

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147484.3 - phase: 0 /pseudo
(526 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
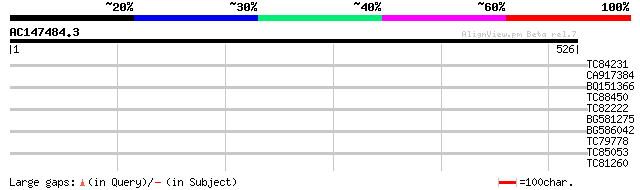
Score E
Sequences producing significant alignments: (bits) Value
TC84231 29 3.9
CA917384 similar to PIR|T06311|T06 hypothetical protein F11C18.9... 29 5.1
BQ151366 similar to GP|20146300|dbj hypothetical protein~similar... 29 5.1
TC88450 similar to PIR|F86427|F86427 auxin response factor 6 (AR... 28 6.6
TC82222 similar to GP|17933299|gb|AAL48232.1 AT4g17800/dl4935c {... 28 6.6
BG581275 28 8.6
BG586042 28 8.6
TC79778 weakly similar to PIR|T39903|T39903 serine-rich protein ... 28 8.6
TC85053 similar to PIR|E85062|E85062 hypothetical protein AT4g04... 28 8.6
TC81260 similar to GP|21536901|gb|AAM61233.1 unknown {Arabidopsi... 28 8.6
>TC84231
Length = 815
Score = 29.3 bits (64), Expect = 3.9
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = +1
Query: 18 CSQGHRSSLNLQTNQGGTICLVCFSNLISNPS 49
CS H+ L L+ G +C+ C SN +NPS
Sbjct: 682 CSYFHQLKL*LRDKGGLWLCITCISNTETNPS 777
>CA917384 similar to PIR|T06311|T06 hypothetical protein F11C18.90 -
Arabidopsis thaliana, partial (41%)
Length = 741
Score = 28.9 bits (63), Expect = 5.1
Identities = 31/158 (19%), Positives = 66/158 (41%)
Frame = +2
Query: 298 SVSQVQIGALDLLFHYLSSVGTSGNQIQVLVEENIADYLFEILRLSVPFHPVQYETLKLI 357
S SQV+ AL L++ + + I ++E ++ +L S+ V L ++
Sbjct: 203 SSSQVKQDALRALYN----LSINQTNISFVLETDLVVFLIN----SIEDMEVSERVLSIL 358
Query: 358 YECISECPGAVSTSQMEELVLVLIKMLTKHSDGEMGMIPETFIMACSVFVALMRSPSCNG 417
+S G + S +++ + VL+ +L PE A + + + +
Sbjct: 359 SNLVSSPEGRKAISAVKDAITVLVDVLNWTDS------PECQEKASYILMIMAHKAYADR 520
Query: 418 ALDLSKSIEEAVKQAILACLYVSERNINQILQCFYLLK 455
+ I ++ + L ++++ ++ILQCF L K
Sbjct: 521 QAMIEAGIVSSLLELTLVGTALAQKRASRILQCFRLDK 634
>BQ151366 similar to GP|20146300|dbj hypothetical protein~similar to Oryza
sativa OSJNBa0058E19.20, partial (9%)
Length = 1137
Score = 28.9 bits (63), Expect = 5.1
Identities = 12/36 (33%), Positives = 21/36 (58%)
Frame = -1
Query: 37 CLVCFSNLISNPSSPTIHVSYALSQLSRSLSIPQFL 72
C VCF+ L+ +PSS S + + + S S+ ++L
Sbjct: 789 CAVCFARLLGSPSSRAFSSSLLVRRFAGSFSVQRWL 682
>TC88450 similar to PIR|F86427|F86427 auxin response factor 6 (ARF6)
[imported] - Arabidopsis thaliana, partial (17%)
Length = 1422
Score = 28.5 bits (62), Expect = 6.6
Identities = 13/54 (24%), Positives = 26/54 (48%)
Frame = +1
Query: 101 EQLVHLILNLSASSDPSVCREFVARVSDRVSSGALGWSSRQLHSLHCLGVLLNC 154
EQ + + + P+ + + + + ++ L WSS H + CL +LL+C
Sbjct: 376 EQALGFLFKIQLVGHPNALQWIHSSLREHLNVFCLRWSS*GRHGIRCLKMLLHC 537
>TC82222 similar to GP|17933299|gb|AAL48232.1 AT4g17800/dl4935c {Arabidopsis
thaliana}, partial (8%)
Length = 1062
Score = 28.5 bits (62), Expect = 6.6
Identities = 23/90 (25%), Positives = 45/90 (49%), Gaps = 2/90 (2%)
Frame = -2
Query: 389 DGEMGMIPETFIMACSVFVALMRSPSCNGALDLSKSIEEAVKQAILACLYVSERN--INQ 446
D +GMI ++A + + L R+ +CN L + S + + A+ +R+ + Q
Sbjct: 704 DN*LGMISN--LLAFLLLLRLWRTKTCNHNLHRTCSYQRTSQWALNNRATYGQRDSQLPQ 531
Query: 447 ILQCFYLLKEAYAYSHDENSTHISKLELRS 476
IL YL++ + + D+ HISK + ++
Sbjct: 530 ILIHLYLVQTSQSRKVDQKFPHISKSQYQN 441
>BG581275
Length = 484
Score = 28.1 bits (61), Expect = 8.6
Identities = 28/92 (30%), Positives = 36/92 (38%)
Frame = +3
Query: 27 NLQTNQGGTICLVCFSNLISNPSSPTIHVSYALSQLSRSLSIPQFLHSIVTFHPHFLVSP 86
NL N T N S SSP++HV P FLH V F P L P
Sbjct: 261 NLSFNNSTTTLATASLNPSSATSSPSMHV-------------PVFLH--VPFKPSSLSLP 395
Query: 87 LVAALSSFDDEPIAEQLVHLILNLSASSDPSV 118
A L S P + ++ +L+A PS+
Sbjct: 396 FAAPLLSNHSTPSS-----MLSSLAADFKPSL 476
>BG586042
Length = 237
Score = 28.1 bits (61), Expect = 8.6
Identities = 17/39 (43%), Positives = 23/39 (58%)
Frame = +2
Query: 190 LFVLYKLFALHSTSDEGDGSDMLIPFCPKLLYLLGDVLL 228
LF+L+KLFALH S SDM++ F +L+ LL
Sbjct: 50 LFMLFKLFALHRCSLH---SDMMLRFFCHILFYEARTLL 157
>TC79778 weakly similar to PIR|T39903|T39903 serine-rich protein - fission
yeast (Schizosaccharomyces pombe), partial (7%)
Length = 789
Score = 28.1 bits (61), Expect = 8.6
Identities = 17/42 (40%), Positives = 24/42 (56%), Gaps = 1/42 (2%)
Frame = -1
Query: 24 SSLNLQTNQGGTICLVCFSNLISN-PSSPTIHVSYALSQLSR 64
S L LQT ++C++ F L S+ PSS +I +S S SR
Sbjct: 720 SKLTLQTLSTSSLCIILFITLNSS*PSSSSISLSSTSSSSSR 595
>TC85053 similar to PIR|E85062|E85062 hypothetical protein AT4g04970
[imported] - Arabidopsis thaliana, partial (14%)
Length = 769
Score = 28.1 bits (61), Expect = 8.6
Identities = 14/42 (33%), Positives = 20/42 (47%)
Frame = -1
Query: 485 HLLPWLATGIMNGEEELFLGLLEIFHSILLLHCSIILGNSQR 526
HL PW T I + + LFL L H + H ++ N Q+
Sbjct: 163 HLRPWPQTWISDSSQHLFLYLPVYIHDVSHCHLAMYN*NQQK 38
>TC81260 similar to GP|21536901|gb|AAM61233.1 unknown {Arabidopsis
thaliana}, partial (34%)
Length = 562
Score = 28.1 bits (61), Expect = 8.6
Identities = 13/46 (28%), Positives = 25/46 (54%)
Frame = -3
Query: 326 VLVEENIADYLFEILRLSVPFHPVQYETLKLIYECISECPGAVSTS 371
+L+ N + L E+L ++ +QY T L+Y + C ++ST+
Sbjct: 437 LLISSNQLENLIELLSINYICFYIQYITYVLVYVS*TNCAASISTT 300
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.137 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,895,606
Number of Sequences: 36976
Number of extensions: 276859
Number of successful extensions: 1557
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1545
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1557
length of query: 526
length of database: 9,014,727
effective HSP length: 101
effective length of query: 425
effective length of database: 5,280,151
effective search space: 2244064175
effective search space used: 2244064175
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147484.3


