

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
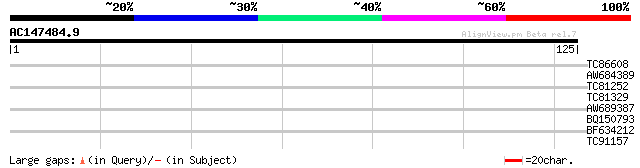
Query= AC147484.9 - phase: 0
(125 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC86608 homologue to SP|O22860|RL38_ARATH 60S ribosomal protein ... 27 2.1
AW684389 similar to GP|9759091|dbj contains similarity to unknow... 26 3.5
TC81252 similar to PIR|T52624|T52624 b-Zip DNA binding protein [... 26 4.6
TC81329 homologue to GP|20466796|gb|AAM20715.1 unknown protein {... 26 4.6
AW689387 similar to GP|10727986|gb CG18331 gene product {Drosoph... 25 6.0
BQ150793 25 6.0
BF634212 weakly similar to GP|10798748|dbj elicitor inducible ge... 25 7.9
TC91157 weakly similar to GP|16604308|gb|AAL24160.1 At1g31300/T1... 25 7.9
>TC86608 homologue to SP|O22860|RL38_ARATH 60S ribosomal protein L38.
[Mouse-ear cress] {Arabidopsis thaliana}, complete
Length = 545
Score = 26.9 bits (58), Expect = 2.1
Identities = 12/42 (28%), Positives = 21/42 (49%)
Frame = +3
Query: 81 LFQAQVIKNESNPKLVDVVIWYCGWIVTVKEKIYVLRVIFVS 122
LF +K E NP+ D+ + +C W Y+ ++F+S
Sbjct: 282 LFCFYFLKLERNPRFCDLELCFCNW--------YLFMIVFLS 383
>AW684389 similar to GP|9759091|dbj contains similarity to unknown
protein~gb|AAF47871.1~gene_id:MJJ3.6 {Arabidopsis
thaliana}, partial (16%)
Length = 656
Score = 26.2 bits (56), Expect = 3.5
Identities = 10/24 (41%), Positives = 15/24 (61%)
Frame = -2
Query: 85 QVIKNESNPKLVDVVIWYCGWIVT 108
++ +N+SN KL+ WY WI T
Sbjct: 247 RIARNKSNSKLI---FWYIAWIAT 185
>TC81252 similar to PIR|T52624|T52624 b-Zip DNA binding protein [imported] -
Arabidopsis thaliana, partial (48%)
Length = 1221
Score = 25.8 bits (55), Expect = 4.6
Identities = 10/22 (45%), Positives = 15/22 (67%)
Frame = -3
Query: 95 LVDVVIWYCGWIVTVKEKIYVL 116
L+ ++IW+ W VT K+YVL
Sbjct: 1045 LLQLIIWHVRWGVTKLGKLYVL 980
>TC81329 homologue to GP|20466796|gb|AAM20715.1 unknown protein {Arabidopsis
thaliana}, partial (20%)
Length = 963
Score = 25.8 bits (55), Expect = 4.6
Identities = 12/31 (38%), Positives = 17/31 (54%)
Frame = -1
Query: 60 HVFSVDFPGKYQDWAATDPLDLFQAQVIKNE 90
HVF GKY+ W T+ LF+ + I N+
Sbjct: 180 HVFLKRLTGKYRMWCVTEA--LFEGKTISNK 94
>AW689387 similar to GP|10727986|gb CG18331 gene product {Drosophila
melanogaster}, partial (1%)
Length = 613
Score = 25.4 bits (54), Expect = 6.0
Identities = 12/30 (40%), Positives = 18/30 (60%)
Frame = -3
Query: 5 PKVLMVAEKPSIALSIATVLSHGKMNTRRG 34
P VL ++ I LSI ++LS + TR+G
Sbjct: 290 PTVLSISSAIIIPLSITSILSKDQSETRKG 201
>BQ150793
Length = 390
Score = 25.4 bits (54), Expect = 6.0
Identities = 11/25 (44%), Positives = 16/25 (64%)
Frame = +3
Query: 101 WYCGWIVTVKEKIYVLRVIFVSLSF 125
WYC V + E I +RVI++ LS+
Sbjct: 27 WYCHTCVMMGEVIRSIRVIWIPLSW 101
>BF634212 weakly similar to GP|10798748|dbj elicitor inducible gene product
EIG-I24 {Nicotiana tabacum}, partial (14%)
Length = 385
Score = 25.0 bits (53), Expect = 7.9
Identities = 10/21 (47%), Positives = 13/21 (61%)
Frame = -1
Query: 20 IATVLSHGKMNTRRGSTEVHE 40
+ TV +GKM RRGS + E
Sbjct: 361 VVTVQQNGKMGRRRGSLRIRE 299
>TC91157 weakly similar to GP|16604308|gb|AAL24160.1 At1g31300/T19E23_12
{Arabidopsis thaliana}, partial (50%)
Length = 763
Score = 25.0 bits (53), Expect = 7.9
Identities = 12/34 (35%), Positives = 17/34 (49%)
Frame = -1
Query: 90 ESNPKLVDVVIWYCGWIVTVKEKIYVLRVIFVSL 123
E N K D+V+ WIV + + +R I V L
Sbjct: 502 EQNTKYKDIVVIKMEWIVDIARLFHSIRWILVRL 401
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.137 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,560,728
Number of Sequences: 36976
Number of extensions: 39616
Number of successful extensions: 213
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 212
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 213
length of query: 125
length of database: 9,014,727
effective HSP length: 84
effective length of query: 41
effective length of database: 5,908,743
effective search space: 242258463
effective search space used: 242258463
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 52 (24.6 bits)
Medicago: description of AC147484.9


