

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147482.6 + phase: 2 /pseudo
(675 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
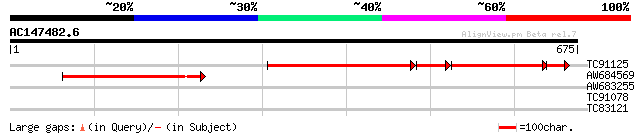
TC91125 similar to PIR|T47659|T47659 spliceosomal-like protein -... 341 0.0
AW684569 similar to PIR|T47659|T4 spliceosomal-like protein - Ar... 187 1e-47
AW683255 similar to PIR|T47659|T47 spliceosomal-like protein - A... 41 0.002
TC91078 homologue to SP|P02828|HS83_DROME Heat shock protein 83 ... 29 6.7
TC83121 similar to PIR|G84487|G84487 probable DNA replication li... 28 8.8
>TC91125 similar to PIR|T47659|T47659 spliceosomal-like protein -
Arabidopsis thaliana, partial (31%)
Length = 1157
Score = 341 bits (874), Expect(4) = 0.0
Identities = 174/177 (98%), Positives = 176/177 (99%)
Frame = +3
Query: 307 RPNGEWRGG*G*R*PPL**ALWLSKS*IR*VGILHQNS*SKNRKYNLSFGASG**GCIQR 366
RPNGEWRGG*G*R*PPL**ALWLSKS*IR*VGILHQNS*SKNRKYNLSFGASG**GCIQR
Sbjct: 33 RPNGEWRGG*G*R*PPL**ALWLSKS*IR*VGILHQNS*SKNRKYNLSFGASG**GCIQR 212
Query: 367 MHSKFS**RIRNSFRRWHSKGTAIYAQKEFNCWIYSYI*VSRGWKVS*TTSQNSG*RCSS 426
MHSKFS**RIRNSF RWHSKGTAIYAQKEFNCWIYSYI*VSRGWK+S*TTSQNSG*RCSS
Sbjct: 213 MHSKFS**RIRNSFSRWHSKGTAIYAQKEFNCWIYSYI*VSRGWKIS*TTSQNSG*RCSS 392
Query: 427 CS*SVSGKITCRNRTCSQVL*FGKKTVIKEV*E*AIPQHNCLYSNLP*SNLCWGHSR 483
CS*SVSGKITCRNRTCSQV+*FGKKTVIKEV*E*AIPQHNCLYSNLP*SNLCWGHSR
Sbjct: 393 CS*SVSGKITCRNRTCSQVI*FGKKTVIKEV*E*AIPQHNCLYSNLP*SNLCWGHSR 563
Score = 215 bits (548), Expect(4) = 0.0
Identities = 109/115 (94%), Positives = 110/115 (94%)
Frame = +2
Query: 526 CHRTCLMR*KKILLVGGSSGSRVS*MELPIRWKKLCNFMLVM*SHACKRPLLYQVVESAF 585
C RTCLMR*KKILLVGGSSGSR S*MELPIR KKLCNFMLVM*SHACKRPLLYQVVESAF
Sbjct: 716 CRRTCLMR*KKILLVGGSSGSRGS*MELPIR*KKLCNFMLVM*SHACKRPLLYQVVESAF 895
Query: 586 *MEQLWEVLEHYMLLLHEMMLISSLIWRCI*GRIIHLCVEEIIWLIDPPIFLLSL 640
MEQLWEVLEHYMLLLHEMMLISSLIWRCI*GRIIHLCVEEIIWLIDPPIFLL +
Sbjct: 896 YMEQLWEVLEHYMLLLHEMMLISSLIWRCI*GRIIHLCVEEIIWLIDPPIFLLRM 1060
Score = 89.0 bits (219), Expect(4) = 0.0
Identities = 39/40 (97%), Positives = 39/40 (97%)
Frame = +1
Query: 485 CKYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG 524
CKYR DENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG
Sbjct: 577 CKYRRDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG 696
Score = 56.2 bits (134), Expect(4) = 0.0
Identities = 26/27 (96%), Positives = 26/27 (96%)
Frame = +1
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEI 666
LCEQFPTLPMDLQRKIADELDRT GEI
Sbjct: 1075 LCEQFPTLPMDLQRKIADELDRTPGEI 1155
>AW684569 similar to PIR|T47659|T4 spliceosomal-like protein - Arabidopsis
thaliana, partial (10%)
Length = 522
Score = 187 bits (474), Expect = 1e-47
Identities = 109/173 (63%), Positives = 115/173 (66%), Gaps = 2/173 (1%)
Frame = +3
Query: 63 SMKCLAMLLVWTLLQCPKADKDPVFLQLDHMTRQSESYHWILMIVCRL*VYRVFLQLQNR 122
SMKCLAMLLVWTLLQCPK DKDPVFLQLDHMTRQSESYHWILMIVCRL*VYRVFLQLQNR
Sbjct: 3 SMKCLAMLLVWTLLQCPKEDKDPVFLQLDHMTRQSESYHWILMIVCRL*VYRVFLQLQNR 182
Query: 123 FFSLKFKHLLAVRMGQITPLAFS*RWFAEWCTI*NCGGYGHRSAF*FSIPILRIETPKAV 182
FFSLKFKHLLAVRMGQITPLAFS* + F P P++
Sbjct: 183 FFSLKFKHLLAVRMGQITPLAFS*TLVCRMVYYLELWWIWSQVCFLILDPDS*D*DPQSC 362
Query: 183 SHCCKREA--CYALPVKSPLAWLHSPRTLPFNTPFV*NP*ICCLIFI*PMCGR 233
+ C L L ++H L PFV NP*ICCLI I*PMCGR
Sbjct: 363 FPLL*EGSVLCSCLSSXPWLGYIHQGHFL-LTPPFVXNP*ICCLIXI*PMCGR 518
>AW683255 similar to PIR|T47659|T47 spliceosomal-like protein - Arabidopsis
thaliana, partial (3%)
Length = 145
Score = 40.8 bits (94), Expect = 0.002
Identities = 27/48 (56%), Positives = 27/48 (56%), Gaps = 7/48 (14%)
Frame = +1
Query: 275 ETFGCD*E*SRSIYCRRA*SCKESVLRLLKLG-------RPNGEWRGG 315
ET G D*E*SRS Y RRA*SCKE V R NGEW GG
Sbjct: 1 ETSGGD*E*SRSTYSRRA*SCKERVF*GCSCWGK*DWK*RSNGEWWGG 144
>TC91078 homologue to SP|P02828|HS83_DROME Heat shock protein 83 (HSP 82).
[Fruit fly] {Drosophila melanogaster}, partial (2%)
Length = 682
Score = 28.9 bits (63), Expect = 6.7
Identities = 17/50 (34%), Positives = 26/50 (52%)
Frame = +3
Query: 589 QLWEVLEHYMLLLHEMMLISSLIWRCI*GRIIHLCVEEIIWLIDPPIFLL 638
+L EVL + L + +++ S +I RCI R L V L P+F+L
Sbjct: 414 ELMEVLSRFQLSIAKLVQPSDIIKRCILSRKPTLAVRYASLLPQAPVFIL 563
>TC83121 similar to PIR|G84487|G84487 probable DNA replication licensing
factor mcm5 [imported] - Arabidopsis thaliana, partial
(12%)
Length = 499
Score = 28.5 bits (62), Expect = 8.8
Identities = 22/67 (32%), Positives = 32/67 (46%)
Frame = -3
Query: 71 LVWTLLQCPKADKDPVFLQLDHMTRQSESYHWILMIVCRL*VYRVFLQLQNRFFSLKFKH 130
L WTLL+ A + +FLQ H + + + ++C Q+RFFSL +
Sbjct: 251 LAWTLLRLSGA*LEGLFLQ*IHSSPFCRDHQSVFFLIC---------GSQSRFFSLS-EL 102
Query: 131 LLAVRMG 137
LLAV G
Sbjct: 101 LLAVSHG 81
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.360 0.160 0.619
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,732,260
Number of Sequences: 36976
Number of extensions: 473532
Number of successful extensions: 4979
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1957
Number of HSP's successfully gapped in prelim test: 224
Number of HSP's that attempted gapping in prelim test: 2943
Number of HSP's gapped (non-prelim): 2353
length of query: 675
length of database: 9,014,727
effective HSP length: 103
effective length of query: 572
effective length of database: 5,206,199
effective search space: 2977945828
effective search space used: 2977945828
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC147482.6


