

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147430.8 + phase: 1 /pseudo
(437 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
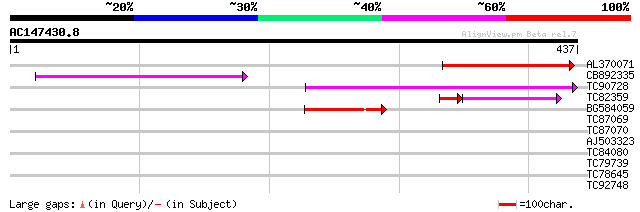
Score E
Sequences producing significant alignments: (bits) Value
AL370071 similar to GP|15795135|dbj transposase-like protein {Ar... 122 2e-28
CB892335 similar to GP|7715599|gb| F20B17.17 {Arabidopsis thalia... 114 7e-26
TC90728 similar to GP|7715599|gb|AAF68117.1| F20B17.17 {Arabidop... 77 1e-14
TC82359 weakly similar to GP|22655127|gb|AAM98154.1 putative pro... 62 4e-14
BG584059 similar to GP|12722947|gb| UNKNOWN PROTEIN {Lactococcus... 74 1e-13
TC87069 weakly similar to GP|9758877|dbj|BAB09431.1 contains sim... 31 0.82
TC87070 weakly similar to GP|9965105|gb|AAG09952.1| resistance p... 31 0.82
AJ503323 weakly similar to PIR|S51391|S513 probable membrane pro... 31 0.82
TC84080 similar to GP|15982773|gb|AAL09734.1 At1g36320/F7F23_4 {... 30 1.8
TC79739 homologue to PIR|G86423|G86423 probable hydrophilic prot... 28 5.3
TC78645 similar to GP|19310705|gb|AAL85083.1 unknown protein {Ar... 28 9.1
TC92748 similar to SP|P53393|SUT3_STYHA Low affinity sulphate tr... 28 9.1
>AL370071 similar to GP|15795135|dbj transposase-like protein {Arabidopsis
thaliana}, partial (3%)
Length = 370
Score = 122 bits (307), Expect = 2e-28
Identities = 56/102 (54%), Positives = 71/102 (68%)
Frame = +3
Query: 334 IIDDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCITRMVEDPEEQAKIEV 393
IID+RWD+ H PLHAA Y+LN Q HY PGF+ D++V++GL CITRMV D EE++KIE+
Sbjct: 63 IIDERWDKQFHSPLHAAGYFLNAQYHYSPGFRDDVKVKRGLQHCITRMVTDHEERSKIEI 242
Query: 394 QMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLELQR 435
Q+ DF KQ FG IA +T D P WW S+V ELQ+
Sbjct: 243 QLDDFDKQANQFGHPIAIITADMEIPPIWWGSLVDGPPELQK 368
>CB892335 similar to GP|7715599|gb| F20B17.17 {Arabidopsis thaliana}, partial
(27%)
Length = 788
Score = 114 bits (285), Expect = 7e-26
Identities = 56/163 (34%), Positives = 95/163 (57%)
Frame = +2
Query: 21 IQRSKEFNIMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIM 80
+ RS + +M D I++ G GF PS ++ L + ++ ++EW T CTI+
Sbjct: 299 VARSSSYQLMIDAITKCGPGFTGPSAEILKTIWLERIKSEVGLQSKDVEKEWATTGCTII 478
Query: 81 TDEWTDRRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQV 140
D WTD + + I+NFLV+SP F KSVDAS K ++ + ++ D+V++E G +NVVQ+
Sbjct: 479 ADTWTDYKSKAIINFLVSSPSRIFFHKSVDASAYFKNTKWLADLFDSVIQEFGPENVVQI 658
Query: 141 VTDNAANYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKI 183
+ D++ NY G +++ ++ +PCA+ L+L+ D K I
Sbjct: 659 IMDSSFNYTGIGNHIVQNYGTIFVSPCASQWLNLIFGDSPKMI 787
>TC90728 similar to GP|7715599|gb|AAF68117.1| F20B17.17 {Arabidopsis
thaliana}, partial (58%)
Length = 1171
Score = 77.4 bits (189), Expect = 1e-14
Identities = 56/211 (26%), Positives = 93/211 (43%), Gaps = 2/211 (0%)
Frame = +3
Query: 229 TSYLTLGCLYENKEALIRMFTSKEWK-SGKCAKLRDGKALEDIILDKKFWKDIVICLKGA 287
+++L+L + + + L MF S E+ A + I D FW+ + C+ +
Sbjct: 12 STFLSLQTMLKLRTRLKHMFHSPEYALDTSYANKPQSLSCIAIAEDGDFWRTVEECVAIS 191
Query: 288 TPLMKVLRLADLDKKTSHGIHL*SNGSSKGANSKEF*WCQK-EPVWNIIDDRWDRMLHRP 346
P +KVLR K T I+ + + + K + +I+D +W LH P
Sbjct: 192 EPFLKVLREVSEGKPTVGSIYELMTRAKESIRTYYIMDENKCKTFLDIVDKKWRDQLHSP 371
Query: 347 LHAAAYYLNPQLHYRPGFKADIEVRKGLMDCITRMVEDPEEQAKIEVQMHDFKKQIGYFG 406
LHAAA +LNP + Y P K +++ + +++ P+ + I Q++ F K G FG
Sbjct: 372 LHAAAAFLNPSIQYNPEIKFLSSIKEDFYHVLEKLLPVPDMRRDITNQIYTFTKAHGMFG 551
Query: 407 TKIAKLTLDKSTPADWWESVVFEYLELQRFA 437
+ K + P WWE LQR A
Sbjct: 552 CSLTKEARNTVAPWLWWEQYGDSAPGLQRVA 644
>TC82359 weakly similar to GP|22655127|gb|AAM98154.1 putative protein
{Arabidopsis thaliana}, partial (14%)
Length = 907
Score = 61.6 bits (148), Expect(2) = 4e-14
Identities = 28/76 (36%), Positives = 42/76 (54%)
Frame = +3
Query: 350 AAYYLNPQLHYRPGFKADIEVRKGLMDCITRMVEDPEEQAKIEVQMHDFKKQIGYFGTKI 409
A +YLNP+ Y E+R G++DCI R+V D Q KI +++ +K G FG K+
Sbjct: 57 AGFYLNPKFFYSIQGDVPNEIRSGMLDCIERLVPDTRVQDKISKELNLYKSAAGDFGRKM 236
Query: 410 AKLTLDKSTPADWWES 425
A D P++WW +
Sbjct: 237 AIRARDNLLPSEWWST 284
Score = 33.9 bits (76), Expect(2) = 4e-14
Identities = 12/18 (66%), Positives = 13/18 (71%)
Frame = +2
Query: 332 WNIIDDRWDRMLHRPLHA 349
WNII RWD + H PLHA
Sbjct: 2 WNIIHQRWDSLWHHPLHA 55
>BG584059 similar to GP|12722947|gb| UNKNOWN PROTEIN {Lactococcus lactis
subsp. lactis}, partial (9%)
Length = 838
Score = 73.6 bits (179), Expect = 1e-13
Identities = 35/63 (55%), Positives = 46/63 (72%)
Frame = -1
Query: 228 ATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKALEDIILDKKFWKDIVICLKGA 287
AT+YLT CL++N ALI MFT KEWKSGK AK +DG +E + D FWK+++IC++G
Sbjct: 796 ATTYLT*ECLHDNYRALIIMFTLKEWKSGKLAKSKDGIIVEKAVKD-NFWKNVMICMRGV 620
Query: 288 TPL 290
PL
Sbjct: 619 FPL 611
>TC87069 weakly similar to GP|9758877|dbj|BAB09431.1 contains similarity to
disease resistance protein~gene_id:K24G6.11 {Arabidopsis
thaliana}, partial (4%)
Length = 1310
Score = 31.2 bits (69), Expect = 0.82
Identities = 16/39 (41%), Positives = 22/39 (56%)
Frame = +3
Query: 37 HGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKT 75
H N F P +Y+I + +E S ALE H+R+ KKT
Sbjct: 291 HNNQFIFPVFYNIEPCTVRKETGSYGMALENHERDNKKT 407
>TC87070 weakly similar to GP|9965105|gb|AAG09952.1| resistance protein LM17
{Glycine max}, partial (4%)
Length = 731
Score = 31.2 bits (69), Expect = 0.82
Identities = 16/39 (41%), Positives = 22/39 (56%)
Frame = +1
Query: 37 HGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKT 75
H N F P +Y+I + +E S ALE H+R+ KKT
Sbjct: 292 HNNQFIFPVFYNIEPCTVRKETGSYGMALENHERDNKKT 408
>AJ503323 weakly similar to PIR|S51391|S513 probable membrane protein YLR373c
- yeast (Saccharomyces cerevisiae), partial (2%)
Length = 512
Score = 31.2 bits (69), Expect = 0.82
Identities = 16/39 (41%), Positives = 22/39 (56%)
Frame = +1
Query: 37 HGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKT 75
H N F P +Y+I + +E S ALE H+R+ KKT
Sbjct: 298 HNNQFIFPVFYNIEPCTVRKETGSYGMALENHERDNKKT 414
>TC84080 similar to GP|15982773|gb|AAL09734.1 At1g36320/F7F23_4 {Arabidopsis
thaliana}, partial (27%)
Length = 594
Score = 30.0 bits (66), Expect = 1.8
Identities = 20/58 (34%), Positives = 25/58 (42%)
Frame = +1
Query: 371 RKGLMDCITRMVEDPEEQAKIEVQMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVF 428
R+G C V E+ A +V D K IG I +DKSTP DW + F
Sbjct: 190 RRGFRPC--NYVASNEDNASSDVV--DESKMIGVCDKLIGVFMVDKSTPTDWRRLLAF 351
>TC79739 homologue to PIR|G86423|G86423 probable hydrophilic protein
29542-30030 [imported] - Arabidopsis thaliana, partial
(95%)
Length = 676
Score = 28.5 bits (62), Expect = 5.3
Identities = 24/99 (24%), Positives = 47/99 (47%), Gaps = 13/99 (13%)
Frame = +2
Query: 107 KSVDASNICKTSEKIFEMIDAVVEEVGEDNVV----QVVTDNAANYKAAGEMLMRK---- 158
K+ D + + K K ++ ++GE+ +V ++ ++A YK G +L+++
Sbjct: 92 KANDLNKLQKDIAKNHQVRKKYTIQLGENELVLKELDLLNEDANVYKLIGPVLVKQDLAE 271
Query: 159 -----RKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPK 192
RK++ + LD ++DLE+K ETI K
Sbjct: 272 ANANVRKRIEYISAELKRLDATVQDLEEKQNSKKETIMK 388
>TC78645 similar to GP|19310705|gb|AAL85083.1 unknown protein {Arabidopsis
thaliana}, partial (70%)
Length = 1101
Score = 27.7 bits (60), Expect = 9.1
Identities = 15/63 (23%), Positives = 33/63 (51%)
Frame = +1
Query: 247 MFTSKEWKSGKCAKLRDGKALEDIILDKKFWKDIVICLKGATPLMKVLRLADLDKKTSHG 306
+F+ K+ CA++ +ED++ + + W+D+ L+K+ +L D+ + TS
Sbjct: 85 LFSCLSSKAFLCAEILKTGVMEDVVSNTRRWEDL-----DTDILVKIFQLLDIFELTSGI 249
Query: 307 IHL 309
H+
Sbjct: 250 AHV 258
>TC92748 similar to SP|P53393|SUT3_STYHA Low affinity sulphate transporter
3. {Stylosanthes hamata}, partial (20%)
Length = 536
Score = 27.7 bits (60), Expect = 9.1
Identities = 11/25 (44%), Positives = 16/25 (64%)
Frame = +2
Query: 155 LMRKRKKLYWTPCAAHCLDLMLEDL 179
+ RK+KKL+W P A L ++L L
Sbjct: 131 IARKKKKLFWLPAIAPLLSVILSTL 205
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.138 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,936,677
Number of Sequences: 36976
Number of extensions: 187083
Number of successful extensions: 1219
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 1213
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1217
length of query: 437
length of database: 9,014,727
effective HSP length: 99
effective length of query: 338
effective length of database: 5,354,103
effective search space: 1809686814
effective search space used: 1809686814
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC147430.8


