

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147483.13 - phase: 0
(208 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
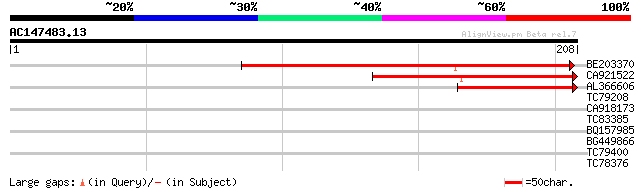
BE203370 121 2e-28
CA921522 87 6e-18
AL366606 69 1e-12
TC79208 similar to PIR|T02652|T02652 probable choline kinase At2... 30 0.50
CA918173 weakly similar to GP|11034537|dbj putative glucosyl tra... 30 0.86
TC83385 27 4.3
BQ157985 27 5.6
BG449866 similar to GP|7298830|gb|A CG1152-PA {Drosophila melano... 26 9.5
TC79400 weakly similar to GP|20161447|dbj|BAB90371. hypothetical... 26 9.5
TC78376 similar to SP|Q9SK33|WR60_ARATH Probable WRKY transcript... 26 9.5
>BE203370
Length = 454
Score = 121 bits (303), Expect = 2e-28
Identities = 63/128 (49%), Positives = 88/128 (68%), Gaps = 6/128 (4%)
Frame = +1
Query: 86 CFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTGPGTSLTPLVVAKDLYLKTSDVSNHLTTK 145
CFTFPD+Q+ PTLE YS+L+ P +K PFTG +A L+L+TS + LT K
Sbjct: 4 CFTFPDYQLAPTLEEYSYLVG*PVLDKVPFTGFEPIPKAATIADALHLETSLIKAKLTIK 183
Query: 146 SHIRGFTSKYLLEQATC------QDALEAILALLIYGLILFPSLDNFVDMNAIKIFHSKN 199
++ +K+L +QA+ DA +ILALLIYGL+LFP++DN+VD++AI++F +KN
Sbjct: 184 GNLPSLPTKFLYQQASDFAKVDNVDAFYSILALLIYGLVLFPNVDNYVDIHAIQMFLTKN 363
Query: 200 PVPTLLAD 207
PVPTLLAD
Sbjct: 364 PVPTLLAD 387
>CA921522
Length = 767
Score = 86.7 bits (213), Expect = 6e-18
Identities = 44/81 (54%), Positives = 57/81 (70%), Gaps = 6/81 (7%)
Frame = +3
Query: 134 KTSDVSNHLTTKSHIRGFTSKYLLEQATCQD------ALEAILALLIYGLILFPSLDNFV 187
+T +V HL T+ GF++ +L E+ T D A +ILALL+YGL+LFPSLDNFV
Sbjct: 3 ETDEVKTHLITRGKFLGFSTDFLYERTTFFDKMGVAYAFNSILALLVYGLVLFPSLDNFV 182
Query: 188 DMNAIKIFHSKNPVPTLLADT 208
D+ AI+IF S+NPVPTLL DT
Sbjct: 183 DIKAIQIFLSRNPVPTLLGDT 245
>AL366606
Length = 393
Score = 69.3 bits (168), Expect = 1e-12
Identities = 34/44 (77%), Positives = 37/44 (83%)
Frame = -1
Query: 165 ALEAILALLIYGLILFPSLDNFVDMNAIKIFHSKNPVPTLLADT 208
A AI L GLILFP+LDNFVDMNAI++FHSKNPVPTLLADT
Sbjct: 390 APSAIXLCLSTGLILFPNLDNFVDMNAIEVFHSKNPVPTLLADT 259
>TC79208 similar to PIR|T02652|T02652 probable choline kinase At2g26830
[imported] - Arabidopsis thaliana, partial (74%)
Length = 1565
Score = 30.4 bits (67), Expect = 0.50
Identities = 21/75 (28%), Positives = 39/75 (52%), Gaps = 3/75 (4%)
Frame = +3
Query: 122 LTPLVV--AKDLYLKTSDVSNHLTTKSHIRGFTSKYLLEQATCQ-DALEAILALLIYGLI 178
+TPLV+ KD++ S + + I G + LL+ + Q D+++ I+ + +YG
Sbjct: 231 MTPLVINLCKDMFKTWSSLGDSCFKVEKISGGITNLLLKVSVKQGDSIDDIITVRLYGPN 410
Query: 179 LFPSLDNFVDMNAIK 193
+D F ++ AIK
Sbjct: 411 TEHIIDRFRELQAIK 455
>CA918173 weakly similar to GP|11034537|dbj putative glucosyl transferase
{Oryza sativa (japonica cultivar-group)}, partial (19%)
Length = 769
Score = 29.6 bits (65), Expect = 0.86
Identities = 13/22 (59%), Positives = 18/22 (81%)
Frame = -3
Query: 72 LVHTLVQFYDPSFRCFTFPDFQ 93
L+HTLVQ + PS RCF FP+++
Sbjct: 113 LIHTLVQKW-PS*RCFPFPNYE 51
>TC83385
Length = 440
Score = 27.3 bits (59), Expect = 4.3
Identities = 18/57 (31%), Positives = 31/57 (53%), Gaps = 5/57 (8%)
Frame = +3
Query: 92 FQVVPTLEAYSHLLDSPTAEKTPFTGPGTSL-----TPLVVAKDLYLKTSDVSNHLT 143
F V+ L +YS+L+ P + F+ P S+ TPL K+++L T + NH++
Sbjct: 120 FIVLFCLSSYSYLI--PFFQFQTFSCPSLSIK*ISSTPLTKKKNIFLPTXN*HNHIS 284
>BQ157985
Length = 908
Score = 26.9 bits (58), Expect = 5.6
Identities = 13/31 (41%), Positives = 20/31 (63%)
Frame = +1
Query: 164 DALEAILALLIYGLILFPSLDNFVDMNAIKI 194
DAL I +IY +ILF +++ ++D N I I
Sbjct: 766 DALFRISIYIIYCIILFRAINRYIDYNYIFI 858
>BG449866 similar to GP|7298830|gb|A CG1152-PA {Drosophila melanogaster},
partial (14%)
Length = 648
Score = 26.2 bits (56), Expect = 9.5
Identities = 20/63 (31%), Positives = 29/63 (45%)
Frame = -1
Query: 82 PSFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTGPGTSLTPLVVAKDLYLKTSDVSNH 141
PS + P +V H +D+ A + TGPGTSL+ V + +L+ S V
Sbjct: 567 PSVVTISCPFVEVSFVSSDVEHEVDTAAATECLTTGPGTSLS-CVTSASTFLRLSLVGPI 391
Query: 142 LTT 144
L T
Sbjct: 390 LRT 382
>TC79400 weakly similar to GP|20161447|dbj|BAB90371. hypothetical
protein~similar to Arabidopsis thaliana chromosome 1
F8K7.2, partial (12%)
Length = 1251
Score = 26.2 bits (56), Expect = 9.5
Identities = 20/52 (38%), Positives = 25/52 (47%), Gaps = 4/52 (7%)
Frame = -3
Query: 87 FTFPDFQVVPTLEAYS--HLLDSPTAEK--TPFTGPGTSLTPLVVAKDLYLK 134
F F F P+ S LL+ P A TPF G GT LVV DL+++
Sbjct: 739 FLFAFFVKTPSATRTSSMELLELPFAFSCFTPFVGLGTRTAVLVVDTDLFVE 584
>TC78376 similar to SP|Q9SK33|WR60_ARATH Probable WRKY transcription factor
60 (WRKY DNA-binding protein 60). [Mouse-ear cress],
partial (38%)
Length = 1121
Score = 26.2 bits (56), Expect = 9.5
Identities = 12/30 (40%), Positives = 16/30 (53%)
Frame = -2
Query: 95 VPTLEAYSHLLDSPTAEKTPFTGPGTSLTP 124
V L +YSH LD+PT + P P + P
Sbjct: 322 VSRLASYSHCLDAPTLQPKPDQQPQQLVCP 233
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.139 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,224,094
Number of Sequences: 36976
Number of extensions: 79029
Number of successful extensions: 424
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 423
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 423
length of query: 208
length of database: 9,014,727
effective HSP length: 92
effective length of query: 116
effective length of database: 5,612,935
effective search space: 651100460
effective search space used: 651100460
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC147483.13


