

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147434.3 - phase: 0
(376 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
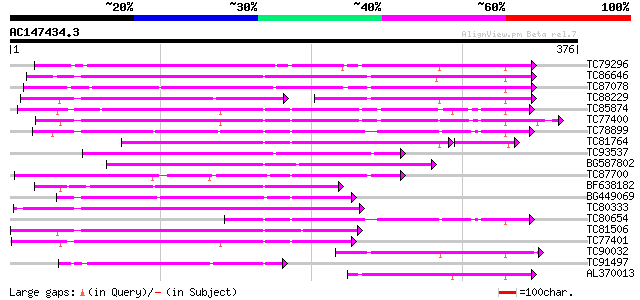
TC79296 weakly similar to GP|14279169|gb|AAK58515.1 beta-1 3-glu... 174 4e-44
TC86646 similar to PIR|E84471|E84471 probable beta-1 3-glucanase... 168 3e-42
TC87078 similar to PIR|T50645|T50645 glucan endo-1 3-beta-D-gluc... 163 9e-41
TC88229 similar to PIR|E84471|E84471 probable beta-1 3-glucanase... 100 7e-36
TC85874 homologue to PIR|T09401|T09401 1 3-beta-glucanase (EC 3.... 140 6e-34
TC77400 similar to GP|1247325|emb|CAA01814.1 beta-1 3-glucanase ... 138 3e-33
TC78899 similar to GP|3900936|emb|CAA10167.1 glucan endo-1 3-bet... 134 7e-32
TC81764 similar to GP|17104805|gb|AAL34291.1 putative glucan end... 122 8e-30
TC93537 weakly similar to GP|6714534|dbj|BAA89481.1 beta-1 3-glu... 122 2e-28
BG587802 similar to GP|20197543|gb putative beta-1 3-glucanase {... 116 1e-26
TC87700 weakly similar to PIR|S31196|S31196 hypothetical protein... 115 3e-26
BF638182 weakly similar to PIR|T45594|T45 glucosidase-like prote... 103 8e-23
BG449069 weakly similar to GP|21741726|em OSJNBa0073L04.6 {Oryza... 103 8e-23
TC80333 similar to GP|9758115|dbj|BAB08587.1 beta-1 3-glucanase-... 102 2e-22
TC80654 similar to GP|3900936|emb|CAA10167.1 glucan endo-1 3-bet... 99 2e-21
TC81506 weakly similar to GP|22748323|gb|AAN05325.1 Putative bet... 98 6e-21
TC77401 similar to GP|1247325|emb|CAA01814.1 beta-1 3-glucanase ... 94 9e-20
TC90032 similar to GP|14090345|dbj|BAB55504. contains EST D15166... 87 1e-17
TC91497 similar to PIR|T05960|T05960 beta-1 3-glucanase (EC 3.2.... 85 5e-17
AL370013 similar to PIR|T05268|T05 hypothetical protein T4L20.60... 82 3e-16
>TC79296 weakly similar to GP|14279169|gb|AAK58515.1 beta-1 3-glucanase-like
protein {Olea europaea}, partial (90%)
Length = 1710
Score = 174 bits (442), Expect = 4e-44
Identities = 122/343 (35%), Positives = 183/343 (52%), Gaps = 10/343 (2%)
Frame = +3
Query: 17 YLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDPSII 76
+ L ++ + S +GV Y T D+ PPP + +Q+ + +R+ DP+II
Sbjct: 132 FSLFAIALSFADSQSFVGVNYGQTA--DNLPPP--EATAKLLQSTTIGKVRIYGADPAII 299
Query: 77 RSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTDVYP 136
+SL + I + + N + +A++ + A WI T+VLP+YP + IT I+VGN +
Sbjct: 300 KSLANSGIGIVIGAANNDIPSLASDPNAATQWINTNVLPYYPASNITLITVGNEVLNSGD 479
Query: 137 ES-INNLLPAISNVHISLRDLGI-RKISVSTSFSFVTAVTSPFPPSSAAFQEIPGV-NLI 193
E ++ L+PAI NV +L + + K+ V+T S S PPSS +F P + N +
Sbjct: 480 EGLVSQLMPAIRNVQTALSSVKLGGKVKVTTVHSMAVLAQSD-PPSSGSFN--PALRNTL 650
Query: 194 GPLLQFLSDTNSSFLINLYPYNLYRL--RPEIPLGIALFQEHPFNFRDDFTTGVRYKNLF 251
LL FL D S F +N YP+ Y+ RPE L LFQ P + R D G Y N+F
Sbjct: 651 NQLLAFLKDNKSPFTVNPYPFFAYQSDPRPE-TLTFCLFQ--PNSGRVDSGNGKLYTNMF 821
Query: 252 DIMVDAVVSAMALEGYETIPVIVTETGWPSSG--SELDANSAYAEIYLKGLVKHLKSGAG 309
D VDAV SA++ YE I ++V ETGWPSSG +E+ + A+ Y L+ HL+S G
Sbjct: 822 DAQVDAVHSALSAMSYEDIEIVVAETGWPSSGDNNEVGPSVENAKAYNGNLITHLRSLVG 1001
Query: 310 TPLLKDGVKGVYLYELFD---KEGAGNGRNWGILYPNGSTKYD 349
TPL+ Y++ L+D K G G+ R +G+ + S YD
Sbjct: 1002TPLIPGKSVDTYIFALYDEDLKPGPGSERAFGLFKTDLSMSYD 1130
>TC86646 similar to PIR|E84471|E84471 probable beta-1 3-glucanase [imported]
- Arabidopsis thaliana, partial (73%)
Length = 1257
Score = 168 bits (426), Expect = 3e-42
Identities = 107/345 (31%), Positives = 186/345 (53%), Gaps = 7/345 (2%)
Frame = +2
Query: 12 TTFSLYLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEP 71
T F+ ++ L+ + T IG+ Y PS ++ +++ +TH+++ +
Sbjct: 89 TLFTFFIFLTFIALTNAG--NIGINYGRIANN----LPSATKVVQLLKSQGITHVKIYDT 250
Query: 72 DPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAF 131
DPS++R+L + I L + +PN + A ++S A +W+ +++ + P I I+VGN
Sbjct: 251 DPSVLRALSGSKIKLTVDLPNQQLFAAAKSQSFALSWVERNIVAYQPNTIIEAIAVGNEV 430
Query: 132 TDVYPESINNLLPAISNVHISLRDLGIRK-ISVSTSFSFVTAVTSPFPPSSAAFQEIPGV 190
S L+PA+ N++ SL+ + I VS+ + ++A+ + +P SS +F+
Sbjct: 431 FVDPNNSTKYLVPAMKNIYRSLQKHNLHNDIKVSSPIA-LSALGNSYPSSSGSFRPELIQ 607
Query: 191 NLIGPLLQFLSDTNSSFLINLYPYNLYRLRPE-IPLGIALFQEHPFNFRDDFTTGVRYKN 249
+ P+L FL +T S ++N+YP+ Y + I L ALF+E+P D G+RY N
Sbjct: 608 PVFKPMLDFLRETGSYLMVNVYPFFAYESNADVISLDYALFRENPGQV--DAGNGLRYLN 781
Query: 250 LFDIMVDAVVSAMALEGYETIPVIVTETGWPS--SGSELDANSAYAEIYLKGLVKHLKSG 307
LFD +DAV +A++ Y+ I V+V+ETGWPS G+E+ A+ A Y LV+ + +
Sbjct: 782 LFDAQIDAVFAALSRLKYDDINVVVSETGWPSKGDGNEVGASVENAAAYNANLVRKILTS 961
Query: 308 AGTPLLKDGVKGVYLYELFD---KEGAGNGRNWGILYPNGSTKYD 349
GTPL V+L+ LF+ K G + RN+G+ YP+ Y+
Sbjct: 962 KGTPLRPKADLTVFLFALFNENQKPGPTSERNFGLFYPDEKKVYN 1096
>TC87078 similar to PIR|T50645|T50645 glucan endo-1 3-beta-D-glucosidase (EC
3.2.1.39) [imported] - garden pea, partial (94%)
Length = 1628
Score = 163 bits (413), Expect = 9e-41
Identities = 118/345 (34%), Positives = 182/345 (52%), Gaps = 5/345 (1%)
Frame = +1
Query: 10 SLTTFSLYLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLE 69
++TT + L L LL + T + +IGV Y T D+ PPP+ N + +++
Sbjct: 16 NMTTKTTLLFLFLLHVSVTVVNSIGVNYG--TLADNLPPPATVA-NFLKTNTIVDRVKIF 186
Query: 70 EPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGN 129
+ P I+++ T IS+ +T PN + + N + AR W+ + PFYP KI I VG+
Sbjct: 187 DVSPQILQAFANTGISVTVTAPNGDIAALG-NINSARQWVQQKIKPFYPATKINYILVGS 363
Query: 130 AFTDVYPES-INNLLPAISNVHISLRDLGIRKISVSTSFSFVTAVTSPFPPSSAAFQEIP 188
+ I L+PA+ +H +L GI I V+T+ S + S PPS+ F+
Sbjct: 364 EVLHWGDGNMIRGLVPAMRTLHSALVAEGINDIKVTTAHSLIIMRQS-LPPSAGKFRPGF 540
Query: 189 GVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRPEIPLGIALFQEHPFNFRDDFTTGVRYK 248
++I P+L+FL +T + F++N YPY Y + + ALF+ + F D T + Y
Sbjct: 541 AKHVIAPMLKFLRETRTPFMVNPYPYFGYNPKN---VNFALFRPNRGLF--DRNTRLTYT 705
Query: 249 NLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANS-AYAEIYLKGLVKHLKSG 307
N FD ++DAV SAM G+ + + V ETGWPS DA S A A+ Y L++HL++G
Sbjct: 706 NQFDALMDAVYSAMKGLGFGDVDIAVGETGWPSVCDGWDACSVANAQSYNGELIRHLEAG 885
Query: 308 AGTPLLKDGVKGVYLYELFD---KEGAGNGRNWGILYPNGSTKYD 349
GTPL+ + +L+ LF+ K G RNWG+ P+ S YD
Sbjct: 886 RGTPLMPNRRFETFLFALFNENQKPGPIAERNWGLFRPDFSPVYD 1020
>TC88229 similar to PIR|E84471|E84471 probable beta-1 3-glucanase [imported]
- Arabidopsis thaliana, partial (69%)
Length = 1368
Score = 100 bits (249), Expect(2) = 7e-36
Identities = 59/153 (38%), Positives = 90/153 (58%), Gaps = 6/153 (3%)
Frame = +3
Query: 203 TNSSFLINLYPYNLYRLRPE-IPLGIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSA 261
T+S ++N+YP+ Y + I L ALF+E+P N D G++Y N+FD +DAV +A
Sbjct: 606 TSSYLMVNVYPFFAYESNADVISLNYALFRENPGNV--DPGNGLKYYNIFDAQIDAVFAA 779
Query: 262 MALEGYETIPVIVTETGWPSSG--SELDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKG 319
+ + Y+ + V+V+ETGWPS G +E+ A+ A Y LVK + + GTPL +
Sbjct: 780 LNVLQYDDVRVVVSETGWPSKGDSNEVGASPQNAAAYNGNLVKKILNNGGTPLRPNANLT 959
Query: 320 VYLYELFD---KEGAGNGRNWGILYPNGSTKYD 349
VYL+ LF+ K G + RN+G+ YP+ YD
Sbjct: 960 VYLFALFNENGKVGLTSERNFGMFYPDMKKVYD 1058
Score = 68.2 bits (165), Expect(2) = 7e-36
Identities = 47/185 (25%), Positives = 92/185 (49%), Gaps = 7/185 (3%)
Frame = +1
Query: 8 MISLTTFSLYLLLSLLPTTTTSLP------TIGVTYSSTTRQDSPPPPSPDRITTAMQNL 61
+I T L+LS+L ++ SLP +IGV Y PS ++ +++
Sbjct: 13 LIPYPTMHCILILSILISSYFSLPLLADGSSIGVNYGRIANN----LPSAFKVVKLLKSQ 180
Query: 62 KLTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAK 121
+ ++L + DP++++SL + I + + +PN + A S A W+ +V+ ++P+ +
Sbjct: 181 GIDRVKLYDTDPAVLKSLSGSGIKVTVNLPNEQLFHTARKLSYALTWLQKNVVVYHPKTQ 360
Query: 122 ITTISVGN-AFTDVYPESINNLLPAISNVHISLRDLGIRKISVSTSFSFVTAVTSPFPPS 180
I I+VGN F D + + L+PA+ N+H +L + +S ++A+ S +P
Sbjct: 361 IEAIAVGNEVFVDTH-NTTKYLIPAMKNIHKALVKFNLHNSIKISSPIALSALGSSYPSF 537
Query: 181 SAAFQ 185
+ Q
Sbjct: 538 NRVIQ 552
>TC85874 homologue to PIR|T09401|T09401 1 3-beta-glucanase (EC 3.2.1.-)
acidic - alfalfa, complete
Length = 1336
Score = 140 bits (354), Expect = 6e-34
Identities = 114/356 (32%), Positives = 182/356 (51%), Gaps = 13/356 (3%)
Frame = +1
Query: 6 TFMISLTTFSLYLLLSLLPTTTTSL-----PTIGVTYSSTTRQDSPPPPSPDRITTAMQN 60
+F FSL L LL T +L IG+ Y + PP+ + I N
Sbjct: 64 SFFAPTMRFSLASPLFLLGLFTINLIHTADAQIGICYGM---MGNNLPPANEVINLYKAN 234
Query: 61 LKLTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRA 120
+ +RL +P+ + + +L + I L L +PN + +ATN AR W+ +VL F+P
Sbjct: 235 -NIKRMRLYDPNQAALNALRNSGIELILGVPNSDLQTLATNSDNARQWVQRNVLNFWPSV 411
Query: 121 KITTISVGNAFTDVYPES--INNLLPAISNVHISLRDLGIR-KISVSTSFSFVTAVTSPF 177
KI I+VGN + V S +LPA N++ ++R G+ +I VST+ +T + + F
Sbjct: 412 KIKYIAVGNEVSPVGGSSWLAQYVLPATQNIYQAIRAQGLHDQIKVSTAID-MTLIGNSF 588
Query: 178 PPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLGIALFQEHPFN 236
PPS +F+ + + P + +L + L+N+YPY + P +I L ALF P
Sbjct: 589 PPSKGSFRN-DVRSYLDPFIGYLVYAGAPLLVNVYPYFSHVGNPRDISLPYALFTS-PGV 762
Query: 237 FRDDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAY--AE 294
D G Y+NLFD M+D+V +A+ G + V+V+E+GWPS G A ++Y A
Sbjct: 763 MVQDGPNG--YQNLFDAMLDSVHAALDNTGIGWVNVVVSESGWPSDGG---AATSYDNAR 927
Query: 295 IYLKGLVKHLKSGAGTPLLKDGVKGVYLYELFD--KEGAGNGRNWGILYPNGSTKY 348
IYL L++H+ G GTP + Y++ +FD ++ +++G+ YPN KY
Sbjct: 928 IYLDNLIRHV--GKGTP-RRPWATETYIFAMFDENQKSPELEKHFGVFYPNKQKKY 1086
>TC77400 similar to GP|1247325|emb|CAA01814.1 beta-1 3-glucanase {Glycine
max}, partial (88%)
Length = 1352
Score = 138 bits (348), Expect = 3e-33
Identities = 116/363 (31%), Positives = 183/363 (49%), Gaps = 13/363 (3%)
Frame = +3
Query: 18 LLLSLLPTTTTSLPT--IGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDPSI 75
LLL L T L + IGV Y PS ++ + +RL +P+ +
Sbjct: 153 LLLFALFTVNLRLTSAQIGVCYGMMGNN----LPSQREAIDLCKSNNIKRMRLYDPNQAA 320
Query: 76 IRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTDVY 135
+ +L + I L L +PN + IATN IA W+ +VL FYP KI I+VGN V
Sbjct: 321 LEALRNSGIELMLGVPNSDLQNIATNNDIAIQWVQKNVLNFYPSVKIKYIAVGNEVNPVG 500
Query: 136 PES--INNLLPAISNVHISLRDLGIR-KISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNL 192
S +LPAI N++ ++R + +I VST+ +T + + +PPS +F+ +
Sbjct: 501 GSSQFAKFVLPAIQNIYQAIRAKNLHDQIKVSTAID-MTMIGTSYPPSKGSFRS-DVRSY 674
Query: 193 IGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLGIALFQEHPFNFRDDFTTGVRYKNLF 251
+ P++ +L N+ N+Y Y Y+ P +I L ALF P D + G Y+NLF
Sbjct: 675 LDPIIGYLVYANAPLFANIYSYFSYKDNPKDISLQYALFTS-PNVVVWDGSRG--YQNLF 845
Query: 252 DIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLKGLVKHLKSGAGTP 311
D ++D++ +A+ G + V+V+E+GWPS G A +YL L++H+K GTP
Sbjct: 846 DALLDSLHAAIDNTGIGFVKVVVSESGWPSDGG-FATTYDNARVYLDNLIRHVK--GGTP 1016
Query: 312 LLKDGVKGVYLYELFD--KEGAGNGRNWGILYPNGSTKY-----DKIDFSSGWKRLVNVG 364
++ G Y++ LFD ++ +++G+ YPN KY KID ++ L NV
Sbjct: 1017-MRSGPIETYIFGLFDENQKNPELEKHFGVFYPNKQKKYPFGFQGKIDGTN----LFNVS 1181
Query: 365 FIL 367
F L
Sbjct: 1182FPL 1190
>TC78899 similar to GP|3900936|emb|CAA10167.1 glucan endo-1
3-beta-d-glucosidase {Cicer arietinum}, partial (94%)
Length = 1296
Score = 134 bits (336), Expect = 7e-32
Identities = 100/340 (29%), Positives = 172/340 (50%), Gaps = 7/340 (2%)
Frame = +3
Query: 16 LYLLLSLLPTTT--TSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDP 73
++LL+ +L T++ +IGV Y PS + ++ + +R+ PD
Sbjct: 72 IFLLVGILSIGLQFTAVESIGVCYGMIGNN----LPSRQDVVNLYKSRGINQMRIFFPDE 239
Query: 74 SIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTD 133
+++L +NI L L + + P N + A W+ +V P+ KI ISVGN
Sbjct: 240 PALQALRGSNIELILDVAKETL-PSLRNANEATNWVNKYVRPYAQNVKIKYISVGNEIKP 416
Query: 134 VYPESINNLLPAISNVHISLRDLGIR-KISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNL 192
E+ +LPA+ N+ ++ ++ +I VST+ +T + FPP+ F +
Sbjct: 417 NDNEA-QYILPAMQNIQNAISSANLQGQIKVSTAID-MTLIGKSFPPNDGVFSD-QAKPY 587
Query: 193 IGPLLQFLSDTNSSFLINLYPYNLY-RLRPEIPLGIALFQEHPFNFRDDFTTGVRYKNLF 251
I P++ FL++ + L N+YPY Y + IPL ALF++ N V Y+NLF
Sbjct: 588 IQPIINFLNNNGAPLLANVYPYFAYIGDKVNIPLDYALFRQQGNN-------AVGYQNLF 746
Query: 252 DIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLKGLVKHLKSGAGTP 311
D +D+V +A+ G + ++V+E+GWPS+ + A++ A Y + L+ H+K+ GTP
Sbjct: 747 DAQLDSVYAALEKVGASGVKIVVSESGWPSAAGD-SASTDNAATYYRNLINHVKN--GTP 917
Query: 312 LLKDGVKGVYLYELFD---KEGAGNGRNWGILYPNGSTKY 348
+ G YL+ +FD K GA +++G+ P+ S KY
Sbjct: 918 -KRPGAIETYLFAMFDENQKTGAATEQHFGLFNPDKSPKY 1034
>TC81764 similar to GP|17104805|gb|AAL34291.1 putative glucan endo-1
3-beta-glucosidase precursor {Arabidopsis thaliana},
partial (71%)
Length = 1560
Score = 122 bits (305), Expect(2) = 8e-30
Identities = 84/225 (37%), Positives = 124/225 (54%), Gaps = 5/225 (2%)
Frame = +2
Query: 75 IIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTDV 134
++++L TNI + + + N V I + S A AWI +V+ + P IT I+VG+
Sbjct: 32 LLQALSKTNIDVMVGVTNEEVLRIGESPSAAAAWINKNVVAYVPSTNITAIAVGSEVLST 211
Query: 135 YPESINNLLPAISNVHISLRDLGIR-KISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNLI 193
P L+PA++++H +L + ++ VST S + + PFPPS+A F + I
Sbjct: 212 IPNVAPVLVPAMNSLHKALVAANLNFRVKVSTPQS-MDIIPKPFPPSTATFNS-SWNSTI 385
Query: 194 GPLLQFLSDTNSSFLINLYPYNLYRLRPEI-PLGIALFQEHP-FNFRDDFTTGVRYKNLF 251
+LQFL +TNSSF++N YPY Y I PL ALF+ P D T Y ++F
Sbjct: 386 YQVLQFLRNTNSSFMLNAYPYYGYTKGDGIFPLEYALFRPLPSVKQIVDPNTLYHYNSMF 565
Query: 252 DIMVDAVVSAMALEGYETIPVIVTETGWPSSG--SELDANSAYAE 294
D MVDA ++ ++ IPV+VTETGWPS G +E DA + AE
Sbjct: 566 DAMVDATYYSIDALNFKDIPVVVTETGWPSFGGANEPDATAENAE 700
Score = 26.2 bits (56), Expect(2) = 8e-30
Identities = 12/46 (26%), Positives = 25/46 (54%), Gaps = 3/46 (6%)
Frame = +3
Query: 296 YLKGLVKHLKSGAGTPLLKDGVKGVYLYELFDKE---GAGNGRNWG 338
Y +++ + + +G P + Y+YELF+++ G + +NWG
Sbjct: 705 YNNNMIQRVLNDSGPPSQPNIPINTYIYELFNEDKRNGPVSEKNWG 842
>TC93537 weakly similar to GP|6714534|dbj|BAA89481.1 beta-1 3-glucanase
{Salix gilgiana}, partial (46%)
Length = 736
Score = 122 bits (307), Expect = 2e-28
Identities = 70/216 (32%), Positives = 125/216 (57%), Gaps = 2/216 (0%)
Frame = +1
Query: 49 PSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAW 108
P+P + ++NLK +++ + +P I+++L T I + + +PN LVT +++N+++A W
Sbjct: 91 PTPTTSVSLIKNLKAKRVKIYDANPQILKALENTGIQVSIMLPNELVTNVSSNQTLANQW 270
Query: 109 IYTHVLPFYPRAKITTISVGN-AFTDVYPESINNLLPAISNVHISLRDLGIRKISVSTSF 167
+ T+++PFY + I + VGN + ++ +++PA+ + SL G+ K+ V T
Sbjct: 271 VQTNLVPFYSKTLIRYLLVGNELISSTTNQTWPHIVPAMYRMKHSLTIFGLHKVKVGTPL 450
Query: 168 SFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLG 226
+ TS FPPS+ F+ ++++ P+++FL TNS F +++YP+ + P I L
Sbjct: 451 AMDVLQTS-FPPSNGTFRNDIALSVMKPMVEFLHVTNSFFFLDVYPFFAWTSDPININLD 627
Query: 227 IALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSAM 262
ALF+ D TG+ Y NLF+ MVDAV AM
Sbjct: 628 YALFESDNITVTDS-GTGLVYTNLFEQMVDAVYFAM 732
>BG587802 similar to GP|20197543|gb putative beta-1 3-glucanase {Arabidopsis
thaliana}, partial (48%)
Length = 753
Score = 116 bits (291), Expect = 1e-26
Identities = 75/222 (33%), Positives = 118/222 (52%), Gaps = 3/222 (1%)
Frame = +1
Query: 65 HLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITT 124
H+RL + D S++ +L T I + +++PN + I + + A W+ +V+ P IT
Sbjct: 4 HVRLFDADKSMLLALAKTGIQVIVSVPNDELLGIGQSNATAANWVARNVIAHVPSTNITA 183
Query: 125 ISVGNAFTDVYPESINNLLPAISNVHISLRDLGIR-KISVSTSFSFVTAVTSPFPPSSAA 183
I+VG+ P + L+ A+ + +L + +I VST S + + FPPS A
Sbjct: 184 IAVGSEVLTSLPNAAPVLVSALQFIQSALVAANLDDQIKVSTPLS-TSIILDSFPPSQAF 360
Query: 184 FQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRPE-IPLGIALFQEHPFNFRD-DF 241
F ++ PLL+FL T S ++N+YPY Y + IPL ALF+ P N D
Sbjct: 361 FNRTWDP-VMSPLLKFLQSTGSYLMLNVYPYYDYMQSNDVIPLDYALFRPLPPNKEAVDS 537
Query: 242 TTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSG 283
T + Y N+FD ++DA AM+ + IP++VTE+GWPS G
Sbjct: 538 NTLLHYTNVFDAVIDAAYFAMSYLKFTNIPILVTESGWPSKG 663
>TC87700 weakly similar to PIR|S31196|S31196 hypothetical protein - potato,
partial (47%)
Length = 791
Score = 115 bits (288), Expect = 3e-26
Identities = 77/266 (28%), Positives = 148/266 (54%), Gaps = 7/266 (2%)
Frame = +3
Query: 4 HSTFMISLTTFSLYLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKL 63
H +++++F +++ L + + + G+ + Q + PS R+ +++L +
Sbjct: 24 HFWVRMAISSFCFRVIMLFLTFSDLFMQSYGLDFGINYGQIANNLPSHSRVAVLIKSLNV 203
Query: 64 THLRLEEPDPSIIRSLLYTNISLFLTIPNY--LVTPIATNRSIARAWIYTHVLPFYPRAK 121
+ ++L + DP+++ + +N+ + + + + PI A++W+ +V P+ P+ K
Sbjct: 204 SRIKLYDADPNVLTTFSNSNVEFMIGLNDLQSMKDPIK-----AQSWVQQNVQPYLPQTK 368
Query: 122 ITTISVGNAF---TDVYPESINNLLPAISNVHISLRDLGI-RKISVSTSFSFVTAVTSPF 177
IT+I+VGN D+ S NNLLPA+ +V+ +L +LG+ ++++V+TS SF+ +++ F
Sbjct: 369 ITSINVGNEVLGNNDI--NSYNNLLPAMKSVYNALVNLGLSQQVTVTTSHSFII-MSNSF 539
Query: 178 PPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRPE-IPLGIALFQEHPFN 236
PPSS AF+E + I PLL F + S FLIN YP+ Y+ P+ + L LFQ + +
Sbjct: 540 PPSSGAFRE-DLIQYIQPLLSFQAQIKSPFLINAYPFFAYKGDPQHVSLNYVLFQPNAGS 716
Query: 237 FRDDFTTGVRYKNLFDIMVDAVVSAM 262
D T + Y N+ +DA +A+
Sbjct: 717 I--DPATNLHYDNMLYAQIDARYAAI 788
>BF638182 weakly similar to PIR|T45594|T45 glucosidase-like protein -
Arabidopsis thaliana, partial (46%)
Length = 674
Score = 103 bits (258), Expect = 8e-23
Identities = 71/208 (34%), Positives = 116/208 (55%), Gaps = 3/208 (1%)
Frame = +2
Query: 17 YLLLSLLPTTTTSLPT--IGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDPS 74
+LLL LL T+ S T IGV Y T + PPP + T +TH+++ + + S
Sbjct: 20 FLLLLLLSTSNFSSATYNIGVNYG-TIANNLPPPTTV--ATFLKTQTTITHIKIFDTNSS 190
Query: 75 IIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTDV 134
I+R+ TNIS+ +T+ N + P T A+ WI T++LPF+P+ I+VGN
Sbjct: 191 ILRAFANTNISVTVTVSNADI-PSLTKLPSAQKWITTNILPFHPKTIFNRIAVGNEILAT 367
Query: 135 YPES-INNLLPAISNVHISLRDLGIRKISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNLI 193
++ I ++LPA++ +H +L + I + + S + ++S PPSSAAF+ V +
Sbjct: 368 SDKTLIAHILPAMNALHQALTLSNLTHIQIVSPNS-LGILSSSSPPSSAAFRRGYDVTIF 544
Query: 194 GPLLQFLSDTNSSFLINLYPYNLYRLRP 221
P+L+F +T S F+IN YP+ Y ++P
Sbjct: 545 TPILKFHRETKSPFMINPYPFFRYFIKP 628
>BG449069 weakly similar to GP|21741726|em OSJNBa0073L04.6 {Oryza sativa},
partial (46%)
Length = 668
Score = 103 bits (258), Expect = 8e-23
Identities = 68/202 (33%), Positives = 114/202 (55%), Gaps = 3/202 (1%)
Frame = +2
Query: 32 TIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISLFLTIP 91
+ G+ Y R PS R+ ++++ +T ++L + DP+++ + +N+ + I
Sbjct: 38 SFGINYGQIARN----LPSHSRVAMLIRSMNVTRIKLYDADPNVLLAFSKSNVEFVIGIG 205
Query: 92 NYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGN-AFTDVYPESINNLLPAISNVH 150
N + + TN S A+ WI HVLP+ + KI I+VGN F + I NLLP++ NVH
Sbjct: 206 NEHLQNM-TNPSKAQNWIQHHVLPYLSQTKINCITVGNEVFNSNDTQLILNLLPSMQNVH 382
Query: 151 ISLRDLGI-RKISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLI 209
SL LG+ + I+V+T+ SF + + +PPSS +F+ + I P+++FL + SSF I
Sbjct: 383 NSLVKLGLDQVITVTTAHSF-NILDNSYPPSSGSFRS-DLIQYIQPIVEFLDEIKSSFHI 556
Query: 210 NLYPYNLYRLRP-EIPLGIALF 230
N YP+ Y+ P E+ L + F
Sbjct: 557 NAYPFFAYKDNPNEVSLTMHYF 622
>TC80333 similar to GP|9758115|dbj|BAB08587.1 beta-1 3-glucanase-like
protein {Arabidopsis thaliana}, partial (46%)
Length = 836
Score = 102 bits (255), Expect = 2e-22
Identities = 64/234 (27%), Positives = 120/234 (50%), Gaps = 1/234 (0%)
Frame = +2
Query: 3 HHSTFMISLTTFSLYLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLK 62
HH+ ++ FS + LL L +++ +IG+ Y P+P ++ ++
Sbjct: 32 HHN---MAFVAFSFFFLLLSLHASSSEAGSIGINYGRIADN----LPTPSKVVELLKAQG 190
Query: 63 LTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKI 122
++L + D +++ +L + I + + +PN L++ A ++S WI +++L YP +I
Sbjct: 191 FNRVKLYDTDATVLTALANSGIKVTVAMPNELLSSAAADQSYTDTWIQSNILNHYPATEI 370
Query: 123 TTISVGNAFTDVYPESINNLLPAISNVHISLRDLGIRKISVSTSFSFVTAVTSPFPPSSA 182
I+VGN + N L+PA+ NVH SL+ + K + +S ++A+ S +P S+
Sbjct: 371 EAIAVGNEVFVDPKNTTNYLVPAMKNVHASLQKQNLDKQILISSPIALSALQSSYPTSTG 550
Query: 183 AFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRPE-IPLGIALFQEHPF 235
+F+ +I P+L+FLS T S ++N YP+ Y + I L AL P+
Sbjct: 551 SFKTELVEPVIKPMLEFLSQTGSYLMVNAYPFFAYAANSDTISLDYALV*AKPW 712
>TC80654 similar to GP|3900936|emb|CAA10167.1 glucan endo-1
3-beta-d-glucosidase {Cicer arietinum}, partial (56%)
Length = 746
Score = 99.4 bits (246), Expect = 2e-21
Identities = 65/211 (30%), Positives = 115/211 (53%), Gaps = 5/211 (2%)
Frame = +3
Query: 143 LPAISNVHISLRDLGIR-KISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLS 201
LPA+ N+ ++ ++ +I VS + +T + + +PP++ F + I P++ FL
Sbjct: 9 LPAMQNIQNAISAANLQGQIKVSIAID-MTLIGNSYPPNNGVFTD-QAKPYIQPIINFLK 182
Query: 202 DTNSSFLINLYPYNLY-RLRPEIPLGIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVS 260
+ + L N+YPY Y + I L ALF++ N V Y+NLFD +D+V +
Sbjct: 183 NNGAPLLANVYPYFAYINNKQSISLDYALFRQQGNN-------QVGYRNLFDAQLDSVYA 341
Query: 261 AMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKGV 320
A+ G + ++V+E+GWPS+G + A++ A Y + L+ H+++ GTP + G
Sbjct: 342 ALEKVGASGVKIVVSESGWPSAGGD-SASTDNAATYYRNLINHVRN--GTP-KRPGAIET 509
Query: 321 YLYELFD---KEGAGNGRNWGILYPNGSTKY 348
YL+ +FD K GA +++G+ PN + KY
Sbjct: 510 YLFAMFDENQKTGAATEQHFGLFNPNRTPKY 602
>TC81506 weakly similar to GP|22748323|gb|AAN05325.1 Putative beta-1
3-glucanase {Oryza sativa (japonica cultivar-group)},
partial (39%)
Length = 850
Score = 97.8 bits (242), Expect = 6e-21
Identities = 70/239 (29%), Positives = 117/239 (48%), Gaps = 5/239 (2%)
Frame = +2
Query: 1 MFHHSTFMISLTTFSLYLLLSLLPTTTTSL---PTIGVTYSSTTRQDSPPPPSPDRITTA 57
+F F ++L+T +L+ LL L + + P IGV ++ P P ++
Sbjct: 143 VFFLGRFELNLSTMALHFLLMLFAVSLVAADDAPFIGVNIGTSLSD----MPHPTQVVAL 310
Query: 58 MQNLKLTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFY 117
++ K+ ++RL + D +++ +L T I + +T+PN + I + + A W+ +V+ Y
Sbjct: 311 LKAQKIQNVRLYDADQAMLVALAKTGIQVVITVPNEQILAIGQSNASAANWVSRNVVAHY 490
Query: 118 PRAKITTISVGNAFTDVYPESINNLLPAISNVHISLRDLGI-RKISVSTSFSFVTAVTSP 176
P IT I VG+ P L+ AI +H +L + R++ VST + +
Sbjct: 491 PATNITAICVGSEVLTTLPNVAKVLVNAIKYIHSALVASNLDRQVKVSTPLP-SSIILDS 667
Query: 177 FPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLY-RLRPEIPLGIALFQEHP 234
FPPS A F LI P+L FL T+S ++N+YPY Y + I L ALF+ P
Sbjct: 668 FPPSQAFFNRSLNSVLI-PILDFLQSTDSYLMLNIYPYYDYMQSNGVIXLXYALFKPLP 841
>TC77401 similar to GP|1247325|emb|CAA01814.1 beta-1 3-glucanase {Glycine
max}, partial (59%)
Length = 737
Score = 94.0 bits (232), Expect = 9e-20
Identities = 73/235 (31%), Positives = 115/235 (48%), Gaps = 6/235 (2%)
Frame = +2
Query: 2 FHHSTFMISLTTFSLYLLLSLLPTTTTSLPT--IGVTYSSTTRQDSPPPPSPDRITTAMQ 59
F S + T + LLL L T T L + IGV Y + PS +
Sbjct: 35 FFPSMITSNSTVMASLLLLFALFTVTLRLTSAQIGVCYG----MEGTNLPSQREAIDLCK 202
Query: 60 NLKLTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPR 119
+ + +RL +P+P + +L + I L L +PN + IA N+ IA W+ +VL FYP
Sbjct: 203 SNNIKRMRLYDPNPDALEALRNSGIELMLGVPNSDLQNIANNKDIANQWVQKNVLNFYPS 382
Query: 120 AKITTISVGNAFTDVYPES--INNLLPAISNVHISLRDLGIR-KISVSTSFSFVTAVTSP 176
KI I+VGN V S +LPAI N++ ++R + +I VST+ +T + +
Sbjct: 383 VKIKYIAVGNEVNPVGGSSQFAKFVLPAIQNIYQAIRAKNFQDQIKVSTAID-MTMIGTS 559
Query: 177 FPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLGIALF 230
+PPS +F+ + + P++ +L N+ N+Y Y Y+ P +I L ALF
Sbjct: 560 YPPSKGSFRS-DVRSYLDPIIGYLVYANAPLFANIYSYFSYKDNPKDISLQYALF 721
>TC90032 similar to GP|14090345|dbj|BAB55504. contains EST
D15166(C0193)~unknown protein {Oryza sativa (japonica
cultivar-group)}, partial (27%)
Length = 1286
Score = 87.0 bits (214), Expect = 1e-17
Identities = 54/144 (37%), Positives = 79/144 (54%), Gaps = 6/144 (4%)
Frame = +2
Query: 217 YRLRP-EIPLGIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVT 275
YR P ++ L ALF+ D TG+ Y N+FD +DA+ A+ + TI V+VT
Sbjct: 2 YRDSPTKVSLDYALFESSSEVI--DPNTGLLYTNMFDAQIDAIYFALTALNFRTIKVMVT 175
Query: 276 ETGWPSSGS--ELDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKGVYLYELFD---KEG 330
ETGWPS GS E A A+ Y L++H+ + GTP +Y++ LF+ K G
Sbjct: 176 ETGWPSKGSPKETAATPDNAQTYNTNLIRHVINETGTPAKPGEELDIYIFSLFNENRKPG 355
Query: 331 AGNGRNWGILYPNGSTKYDKIDFS 354
+ RNWGI+YP+ + Y +DF+
Sbjct: 356 LESERNWGIVYPDLTNVY-SLDFT 424
>TC91497 similar to PIR|T05960|T05960 beta-1 3-glucanase (EC 3.2.1.-) 7 -
soybean (fragment), partial (63%)
Length = 651
Score = 84.7 bits (208), Expect = 5e-17
Identities = 51/154 (33%), Positives = 91/154 (58%), Gaps = 2/154 (1%)
Frame = +2
Query: 33 IGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISLFLTIPN 92
+G+ Y D PPP D+++ +Q+ K+ ++R+ + + +++S T + L + IPN
Sbjct: 176 VGICYGRNA-DDLPPP---DKVSQLVQDHKIKYVRIYDSNIQVLKSFANTGVELMIGIPN 343
Query: 93 YLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTDVYPESINNL-LPAISNVHI 151
+ P + ++ A W+ +LP+YP KIT I+VG T+ PE+I+ L +PA++NV
Sbjct: 344 LDLLPFSQFQTNADTWLRNSILPYYPATKITYITVGAEVTE-SPENISALVVPAMTNVLA 520
Query: 152 SLRDLGI-RKISVSTSFSFVTAVTSPFPPSSAAF 184
+L+ G+ +KI VS++ S + + FPPS AF
Sbjct: 521 ALKKAGLHKKIKVSSTHS-LGVLXRSFPPSPGAF 619
>AL370013 similar to PIR|T05268|T05 hypothetical protein T4L20.60 -
Arabidopsis thaliana (fragment), partial (39%)
Length = 461
Score = 82.0 bits (201), Expect = 3e-16
Identities = 49/130 (37%), Positives = 71/130 (53%), Gaps = 5/130 (3%)
Frame = +2
Query: 225 LGIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGS 284
L LFQ P R D T + Y N+FD VDAV SA+ G++ + ++V ETGWP G
Sbjct: 38 LAFCLFQ--PNAGRVDANTKLNYMNMFDAQVDAVRSALDSMGFKDVEIVVAETGWPYKGD 211
Query: 285 ELDANSAY--AEIYLKGLVKHLKSGAGTPLLKDGVKGVYLYELFD---KEGAGNGRNWGI 339
+A + A+ Y L+KHL+S GTPL+ Y++ L+D K GAG+ + +G+
Sbjct: 212 NDEAGPSIENAKAYNGNLIKHLRSKVGTPLMPGKSVDTYIFALYDEDLKPGAGSEKAFGL 391
Query: 340 LYPNGSTKYD 349
+ S YD
Sbjct: 392 YNTDQSMIYD 421
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.139 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,175,092
Number of Sequences: 36976
Number of extensions: 192401
Number of successful extensions: 1970
Number of sequences better than 10.0: 96
Number of HSP's better than 10.0 without gapping: 1762
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1908
length of query: 376
length of database: 9,014,727
effective HSP length: 98
effective length of query: 278
effective length of database: 5,391,079
effective search space: 1498719962
effective search space used: 1498719962
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC147434.3


