

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147435.1 - phase: 0 /pseudo
(346 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
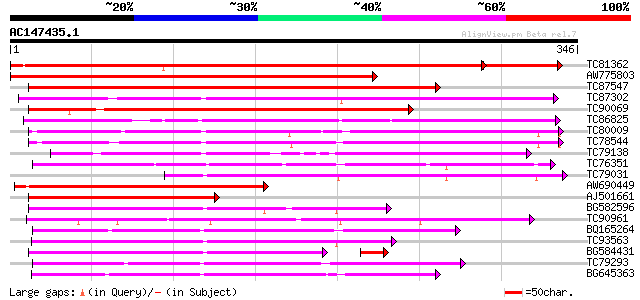
Score E
Sequences producing significant alignments: (bits) Value
TC81362 similar to GP|19911201|dbj|BAB86927. glucosyltransferase... 462 e-147
AW775803 similar to GP|19911201|db glucosyltransferase-9 {Vigna ... 449 e-127
TC87547 weakly similar to PIR|F84784|F84784 probable glucosyl tr... 267 4e-72
TC87302 similar to GP|19911199|dbj|BAB86926. glucosyltransferase... 255 2e-68
TC90069 similar to GP|19911211|dbj|BAB86932. glucosyltransferase... 240 5e-64
TC86825 similar to GP|18151384|dbj|BAB83692. ABA-glucosyltransfe... 201 4e-52
TC80009 weakly similar to PIR|T03747|T03747 glucosyltransferase ... 197 6e-51
TC78544 weakly similar to PIR|T03747|T03747 glucosyltransferase ... 188 3e-48
TC79138 weakly similar to GP|4115534|dbj|BAA36410.1 UDP-glycose:... 186 1e-47
TC76351 similar to GP|7635494|emb|CAB88666.1 putative UDP-glycos... 175 3e-44
TC79031 weakly similar to GP|13492674|gb|AAK28303.1 phenylpropan... 152 2e-37
AW690449 weakly similar to PIR|T07786|T077 UDP-glucose glucosylt... 149 1e-36
AJ501661 weakly similar to PIR|F84784|F847 probable glucosyl tra... 148 3e-36
BG582596 weakly similar to GP|18151384|dbj ABA-glucosyltransfera... 146 1e-35
TC90961 similar to GP|20067056|gb|AAM09517.1 putative glucosyltr... 144 7e-35
BQ165264 weakly similar to PIR|T07786|T077 UDP-glucose glucosylt... 135 3e-32
TC93563 weakly similar to PIR|T07786|T07786 UDP-glucose glucosyl... 130 7e-31
BG584431 weakly similar to GP|19911201|dbj glucosyltransferase-9... 120 5e-29
TC79293 weakly similar to GP|15290056|dbj|BAB63750. putative glu... 122 2e-28
BG645363 weakly similar to GP|15290080|dbj putative salicylate-i... 117 5e-27
>TC81362 similar to GP|19911201|dbj|BAB86927. glucosyltransferase-9 {Vigna
angularis}, partial (70%)
Length = 1347
Score = 462 bits (1189), Expect(2) = e-147
Identities = 231/295 (78%), Positives = 258/295 (87%), Gaps = 4/295 (1%)
Frame = +1
Query: 1 MVLPANINDVPHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRA 60
MVL ANIN VPHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRA
Sbjct: 76 MVLQANIN-VPHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRA 252
Query: 61 VSSGLQIKIVTLNFPTKQAGLPEGCENFDMVD---SIDMRMNLFHAITLLQKPAEELFDA 117
VSSGLQIKIVTLNFP+KQ GLP+GCENFDMV+ ++M+ NLFHA++LLQK E+LFD
Sbjct: 253 VSSGLQIKIVTLNFPSKQVGLPDGCENFDMVNISKDMNMKYNLFHAVSLLQKEGEDLFDK 432
Query: 118 LTPKPSCIISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEY 177
L+PKPSCIISDF I WT QIAE H IPRISFHGF CF LHCM K+ TS ILE ++SE+E+
Sbjct: 433 LSPKPSCIISDFCITWTSQIAEKHHIPRISFHGFCCFTLHCMFKVHTSNILESINSETEF 612
Query: 178 FTVPGIPDQIQVTKEQIPGILKGE-LKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYK 236
F++PGIPD+IQVTKEQIPG +K E +K F EKM +AEMKSYG IIN+FEELEK YV DYK
Sbjct: 613 FSIPGIPDKIQVTKEQIPGTVKEEKMKGFAEKMQEAEMKSYGVIINSFEELEKEYVNDYK 792
Query: 237 KEKNGKVWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLG 291
K +N KVW VGPV+LCNKDGLDKAQRG IASISEH+CL +LDLH+PKSVVY CLG
Sbjct: 793 KVRNDKVWCVGPVALCNKDGLDKAQRGNIASISEHNCLNFLDLHKPKSVVYVCLG 957
Score = 76.3 bits (186), Expect(2) = e-147
Identities = 37/48 (77%), Positives = 41/48 (85%)
Frame = +2
Query: 290 LGSLCNLIPSQLMELALALEATNRPFIWVIREGNKSSEELEKWISEER 337
L SLCNLIPSQL+ELAL L+AT PFIWVIREG SEELEKWIS+E+
Sbjct: 953 LESLCNLIPSQLIELALGLQATKIPFIWVIREGIYKSEELEKWISDEK 1096
>AW775803 similar to GP|19911201|db glucosyltransferase-9 {Vigna angularis},
partial (34%)
Length = 716
Score = 449 bits (1156), Expect = e-127
Identities = 223/224 (99%), Positives = 223/224 (99%)
Frame = +2
Query: 1 MVLPANINDVPHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRA 60
MVLPANINDVPHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRA
Sbjct: 44 MVLPANINDVPHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRA 223
Query: 61 VSSGLQIKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTP 120
VSSGLQIKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTP
Sbjct: 224 VSSGLQIKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTP 403
Query: 121 KPSCIISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTV 180
KPSCIISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTV
Sbjct: 404 KPSCIISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTV 583
Query: 181 PGIPDQIQVTKEQIPGILKGELKEFGEKMHDAEMKSYGEIINTF 224
PGIPDQIQVTKEQIPGILKGEL EFGEKMHDAEMKSYGEIINTF
Sbjct: 584 PGIPDQIQVTKEQIPGILKGELXEFGEKMHDAEMKSYGEIINTF 715
>TC87547 weakly similar to PIR|F84784|F84784 probable glucosyl transferase
[imported] - Arabidopsis thaliana, partial (29%)
Length = 878
Score = 267 bits (683), Expect = 4e-72
Identities = 137/256 (53%), Positives = 182/256 (70%), Gaps = 4/256 (1%)
Frame = +2
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQ-RGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIV 70
HFVLFP++AQGH+IPM+DIAK+LAQ + VIVTI TTPKNASRFTS+++R V GL I++V
Sbjct: 80 HFVLFPMMAQGHMIPMMDIAKILAQHQNVIVTIVTTPKNASRFTSIVARCVEYGLDIQLV 259
Query: 71 TLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFF 130
L FP K++GLPEGCEN DM+ ++ M N +A+ Q+ E+LF+ T +CIISD
Sbjct: 260 QLEFPCKESGLPEGCENLDMLPALGMASNFLNALKFFQQEVEKLFEEFTTPATCIISDMC 439
Query: 131 IPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILE-RVDSESEYFTVPGIPDQIQV 189
+P+T +A IPRI+F G SCF L M + + E + ESEYF +PGIPD+I++
Sbjct: 440 LPYTSHVARKFNIPRITFLGVSCFHLFNMHNFHVNNMAEIMANKESEYFELPGIPDKIEM 619
Query: 190 TKEQIP-GILKGEL-KEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVG 247
T Q G LKGE+ K+F + + +AE+ SYG ++N+FEELE Y +DYKK +N KVW +G
Sbjct: 620 TIAQTGLGGLKGEVWKQFNDDLLEAEIGSYGMLVNSFEELEPTYARDYKKVRNDKVWCIG 799
Query: 248 PVSLCNKDGLDKAQRG 263
PVSL N D LDK QRG
Sbjct: 800 PVSLSNTDYLDKVQRG 847
>TC87302 similar to GP|19911199|dbj|BAB86926. glucosyltransferase-8 {Vigna
angularis}, partial (53%)
Length = 1787
Score = 255 bits (652), Expect = 2e-68
Identities = 134/333 (40%), Positives = 196/333 (58%), Gaps = 3/333 (0%)
Frame = +1
Query: 6 NINDVPHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGL 65
N N H + FP +A GHIIP +D+A++ + RG+ VTI TT N + + +A
Sbjct: 40 NENQELHIIFFPFLANGHIIPCVDLARVFSSRGLKVTIVTTHLNVPLISRTIGKA----- 204
Query: 66 QIKIVTLNFPT-KQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSC 124
+I I T+ FP+ ++ GLPEGCEN + + D + + LL++P E + + KP C
Sbjct: 205 KINIKTIKFPSPEETGLPEGCENSESALAPDKFIKFMKSTLLLREPLEHVLEQ--EKPDC 378
Query: 125 IISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIP 184
+++D F PW+ A IPRI FHG F L + + K ++V S +E F VP +P
Sbjct: 379 LVADMFFPWSTDSAAKFNIPRIVFHGLGFFPLCVLACTRQYKPQDKVSSYTEPFVVPNLP 558
Query: 185 DQIQVTKEQIPGILKGE--LKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGK 242
+I +TK Q+P + + + + E+ +++E+KS+G I NTF ELE Y Y+ E K
Sbjct: 559 GEITLTKMQLPQLPQHDKVFTKLLEESNESEVKSFGVIANTFYELEPVYADHYRNELGRK 738
Query: 243 VWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLM 302
W +GPVSLCN+D +KA RG ASI EH CLKWL +P SV+Y C GS+ +QL
Sbjct: 739 AWHLGPVSLCNRDTEEKACRGREASIDEHECLKWLQSKEPNSVIYVCFGSMTVFSDAQLK 918
Query: 303 ELALALEATNRPFIWVIREGNKSSEELEKWISE 335
E+A+ LEA+ PFIWV+R+ KS E +W+ E
Sbjct: 919 EIAMGLEASEVPFIWVVRKSAKSEGENLEWLPE 1017
>TC90069 similar to GP|19911211|dbj|BAB86932. glucosyltransferase-14 {Vigna
angularis}, partial (22%)
Length = 734
Score = 240 bits (613), Expect = 5e-64
Identities = 125/238 (52%), Positives = 162/238 (67%), Gaps = 3/238 (1%)
Frame = +3
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLA--QRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKI 69
HFVLFPL++ GH++PMID+A LA ++ +IVTI TTP NASRF S+ S QI++
Sbjct: 33 HFVLFPLMSPGHMLPMIDLATTLAHQKQNIIVTIVTTPHNASRF----SQTFSQNSQIQL 200
Query: 70 VTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFH-AITLLQKPAEELFDALTPKPSCIISD 128
+ L FP+K AG PEGCENFDM+ S+ M F A TLLQ AEE F+ LTPKPSCIISD
Sbjct: 201 LQLQFPSKDAGFPEGCENFDMLPSMSMAHTFFKVANTLLQDQAEEAFEKLTPKPSCIISD 380
Query: 129 FFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQ 188
P+T +IA IPRISF+G SCFCL K+ S ++E++ ++SEYF +P IP +I
Sbjct: 381 VGFPYTSKIATKFNIPRISFYGVSCFCLVWQQKLIVSNVMEKIATDSEYFLIPEIPHKIM 560
Query: 189 VTKEQIPGILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFV 246
+TK Q P + K+F ++M AEM SYG ++N+FEELE Y D K +NGKVW V
Sbjct: 561 ITKAQTPSSNDEDWKDFVDQMAAAEMVSYGVVVNSFEELET*YASDLKNTRNGKVWCV 734
>TC86825 similar to GP|18151384|dbj|BAB83692. ABA-glucosyltransferase {Vigna
angularis}, partial (87%)
Length = 1704
Score = 201 bits (510), Expect = 4e-52
Identities = 119/329 (36%), Positives = 178/329 (53%), Gaps = 1/329 (0%)
Frame = +3
Query: 9 DVPHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIK 68
D FP + GH IPM+D A++ A+ G TI TTP NA +F + ++R S I
Sbjct: 87 DAVKIYFFPFVGGGHQIPMVDTARVFAKHGATSTIITTPSNALQFQNSITRDQKSNHPIT 266
Query: 69 IVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISD 128
I L P EN ++ D+ DM ++L +P ++ + +P I+ D
Sbjct: 267 IQILTTP----------ENTEVTDT-DMSAGPMIDTSILLEPLKQFL--VQHRPDGIVVD 407
Query: 129 FFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQ 188
F W I + KIPRI F+G CF + + +LE + S+SE F VPG+PD I+
Sbjct: 408 MFHRWAGDIIDELKIPRIVFNGNGCFPRCVIENTRKHVVLENLSSDSEPFIVPGLPDIIE 587
Query: 189 VTKEQIPGILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGK-VWFVG 247
+T+ Q P ++ +F +++ E S G +IN+F +LE AY DY + K GK W VG
Sbjct: 588 MTRSQTPIFMRNP-SQFSDRIKQLEENSLGTLINSFYDLEPAYA-DYVRNKLGKKAWLVG 761
Query: 248 PVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALA 307
PVSLCN+ DK +RG +I E CL WL+ +P SV+Y GS+ + QL E+A
Sbjct: 762 PVSLCNRSVEDKKERGKQPTIDEQSCLNWLNSKKPNSVLYISFGSVARVPMKQLKEIAYG 941
Query: 308 LEATNRPFIWVIREGNKSSEELEKWISEE 336
LEA+++ FIWV+ + SS+ E W+ ++
Sbjct: 942 LEASDQSFIWVVGKILNSSKNEEDWVLDK 1028
>TC80009 weakly similar to PIR|T03747|T03747 glucosyltransferase IS5a (EC
2.4.1.-) salicylate-induced - common tobacco, partial
(19%)
Length = 1118
Score = 197 bits (500), Expect = 6e-51
Identities = 122/338 (36%), Positives = 179/338 (52%), Gaps = 11/338 (3%)
Frame = +3
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
HF+ P +A GH+IP+ DIA + A G VT+ TTP NA T LS A S L++ T
Sbjct: 114 HFI--PFLASGHMIPLFDIATMFASHGHQVTVITTPSNAQSLTKSLSSAASFFLRLH--T 281
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFI 131
++FP++Q LP+G E+ + LL P E + P CIISD
Sbjct: 282 VDFPSEQVDLPKGIESMSSTTDSITSWKIHRGAMLLHGPIENFMEK--DPPDCIISDSTY 455
Query: 132 PWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILE---RVDSESEYFTVPGIPDQIQ 188
PW +A +IP ++F+G S F + + + + +L DS+S F VP P +I
Sbjct: 456 PWANDLAHKLQIPNLTFNGLSLFTISLVESLIRNNLLHSDTNSDSDSSSFLVPNFPHRIT 635
Query: 189 VTKEQIPGILKGELKEFGEKMHDAEMKSYGEIINTFEELE-KAYVKDYKKEKNGKVWFVG 247
++ E+ P +L LK M + +KS IIN F EL+ + ++ Y+K KVW +G
Sbjct: 636 LS-EKPPKVLSKFLK----MMLETVLKSKALIINNFAELDGEECIQHYEKTTGRKVWHLG 800
Query: 248 PVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALA 307
P SL + +KA+RG ++ H CL WL+ + +V+Y C GS+ L QL E+A A
Sbjct: 801 PTSLIRRTIQEKAERGNEGEVNMHECLSWLNSQRVNAVLYICFGSINYLSDKQLYEMACA 980
Query: 308 LEATNRPFIWVIRE----GNKSSEELEKWIS---EERN 338
+E + PFIWV+ E ++S EE EKW+ EERN
Sbjct: 981 IETSGHPFIWVVPEKKGKEDESEEEKEKWLPKGFEERN 1094
>TC78544 weakly similar to PIR|T03747|T03747 glucosyltransferase IS5a (EC
2.4.1.-) salicylate-induced - common tobacco, partial
(26%)
Length = 1248
Score = 188 bits (477), Expect = 3e-48
Identities = 119/338 (35%), Positives = 172/338 (50%), Gaps = 11/338 (3%)
Frame = +3
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
HF+ +P A GH++P+ DIA L A RG VTI TTP NA T LS A +++ T
Sbjct: 108 HFIPYP--ASGHMMPLCDIATLFASRGQHVTIITTPSNAQSLTKTLSSAA-----LRLHT 266
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFI 131
+ FP +Q LP+G E+ + + LL + + + P CII+D
Sbjct: 267 VEFPYQQVDLPKGVESMTSTTDPITTWKIHNGAMLLNEAVGDFVEK--NPPDCIIADSAF 440
Query: 132 PWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILER---VDSESEYFTVPGIPDQIQ 188
W +A +IP ++F+G S F + ++T+ +L DS+S + VP +
Sbjct: 441 SWANDLAHKLQIPNLTFNGSSLFAVSIFHSLRTNNLLHTDADADSDSSSYVVPNLHHDNI 620
Query: 189 VTKEQIPGILKGELKEFGEKMHDAEMKSYGEIINTFEELE-KAYVKDYKKEKNGKVWFVG 247
+ P +L F M D +KS G IIN F EL+ + VK Y+K K W +G
Sbjct: 621 TLCSKPPKVLS----MFIGMMLDTVLKSTGYIINNFVELDGEECVKHYEKTTGHKAWHLG 788
Query: 248 PVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALA 307
P S K +KA++G + +SEH CL WL + SVVY C GS+ + QL E+A A
Sbjct: 789 PTSFIRKTVQEKAEKGNKSDVSEHECLNWLKSQRVNSVVYICFGSINHFSDKQLYEIACA 968
Query: 308 LEATNRPFIWVIRE----GNKSSEELEKWIS---EERN 338
+EA+ PFIWV+ E ++ EE EKW+ EERN
Sbjct: 969 VEASGHPFIWVVPEKKGKEDEIEEEKEKWLPKGFEERN 1082
>TC79138 weakly similar to GP|4115534|dbj|BAA36410.1 UDP-glycose:flavonoid
glycosyltransferase {Vigna mungo}, partial (36%)
Length = 1575
Score = 186 bits (471), Expect = 1e-47
Identities = 109/294 (37%), Positives = 161/294 (54%), Gaps = 1/294 (0%)
Frame = +2
Query: 26 PMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVTLNFPTKQAGLPEGC 85
P+ DIA L A G VTI TTP NA ++ +++ S +++ T+ FP+ Q GLP G
Sbjct: 59 PLCDIATLFASCGHHVTIITTPSNAQ----IILKSIPSHNHLRLHTVPFPSHQVGLPLGV 226
Query: 86 ENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFIPWTIQIAENHKIPR 145
EN V+++D + HA LL+ P + P CI++DF W ++A IPR
Sbjct: 227 ENLAFVNNVDNSCKIHHATMLLRSPINHFVEQ--DSPDCIVADFMFLWVDELANRLHIPR 400
Query: 146 ISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVTKEQIPGILKGELKEF 205
++F+GFS F + M + L+ D ES + G+P I T +P L +F
Sbjct: 401 LAFNGFSLFAICAM------ESLKARDFESSI--IQGLPHCI--TLNAMP---PKALTKF 541
Query: 206 GEKMHDAEMKSYGEIINTFEELE-KAYVKDYKKEKNGKVWFVGPVSLCNKDGLDKAQRGI 264
E + + E+KSYG I+N F EL+ + Y++ Y+K + W +GP SL + +KA RG
Sbjct: 542 MEPLLETELKSYGLIVNNFTELDGEEYIEHYEKTIGHRAWHLGPSSLICRTTQEKADRGQ 721
Query: 265 IASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALALEATNRPFIWV 318
+ + H CL WL+ QP SV+Y C GSLC+ QL E+A A+EA+ FIWV
Sbjct: 722 TSVVDVHECLSWLNSKQPNSVLYICFGSLCHFTNKQLYEIASAIEASGHQFIWV 883
>TC76351 similar to GP|7635494|emb|CAB88666.1 putative UDP-glycose {Cicer
arietinum}, partial (92%)
Length = 1645
Score = 175 bits (443), Expect = 3e-44
Identities = 107/323 (33%), Positives = 167/323 (51%), Gaps = 4/323 (1%)
Frame = +2
Query: 15 LFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVTLNF 74
+ P AQGH+IP++++A+L+A + VTI TTP NA F + ++G I++ + F
Sbjct: 107 MLPFFAQGHLIPLVNLARLVASKNQHVTIITTPSNAQLFDKTIEEEKAAGHHIRVHIIKF 286
Query: 75 PTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFIPWT 134
P+ Q GLP G EN S + H K E F P P ISD W+
Sbjct: 287 PSAQLGLPTGVENL-FAASDNQTAGKIHMAAHFVKADIEEFMKENP-PDVFISDIIFTWS 460
Query: 135 IQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVTKEQI 194
A+N +IPR+ F+ S F + + IQ+ E S+S + + G+P + + +
Sbjct: 461 ESTAKNLQIPRLVFNPISIFDVCMIQAIQSHP--ESFVSDSGPYQIHGLPHPLTLPIKPS 634
Query: 195 PGILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVGPVSLCNK 254
PG + E + +AE S+G I+N+F EL++ Y + Y+ KVW VGP SL +
Sbjct: 635 PGFAR-----LTESLIEAENDSHGVIVNSFAELDEGYTEYYENLTGRKVWHVGPTSLMVE 799
Query: 255 DGLDKAQRGII----ASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALALEA 310
+ K ++ + +SI++H L WLD +P SV+Y GSLC L QL E+A +EA
Sbjct: 800 --IPKKKKVVSTENDSSITKHQSLTWLDTKEPSSVLYISFGSLCRLSNEQLKEMANGIEA 973
Query: 311 TNRPFIWVIREGNKSSEELEKWI 333
+ F+WV+ K E+ + W+
Sbjct: 974 SKHQFLWVVH--GKEGEDEDNWL 1036
>TC79031 weakly similar to GP|13492674|gb|AAK28303.1
phenylpropanoid:glucosyltransferase 1 {Nicotiana
tabacum}, partial (32%)
Length = 1385
Score = 152 bits (383), Expect = 2e-37
Identities = 90/258 (34%), Positives = 139/258 (52%), Gaps = 12/258 (4%)
Frame = +1
Query: 95 DMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFIPWTIQIAENHKIPRISFHGFSCF 154
+M ++ + +LQ E+LF+ L +P I++D F PW+ +A+ IPRI FHG S
Sbjct: 16 EMIPKIYMGLYILQPDIEKLFETL--QPDFIVTDMFFPWSADVAKKLGIPRIMFHGASYL 189
Query: 155 CLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVTKEQIPGILK--GELKEFGEKMHDA 212
++ + + +S+++ F +P +PD++++T+ Q+P L+ + E + + ++
Sbjct: 190 ARSAAHSVEVYRPHLKAESDTDKFVIPDLPDELEMTRLQLPDWLRSPNQYAELMKVIKES 369
Query: 213 EMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVGPVSL-CNKDGLDKAQRGII---ASI 268
E KS+G + N+F +LE Y YKK K W +GPVSL N+D DKA RG
Sbjct: 370 EKKSFGSVFNSFYKLESEYYDHYKKVMGTKSWGLGPVSLWANQDDSDKAARGYARKEEGA 549
Query: 269 SEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALALEATNRPFIWVIR------EG 322
E LKWL+ SV+Y GS+ SQL+E+A ALE + FIWV+R EG
Sbjct: 550 KEEGWLKWLNSKPDGSVLYVSFGSMNKFPYSQLVEIAHALENSGHNFIWVVRKNEENEEG 729
Query: 323 NKSSEELEKWISEERNGY 340
EE EK + E GY
Sbjct: 730 GVFLEEFEKKMKESGKGY 783
>AW690449 weakly similar to PIR|T07786|T077 UDP-glucose glucosyltransferase
(EC 2.4.1.-) - potato, partial (14%)
Length = 573
Score = 149 bits (377), Expect = 1e-36
Identities = 73/155 (47%), Positives = 103/155 (66%)
Frame = +3
Query: 4 PANINDVPHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSS 63
P ND+ HF+ PL+A GH++PM+D+AKLLA+R V VTI TTP N+ RF S + R +
Sbjct: 27 PKCYNDL-HFIFIPLMAPGHLLPMVDMAKLLARRKVKVTILTTPLNSIRFQSTIDREIQL 203
Query: 64 GLQIKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPS 123
G QI+IV + FP+ ++G+PEGCE+ D + S+D+ N + A+ +Q E +F+ L P PS
Sbjct: 204 GSQIQIVHIKFPSVESGIPEGCESVDTLPSMDLMSNFYIALCKMQNSLENVFEKLXPIPS 383
Query: 124 CIISDFFIPWTIQIAENHKIPRISFHGFSCFCLHC 158
C+ISD I +IA K+PRI F G +CF L C
Sbjct: 384 CVISDKHISCVAEIAMKFKVPRIIFDGTNCFHLLC 488
>AJ501661 weakly similar to PIR|F84784|F847 probable glucosyl transferase
[imported] - Arabidopsis thaliana, partial (16%)
Length = 454
Score = 148 bits (373), Expect = 3e-36
Identities = 71/118 (60%), Positives = 93/118 (78%), Gaps = 1/118 (0%)
Frame = +3
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQR-GVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIV 70
HFVLFP++AQGH+IPM+DIAK+LAQ V+VTI TTP NASR+TS+L+R + SGL I++V
Sbjct: 96 HFVLFPMMAQGHMIPMMDIAKILAQHHNVMVTIVTTPHNASRYTSILARYLESGLHIQLV 275
Query: 71 TLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISD 128
L FP K++GLPEGCEN DM+ S+ N F++ LQ+ E+LF+ LTP P+CIISD
Sbjct: 276 QLKFPFKESGLPEGCENLDMLPSLGAATNFFNSSKFLQQEVEKLFEELTPSPTCIISD 449
>BG582596 weakly similar to GP|18151384|dbj ABA-glucosyltransferase {Vigna
angularis}, partial (8%)
Length = 729
Score = 146 bits (369), Expect = 1e-35
Identities = 87/227 (38%), Positives = 123/227 (53%), Gaps = 5/227 (2%)
Frame = +1
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
H P GH+IPMID A+L A+ GV VTI T NAS F + +SG IK
Sbjct: 37 HVTFLPYPTPGHMIPMIDTARLFAKHGVNVTIIATHANASTFQKSIDSDFNSGYSIKTQL 216
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFI 131
+ FP+ Q GLP+G EN S++M + I +LQ P E LF L +P CI++D
Sbjct: 217 IPFPSAQVGLPDGVENIKDGTSLEMLGKISSGILMLQDPIENLFHDL--RPDCIVTDQMY 390
Query: 132 PWTIQIAENHKIPRISFHGFSCF---CLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQ 188
WT++ A IPRI ++ S F H ++K + L S+++ FTVPG+P I+
Sbjct: 391 AWTVEAAAKLGIPRIHYYSSSYFSNCVFHFIMKYRPHNNLV---SDTQKFTVPGLPHTIE 561
Query: 189 VTKEQIPGIL--KGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVK 233
+T Q+P L K + + E M ++E +SYG + N+F EL YVK
Sbjct: 562 MTPLQLPDWLRTKNSVTAYFEPMFESEKRSYGTLYNSFHELGSDYVK 702
>TC90961 similar to GP|20067056|gb|AAM09517.1 putative glucosyltransferase
{Phaseolus lunatus}, partial (9%)
Length = 1130
Score = 144 bits (362), Expect = 7e-35
Identities = 104/326 (31%), Positives = 152/326 (45%), Gaps = 16/326 (4%)
Frame = +2
Query: 11 PHFVLFPLIAQGHIIPMIDIAKLLAQRGVI--VTIFTTPKNASRFTSVLSRAVSSG---L 65
PH V+ P +A GH+IP + +A+ + + +TI TTP N S +S SS +
Sbjct: 14 PHIVMTPFMAHGHLIPFLALARKIQETTTTFKITIATTPLNIQHLKSAISNTFSSSNNDI 193
Query: 66 QIKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPK---- 121
I + L F Q GLP EN + + D+ + FHA T L+ P L +T +
Sbjct: 194 SINLAELPFNHSQYGLPPNVENTEKLPLTDI-IKPFHASTSLEAPLSSLISKITQQEGQP 370
Query: 122 PSCIISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVP 181
P CIISD F+ W +A++ ISF + + I + + DS+ F VP
Sbjct: 371 PMCIISDVFLGWATNVAKSLGTRNISFTTCGAYGT*AHISIWCNLPHRKTDSDE--FWVP 544
Query: 182 GIPDQIQVTKEQIPGILKG-----ELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYK 236
G P + Q+ L+ + +F MKS G I NT EE+E ++ K
Sbjct: 545 GFPQNYRFHISQMHRYLRAADGTDDWSKFFPPQIALSMKSDGWICNTVEEIENLGLQLLK 724
Query: 237 KEKNGKVWFVGPV--SLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLC 294
VW +GP+ S K K + G + I+ C++WLDL SV+Y GS
Sbjct: 725 NYLQLPVWCIGPLLPSTTLKGSNSKYRAGKESGIALEECMEWLDLKDENSVLYISFGSQN 904
Query: 295 NLIPSQLMELALALEATNRPFIWVIR 320
+ SQ+M LA LE + + FIWVIR
Sbjct: 905 TVSASQMMALAEGLEESEKLFIWVIR 982
>BQ165264 weakly similar to PIR|T07786|T077 UDP-glucose glucosyltransferase
(EC 2.4.1.-) - potato, partial (7%)
Length = 847
Score = 135 bits (339), Expect = 3e-32
Identities = 86/262 (32%), Positives = 128/262 (48%), Gaps = 1/262 (0%)
Frame = +2
Query: 15 LFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVTLNF 74
L P ++ GH+IP+ DIA L A G VTI TTP NA FT LS +++ T++F
Sbjct: 89 LLPFLSPGHMIPLGDIATLFASHGQQVTIITTPSNAHFFTKSLSSV--DPFFLRLHTVDF 262
Query: 75 PTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFIPWT 134
P++Q GLP+G E+ D ++ LL P +E + P II D PW
Sbjct: 263 PSQQVGLPDGVESLSSNIDTDTTHKIYVGSMLLHGPIKEFIE--KDPPDYIIGDCVFPWI 436
Query: 135 IQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVTKEQI 194
+A I ++F G+S F + + ++ + +S+S F VP P I
Sbjct: 437 HDLANKPHISTLAFTGYSLFSVSLIEALRVHRSNSHTNSDSSSFVVPNFPHSITFNSGPP 616
Query: 195 PGILKGELKEFGEKMHDAEMKSYGEIINTFEELE-KAYVKDYKKEKNGKVWFVGPVSLCN 253
+ EF E M +KS G IIN F EL+ + +K Y+K K W +GP L +
Sbjct: 617 KTFI-----EFEEGMLKTIIKSKGLIINNFVELDGEDCIKHYEKTMGHKAWHLGPACLIH 781
Query: 254 KDGLDKAQRGIIASISEHHCLK 275
+ +KA+RG + +S H CL+
Sbjct: 782 ESVQEKAERGNESVVSMHECLR 847
>TC93563 weakly similar to PIR|T07786|T07786 UDP-glucose glucosyltransferase
(EC 2.4.1.-) - potato, partial (7%)
Length = 855
Score = 130 bits (327), Expect = 7e-31
Identities = 79/226 (34%), Positives = 114/226 (49%), Gaps = 3/226 (1%)
Frame = +3
Query: 14 VLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVTLN 73
+ P + GH+ PMID A+L A+ G+ VTI TT NA F + G I+ +
Sbjct: 165 IFLPYLTPGHMNPMIDTARLFAKHGINVTIITTHANALLFKKAIDNDTCCGYSIRTCVIQ 344
Query: 74 FPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFIPW 133
FP+ Q GLPEG EN S++M + H I+LLQ E LF L +P CI+SD F PW
Sbjct: 345 FPSAQVGLPEGVENIKDGTSLEMLGKIGHGISLLQDQIEILFQDL--QPDCIVSDMFYPW 518
Query: 134 TIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVTKEQ 193
T++ A +PRI ++ S F C I+ K E + S+ + F++P +P I++T Q
Sbjct: 519 TVESAAKLGVPRIYYYSSSYFSSCCAHFIRKYKPHENLVSDGQLFSIPELPHNIEITSLQ 698
Query: 194 IPGIL---KGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYK 236
+ K L+ F + + K+ I F LE Y YK
Sbjct: 699 LEEWCRT*KSILRLFWMWYMNQKAKATEHCIIAFMNLESDYEPLYK 836
>BG584431 weakly similar to GP|19911201|dbj glucosyltransferase-9 {Vigna
angularis}, partial (15%)
Length = 759
Score = 120 bits (302), Expect(2) = 5e-29
Identities = 70/183 (38%), Positives = 100/183 (54%)
Frame = +2
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
H V P A GH+ PMID A+L A+ GV VTI T NASRF + VS G IK
Sbjct: 56 HVVFLPYPAIGHMNPMIDTARLFAKHGVNVTIILTHANASRFQKSIDSDVSLGYSIKTQL 235
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFI 131
L FP+ Q GLPEG EN + S +M + + +L+ E LF L +P CI++D
Sbjct: 236 LQFPSAQVGLPEGIENMNDATSREMLSKVTRGVWMLKDSFEVLFKDL--QPDCIVTDMMY 409
Query: 132 PWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVTK 191
PWT++ A IPRI F S F + ++ K + S+++ FT+P +P +++T+
Sbjct: 410 PWTVESAAKLNIPRIHFCSSSYFSDCGIYFVRKYKPHYNLVSDTQKFTIPCLPHTVEMTR 589
Query: 192 EQI 194
Q+
Sbjct: 590 LQL 598
Score = 24.6 bits (52), Expect(2) = 5e-29
Identities = 9/17 (52%), Positives = 12/17 (69%)
Frame = +1
Query: 215 KSYGEIINTFEELEKAY 231
K+YG + N+F ELE Y
Sbjct: 664 KNYGSLYNSFHELESDY 714
>TC79293 weakly similar to GP|15290056|dbj|BAB63750. putative
glucosyltransferase IS5a salicylate-induced {Oryza
sativa (japonica cultivar-group)}, partial (7%)
Length = 866
Score = 122 bits (306), Expect = 2e-28
Identities = 81/265 (30%), Positives = 123/265 (45%), Gaps = 1/265 (0%)
Frame = +3
Query: 15 LFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVTLNF 74
+ P ++ GH+IP+ DIA L A G VTI TTP NA F ++ L++ IV +F
Sbjct: 90 MLPFLSPGHMIPLGDIAALFASHGQQVTIITTPSNAHFFDKSIASVDPFFLRLHIV--DF 263
Query: 75 PTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFIPWT 134
P++Q LP+G E+ + LL +P E + +P II+D PW
Sbjct: 264 PSQQVDLPDGVESLSSTTGPATMAKICKGANLLHEPIREFVE--KDQPDYIIADCVYPWI 437
Query: 135 IQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVTKEQI 194
+ I I+F GFS F + + ++ ++ +S F P P I
Sbjct: 438 NDLTNKPHISTIAFTGFSLFTISLIESLRINRSYFDKNSSLSSFVDPNFPHSITFC---- 605
Query: 195 PGILKGELKEFGEKMHDAEMKSYGEIINTFEELE-KAYVKDYKKEKNGKVWFVGPVSLCN 253
+L F E+M + KS G I+N F EL+ + +K Y+K K W +GP L
Sbjct: 606 -ATTPKQLIAFEERMLETIRKSKGLIVNNFAELDGEDCIKHYEKTMGYKAWHLGPACLIR 782
Query: 254 KDGLDKAQRGIIASISEHHCLKWLD 278
K +K+ RG + +S H C WL+
Sbjct: 783 KTFQEKSVRGNESVVSVHECPSWLN 857
>BG645363 weakly similar to GP|15290080|dbj putative salicylate-induced
glucosyltransferase {Oryza sativa (japonica
cultivar-group)}, partial (9%)
Length = 796
Score = 117 bits (294), Expect = 5e-27
Identities = 81/251 (32%), Positives = 119/251 (47%), Gaps = 1/251 (0%)
Frame = +3
Query: 14 VLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVTLN 73
++ P +A GH+IP+ DIA L A G VTI TTP NA FT LS +++ T++
Sbjct: 48 LILPFLAPGHMIPLGDIAALFASHGQQVTIITTPSNAHFFTKSLS--FVDPFFLRLHTVD 221
Query: 74 FPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFIPW 133
FP++Q LP+G E+ + D + LL +P E + +P II+D PW
Sbjct: 222 FPSQQVDLPDGVESLSSTTNPDTMAKICKGAMLLHEPIREFVE--KDQPDYIIADCVYPW 395
Query: 134 TIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVTKEQ 193
+A I I+F F F + + ++ + +S S F P P I
Sbjct: 396 INDLANKPHISTIAFTPFFLFTVSLIESLRIKRSYSDKNSSSSSFVDPNFPHSINFCSTP 575
Query: 194 IPGILKGELKEFGEKMHDAEMKSYGEIINTFEELE-KAYVKDYKKEKNGKVWFVGPVSLC 252
P + F E+M +A KS G IIN F EL+ + +K Y+K K W +GP L
Sbjct: 576 -PKLYVA----FEERMLEAIRKSKGLIINNFVELDGEDCIKHYEKTMGYKAWHLGPTCLI 740
Query: 253 NKDGLDKAQRG 263
K +K+ RG
Sbjct: 741 RKTF*EKSMRG 773
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.139 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,670,564
Number of Sequences: 36976
Number of extensions: 175093
Number of successful extensions: 1222
Number of sequences better than 10.0: 170
Number of HSP's better than 10.0 without gapping: 1100
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1112
length of query: 346
length of database: 9,014,727
effective HSP length: 97
effective length of query: 249
effective length of database: 5,428,055
effective search space: 1351585695
effective search space used: 1351585695
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC147435.1


