93746 weakly similar to GP|22830935|dbj|BAC15800. hypothetical... 105 4e-23
TC82426 104 8e-23
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 89 4e-18
BG646295 similar to GP|13992688|gb Putative retroelement {Oryza ... 47 2e-05
TC81146 similar to GP|13543783|gb|AAH06040.1 Unknown (protein fo... 34 0.13
CA990769 weakly similar to GP|21594005|gb| unknown {Arabidopsis ... 33 0.22
TC85442 similar to GP|13543783|gb|AAH06040.1 Unknown (protein fo... 32 0.48
TC86251 similar to GP|22136176|gb|AAM91166.1 unknown protein {Ar... 31 1.1
TC88253 weakly similar to GP|15810597|gb|AAL07186.1 unknown prot... 31 1.1
TC84066 similar to GP|23498475|emb|CAD50426. ribonuclease putat... 31 1.1
BQ141544 similar to GP|7022293|dbj| unnamed protein product {Hom... 31 1.1
TC85290 similar to GP|7767653|gb|AAF69150.1| F27F5.2 {Arabidopsi... 31 1.1
TC79306 similar to PIR|F86397|F86397 protein T7N9.16 [imported] ... 30 1.8
TC83069 30 1.8
TC82397 similar to GP|18377877|gb|AAL67124.1 AT3g53630/F4P12_330... 30 2.4
AW560664 similar to GP|23495077|gb hypothetical protein {Plasmod... 30 2.4
TC79030 similar to PIR|H86366|H86366 protein F26F24.12 [imported... 30 3.1
BG645417 weakly similar to GP|14586370|em putative protein {Arab... 29 4.1
>CB893783 weakly similar to GP|22830935|dbj hypothetical protein~similar to
gag-pol polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 853
Score = 150 bits (379), Expect = 1e-36
Identities = 87/193 (45%), Positives = 112/193 (57%), Gaps = 4/193 (2%)
Frame = +2
Query: 298 ENHEGYVEEEAEEIPSGDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGG 357
E E +EE SG+ ++RR L +EE R +FHTR +QG VC +IID G
Sbjct: 104 EMGEQTYDEEPVLADSGESLIVRRSLHTTTVKEEPWLRHNIFHTR*TLQGIVCDVIIDSG 283
Query: 358 SRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIG-KYEDDVLCDV 416
S NV S +V K+EL TK HP YKLQWL + E+ V K V F IG KY+D+V CDV
Sbjct: 284 SCENVVSNYMVEKLELPTKDHPHRYKLQWLKKGNEVRVSKCCLVSFSIGQKYKDNVWCDV 463
Query: 417 VPMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPS---EVRDDQKKRKE 473
+ M+A H+LLGRP Q+DR L+DG N Y+F+ G KI L PL P+ E + D K
Sbjct: 464 ISMDACHMLLGRPWQYDRHALYDGHANTYTFVKYGVKIKLVPLPPNAFDEGKKDFKPIVS 643
Query: 474 KYEKEKRKIKRKE 486
KE K+ K+
Sbjct: 644 LVSKEPFKVTTKD 682
>TC93746 weakly similar to GP|22830935|dbj|BAC15800. hypothetical
protein~similar to gag-pol polyprotein {Oryza sativa
(japonica cultivar-group)}, partial (4%)
Length = 1019
Score = 105 bits (262), Expect = 4e-23
Identities = 57/143 (39%), Positives = 82/143 (56%), Gaps = 1/143 (0%)
Frame = +2
Query: 344 LVQGKVCFLIIDGGSRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCF 403
+ + VC +IID NV S +V K+++ TK HP PYKLQWLN++ E+ + + V F
Sbjct: 29 ITKDMVCNVIIDNKICENVESNYMVEKLKMPTKEHPHPYKLQWLNKDNEVKLSQHSAVSF 208
Query: 404 KI-GKYEDDVLCDVVPMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPS 462
I Y ++V CDV+PM+ H+ LGRP Q+DR L DG N +F+ KI APL +
Sbjct: 209 SIDNNYNENVWCDVIPMDTCHIHLGRPWQYDRHALFDGYANTCTFVTDAIKIKFAPLPLN 388
Query: 463 EVRDDQKKRKEKYEKEKRKIKRK 485
E +++ K + E K K K
Sbjct: 389 EGKEEFKPLGFLADMEPFKDKTK 457
>TC82426
Length = 1017
Score = 104 bits (260), Expect = 8e-23
Identities = 66/174 (37%), Positives = 94/174 (53%), Gaps = 1/174 (0%)
Frame = +3
Query: 313 SGDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKME 372
SG + RR L +EES + +FHT QGKVC +I++GGS NV S +V K+
Sbjct: 33 SGKSLVARRSLQTTTDKEESCLKHNIFHTSNTSQGKVCDVIVNGGSCENVVSNYMVEKLN 212
Query: 373 LETKPHPKPYKLQWLNENVEILVDKQVEVCFKIG-KYEDDVLCDVVPMEASHLLLGRP*Q 431
L TK PYKLQWLN++ E+ V + + F IG Y++++ CDV+ M+ RP Q
Sbjct: 213 LPTKERLHPYKLQWLNKDNEVKVSQHSIISFSIGNNYKENLWCDVISMDT-----WRPCQ 377
Query: 432 FDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQKKRKEKYEKEKRKIKRK 485
+DR L+D N +F+ I LAPL ++ + + K KE K K K
Sbjct: 378 YDRCALYDDYVNTCTFVK--DVIKLAPLPLNDGKAEFKLLGLSLAKEPFKDKTK 533
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 89.0 bits (219), Expect = 4e-18
Identities = 48/122 (39%), Positives = 72/122 (58%), Gaps = 1/122 (0%)
Frame = +3
Query: 352 LIIDGGSRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGK-YED 410
+IID S NVAS + +++ H PYKLQWLN++ E+ V + + F I K Y++
Sbjct: 1428 VIIDNKSGENVASNYIEEELKFRMINHQDPYKLQWLNKDNEVKVSQHSIISFSIDKNYKE 1607
Query: 411 DVLCDVVPMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQKK 470
V CD++PM+ H LG P Q+DR L+DG N +F+ G KI LA L +E + +++
Sbjct: 1608 KVWCDIIPMDTCHTHLGMPWQYDRRALYDGYANINTFVKYGIKIKLARLPLNEFIEGKEE 1787
Query: 471 RK 472
K
Sbjct: 1788 FK 1793
>BG646295 similar to GP|13992688|gb Putative retroelement {Oryza sativa}
[Oryza sativa (japonica cultivar-group)], partial (0%)
Length = 640
Score = 47.0 bits (110), Expect = 2e-05
Identities = 23/54 (42%), Positives = 32/54 (58%)
Frame = +3
Query: 339 FHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKPHPKPYKLQWLNENVE 392
FHT QGKV + I+ G N+ S +V ++L+TK H YKL WLN++ E
Sbjct: 3 FHTSNTSQGKVYDIYINSGICQNILSNYMVENLKLQTKEHLHRYKLHWLNKDNE 164
>TC81146 similar to GP|13543783|gb|AAH06040.1 Unknown (protein for MGC:7642)
{Mus musculus}, partial (77%)
Length = 1023
Score = 34.3 bits (77), Expect = 0.13
Identities = 18/39 (46%), Positives = 26/39 (66%), Gaps = 1/39 (2%)
Frame = +2
Query: 468 QKKRKEKYEKEKRKIKRK-EREFA*KKKKRRVSRNSHEQ 505
+KK+ K EK K+K +RK ER+ +KKKRR ++ H Q
Sbjct: 587 RKKKPRKMEKRKKKERRKRERKKMERKKKRRRKKHPHHQ 703
>CA990769 weakly similar to GP|21594005|gb| unknown {Arabidopsis thaliana},
partial (20%)
Length = 488
Score = 33.5 bits (75), Expect = 0.22
Identities = 17/62 (27%), Positives = 34/62 (54%)
Frame = +2
Query: 443 NKYSFMHSGQKISLAPLSPSEVRDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNS 502
N Y +H+ + + +R ++KRK+K +KE++K KR+ + +KKR+ +
Sbjct: 146 NCYLILHTPKWARKQNRNNQRLRRSKRKRKKKRQKEEKKRKRRRKAKRKSQKKRKRKSLN 325
Query: 503 HE 504
H+
Sbjct: 326 HQ 331
>TC85442 similar to GP|13543783|gb|AAH06040.1 Unknown (protein for MGC:7642)
{Mus musculus}, partial (62%)
Length = 585
Score = 32.3 bits (72), Expect = 0.48
Identities = 15/36 (41%), Positives = 23/36 (63%)
Frame = +1
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSH 503
+KK K + + K+K KRK R+ K+KKRR ++ H
Sbjct: 145 RKKGKRRKNQRKKKRKRKRRKKKGKRKKRRKRKSQH 252
Score = 31.6 bits (70), Expect = 0.83
Identities = 15/33 (45%), Positives = 23/33 (69%)
Frame = +1
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSR 500
+KKRK+K ++KR+ K K R+ KKK++R R
Sbjct: 106 KKKRKKKQRRKKRRKKGKRRKNQRKKKRKRKRR 204
>TC86251 similar to GP|22136176|gb|AAM91166.1 unknown protein {Arabidopsis
thaliana}, partial (37%)
Length = 1636
Score = 31.2 bits (69), Expect = 1.1
Identities = 12/23 (52%), Positives = 18/23 (78%)
Frame = +1
Query: 466 DDQKKRKEKYEKEKRKIKRKERE 488
DD+KK+K+K EK K+K K++E
Sbjct: 685 DDEKKKKKKNVNEKEKVKEKKKE 753
>TC88253 weakly similar to GP|15810597|gb|AAL07186.1 unknown protein
{Arabidopsis thaliana}, partial (51%)
Length = 1122
Score = 31.2 bits (69), Expect = 1.1
Identities = 15/42 (35%), Positives = 24/42 (56%)
Frame = +1
Query: 465 RDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQT 506
R ++KRK +K K+K+K+K +E K K V H++T
Sbjct: 724 RKKKRKRKRSKKKRKKKVKKKTKEKMIIKLKSNVVNIGHQKT 849
Score = 30.8 bits (68), Expect = 1.4
Identities = 15/38 (39%), Positives = 26/38 (67%)
Frame = +1
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQ 505
+KKR++K K+KRK KR + KK+K++V + + E+
Sbjct: 700 RKKRRKKRRKKKRKRKRSK-----KKRKKKVKKKTKEK 798
>TC84066 similar to GP|23498475|emb|CAD50426. ribonuclease putative
{Plasmodium falciparum 3D7}, partial (0%)
Length = 481
Score = 31.2 bits (69), Expect = 1.1
Identities = 15/30 (50%), Positives = 21/30 (70%)
Frame = +2
Query: 467 DQKKRKEKYEKEKRKIKRKEREFA*KKKKR 496
+ K RKEK + +K+K K KER+ KKKK+
Sbjct: 323 NSKPRKEKEKGKKKKRKEKERKTVMKKKKK 412
Score = 29.6 bits (65), Expect = 3.1
Identities = 20/75 (26%), Positives = 38/75 (50%), Gaps = 2/75 (2%)
Frame = +2
Query: 435 SVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDD--QKKRKEKYEKEKRKIKRKEREFA*K 492
+++H R+ F S +I+ P + ++ +KKRKEK K K K+K+R+ K
Sbjct: 254 AMIHKLRSRTQKFNDSKTRIN*MNSKPRKEKEKGKKKKRKEKERKTVMKKKKKDRKGKGK 433
Query: 493 KKKRRVSRNSHEQTA 507
K + + ++T+
Sbjct: 434 GKGKNENAGKSKKTS 478
>BQ141544 similar to GP|7022293|dbj| unnamed protein product {Homo sapiens},
partial (5%)
Length = 1284
Score = 31.2 bits (69), Expect = 1.1
Identities = 16/33 (48%), Positives = 22/33 (66%)
Frame = +2
Query: 465 RDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRR 497
R+ +KR +KY+ E K KR+EREF +K RR
Sbjct: 311 REGARKRIKKYKNE*GKGKREEREFRHEKDPRR 409
>TC85290 similar to GP|7767653|gb|AAF69150.1| F27F5.2 {Arabidopsis
thaliana}, partial (32%)
Length = 1558
Score = 31.2 bits (69), Expect = 1.1
Identities = 14/38 (36%), Positives = 25/38 (64%)
Frame = +3
Query: 469 KKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQT 506
KK KE+ EKEKRK K K+ + ++K++ R+ +++
Sbjct: 768 KKEKEREEKEKRKEKEKKEKEREREKEKSKERHKKDES 881
Score = 30.4 bits (67), Expect = 1.8
Identities = 15/37 (40%), Positives = 23/37 (61%)
Frame = +3
Query: 465 RDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRN 501
R++++KRKEK +KEK + + KE+ KK S N
Sbjct: 783 REEKEKRKEKEKKEKEREREKEKSKERHKKDESDSDN 893
>TC79306 similar to PIR|F86397|F86397 protein T7N9.16 [imported] -
Arabidopsis thaliana, partial (2%)
Length = 826
Score = 30.4 bits (67), Expect = 1.8
Identities = 12/27 (44%), Positives = 18/27 (66%)
Frame = -2
Query: 462 SEVRDDQKKRKEKYEKEKRKIKRKERE 488
S DD+K+RK E++ RK++R RE
Sbjct: 627 SRYEDDRKERKNSREEDDRKVRRNSRE 547
>TC83069
Length = 894
Score = 30.4 bits (67), Expect = 1.8
Identities = 21/62 (33%), Positives = 36/62 (57%)
Frame = +3
Query: 454 ISLAPLSPSEVRDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQTAPIFTTL 513
IS P SPS++ D + + K K + + IK++ +EF KKKK + R ++ IFT+
Sbjct: 528 ISTTP-SPSDILDKEDQIKNK*SSDLQ-IKKE*KEFRLKKKKPKDKRK*KQKIIIIFTSA 701
Query: 514 QK 515
++
Sbjct: 702 KE 707
>TC82397 similar to GP|18377877|gb|AAL67124.1 AT3g53630/F4P12_330
{Arabidopsis thaliana}, partial (9%)
Length = 716
Score = 30.0 bits (66), Expect = 2.4
Identities = 17/52 (32%), Positives = 29/52 (55%)
Frame = +3
Query: 439 DGRTNKYSFMHSGQKISLAPLSPSEVRDDQKKRKEKYEKEKRKIKRKEREFA 490
DG ++ G K+ L V+ +++KR+ ++ KEK + KRKE EF+
Sbjct: 459 DGDVSEEELKRLGVKV--LKLREVMVKREERKREREFLKEKEEWKRKEFEFS 608
>AW560664 similar to GP|23495077|gb hypothetical protein {Plasmodium
falciparum 3D7}, partial (0%)
Length = 545
Score = 30.0 bits (66), Expect = 2.4
Identities = 17/48 (35%), Positives = 28/48 (57%)
Frame = -3
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQTAPIFTTLQK 515
+K K K EK+++ K+KE+ KKKK R + + + P TTL++
Sbjct: 393 KKVTKNKGEKKEKSKKKKEK----KKKKERKEKKTSKAALPRGTTLRE 262
>TC79030 similar to PIR|H86366|H86366 protein F26F24.12 [imported] -
Arabidopsis thaliana, partial (57%)
Length = 1598
Score = 29.6 bits (65), Expect = 3.1
Identities = 17/49 (34%), Positives = 22/49 (44%)
Frame = +1
Query: 288 DDLGE*NVEVENHEGYVEEEAEEIPSGDLFMIRRFLGNQAKEEESNQRE 336
D+ E EVE EGY EE +++ F I GN A E + E
Sbjct: 829 DEEDEEEAEVEYVEGYELEEEDDMEDFGAFAIHESRGNDADHENAGSSE 975
>BG645417 weakly similar to GP|14586370|em putative protein {Arabidopsis
thaliana}, partial (3%)
Length = 805
Score = 29.3 bits (64), Expect = 4.1
Identities = 21/67 (31%), Positives = 30/67 (44%)
Frame = +3
Query: 324 GNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKPHPKPYK 383
G + E S R + LVQGK+C+ II G + V K +ET+ P+ +
Sbjct: 570 GKGQETEASAFRNANICDKKLVQGKICYGIIFGSQKHLPIR---VMKSAVETEQGPERRE 740
Query: 384 LQWLNEN 390
W EN
Sbjct: 741 KYWFFEN 761
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.358 0.160 0.569
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,023,677
Number of Sequences: 36976
Number of extensions: 214799
Number of successful extensions: 2519
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 1442
Number of HSP's successfully gapped in prelim test: 119
Number of HSP's that attempted gapping in prelim test: 973
Number of HSP's gapped (non-prelim): 1658
length of query: 550
length of database: 9,014,727
effective HSP length: 101
effective length of query: 449
effective length of database: 5,280,151
effective search space: 2370787799
effective search space used: 2370787799
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147496.2


