

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147429.11 - phase: 0
(149 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
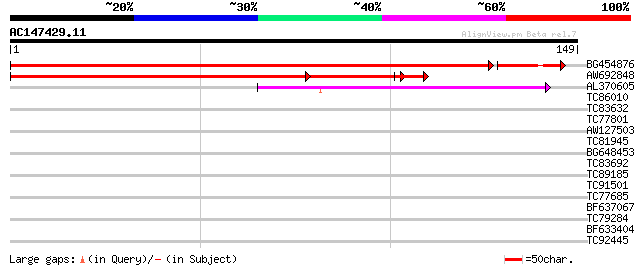
Score E
Sequences producing significant alignments: (bits) Value
BG454876 234 3e-65
AW692848 157 5e-49
AL370605 weakly similar to SP|P27061|PPA1 Acid phosphatase precu... 40 3e-04
TC86010 homologue to PIR|T00795|T00795 26S proteasome regulatory... 32 0.13
TC83632 similar to PIR|B86248|B86248 protein T23J18.11 [imported... 31 0.21
TC77801 weakly similar to GP|21536807|gb|AAM61139.1 unknown {Ara... 30 0.36
AW127503 27 3.1
TC81945 weakly similar to GP|10177640|dbj|BAB10787. contains sim... 27 3.1
BG648453 similar to GP|15293215|gb putative oligopeptide transpo... 26 5.3
TC83692 weakly similar to GP|8809708|dbj|BAA97249.1 gene_id:MQM1... 26 5.3
TC89185 weakly similar to GP|13605746|gb|AAK32866.1 AT4g31930/F1... 26 5.3
TC91501 N7 protein 26 6.9
TC77685 apyrase-like protein [Medicago truncatula] 26 6.9
BF637067 weakly similar to GP|21618058|gb| unknown {Arabidopsis ... 25 9.0
TC79284 similar to GP|10178077|dbj|BAB11496. gene_id:K19B1.7~unk... 25 9.0
BF633404 similar to GP|11994779|dbj gene_id:T19N8.1~pir||B71428~... 25 9.0
TC92445 weakly similar to PIR|T01004|T01004 hypothetical protein... 25 9.0
>BG454876
Length = 692
Score = 234 bits (598), Expect(2) = 3e-65
Identities = 115/127 (90%), Positives = 120/127 (93%)
Frame = +1
Query: 1 MGSSFVLESGFYITSFSATIFIAGFAALGLLLVTLLVSMAMMLQSCQNSNGGIIELRNIN 60
MG +F LESGFYITSFSATIFIAGFA LGLLLVTLLVSMAMMLQSCQNSNGG+IELRNIN
Sbjct: 187 MGXNFXLESGFYITSFSATIFIAGFATLGLLLVTLLVSMAMMLQSCQNSNGGVIELRNIN 366
Query: 61 NDYSYCKIHSLHAKLNNLEEHNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKP 120
+DYSYCKIHSLHAKLNNLEE+NVPNICKDLALQYIKGGQY DLDSTKSVIEDYFNGVKP
Sbjct: 367 DDYSYCKIHSLHAKLNNLEEYNVPNICKDLALQYIKGGQYVXDLDSTKSVIEDYFNGVKP 546
Query: 121 SEDGFDV 127
S+ G DV
Sbjct: 547 SKXGLDV 567
Score = 30.0 bits (66), Expect(2) = 3e-65
Identities = 14/18 (77%), Positives = 16/18 (88%)
Frame = +3
Query: 129 LIDIDSLFQWNPPHSSNL 146
+I+I SLFQWN PHSSNL
Sbjct: 570 VINIXSLFQWN-PHSSNL 620
>AW692848
Length = 611
Score = 157 bits (396), Expect(3) = 5e-49
Identities = 79/79 (100%), Positives = 79/79 (100%)
Frame = +2
Query: 1 MGSSFVLESGFYITSFSATIFIAGFAALGLLLVTLLVSMAMMLQSCQNSNGGIIELRNIN 60
MGSSFVLESGFYITSFSATIFIAGFAALGLLLVTLLVSMAMMLQSCQNSNGGIIELRNIN
Sbjct: 257 MGSSFVLESGFYITSFSATIFIAGFAALGLLLVTLLVSMAMMLQSCQNSNGGIIELRNIN 436
Query: 61 NDYSYCKIHSLHAKLNNLE 79
NDYSYCKIHSLHAKLNNLE
Sbjct: 437 NDYSYCKIHSLHAKLNNLE 493
Score = 53.1 bits (126), Expect(3) = 5e-49
Identities = 23/26 (88%), Positives = 25/26 (95%)
Frame = +3
Query: 79 EEHNVPNICKDLALQYIKGGQYARDL 104
+EHNVPNICKDLALQYIKGGQYA+ L
Sbjct: 492 KEHNVPNICKDLALQYIKGGQYAKRL 569
Score = 21.2 bits (43), Expect(3) = 5e-49
Identities = 9/9 (100%), Positives = 9/9 (100%)
Frame = +1
Query: 102 RDLDSTKSV 110
RDLDSTKSV
Sbjct: 562 RDLDSTKSV 588
>AL370605 weakly similar to SP|P27061|PPA1 Acid phosphatase precursor 1 (EC
3.1.3.2). [Tomato] {Lycopersicon esculentum}, partial
(42%)
Length = 472
Score = 40.4 bits (93), Expect = 3e-04
Identities = 23/78 (29%), Positives = 33/78 (41%), Gaps = 1/78 (1%)
Frame = +2
Query: 66 CKIHSLHAKLNNLEE-HNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSEDG 124
C+ + NNL VP C + +Y+ G Y DL+ ++ VK EDG
Sbjct: 167 CRSWRFAGEANNLSPWKTVPKECAEHVKEYMNGKGYVYDLEIANKEAGEFAKSVKLKEDG 346
Query: 125 FDVVLIDIDSLFQWNPPH 142
D + DID N P+
Sbjct: 347 LDAWVFDIDETLLSNLPY 400
>TC86010 homologue to PIR|T00795|T00795 26S proteasome regulatory subunit
[imported] - Arabidopsis thaliana, partial (35%)
Length = 1865
Score = 31.6 bits (70), Expect = 0.13
Identities = 15/51 (29%), Positives = 27/51 (52%)
Frame = -2
Query: 33 VTLLVSMAMMLQSCQNSNGGIIELRNINNDYSYCKIHSLHAKLNNLEEHNV 83
+ +LV + S + SN I N+N+D+S+C +L K+NN+ +
Sbjct: 433 ILMLVKDNFLQLSPEGSNTRITSFTNLNHDHSHCYEGTLTNKINNIRSQRL 281
>TC83632 similar to PIR|B86248|B86248 protein T23J18.11 [imported] -
Arabidopsis thaliana, partial (11%)
Length = 654
Score = 30.8 bits (68), Expect = 0.21
Identities = 19/71 (26%), Positives = 30/71 (41%)
Frame = +2
Query: 30 LLLVTLLVSMAMMLQSCQNSNGGIIELRNINNDYSYCKIHSLHAKLNNLEEHNVPNICKD 89
+ + LL SMA SC S+ G + +Y Y ++ S N ++N PN D
Sbjct: 158 VFIFNLL*SMAYYYSSCSGSDYGENNFNSYTFNYDYAQVPSF--MTYNSYDYNQPNYAYD 331
Query: 90 LALQYIKGGQY 100
+L + Y
Sbjct: 332 SSLYFAPSNNY 364
>TC77801 weakly similar to GP|21536807|gb|AAM61139.1 unknown {Arabidopsis
thaliana}, partial (12%)
Length = 1145
Score = 30.0 bits (66), Expect = 0.36
Identities = 22/79 (27%), Positives = 39/79 (48%), Gaps = 3/79 (3%)
Frame = +2
Query: 53 IIELRNINNDYSYCKIHSLHAKLNNLEEHNVPNICKDLALQYIKGGQYARDLDSTKSVIE 112
I+EL+N ++D S I S H +N ++ HN+PN ++ + R+ D + +
Sbjct: 314 IVELKNSDSDSSI--IRSDHFSINPIQIHNIPN---SISSSCSSNSEIDRNFDDSYVANQ 478
Query: 113 DYFNGVK---PSEDGFDVV 128
Y N ++ S+ DVV
Sbjct: 479 IYPNNLRRWNDSDSDSDVV 535
>AW127503
Length = 440
Score = 26.9 bits (58), Expect = 3.1
Identities = 14/46 (30%), Positives = 22/46 (47%), Gaps = 2/46 (4%)
Frame = +2
Query: 52 GIIELRNINNDYSYCKIHSLHAKLNNLEEHN--VPNICKDLALQYI 95
G + ++N+ Y Y IH+ H + N E N P +C L + I
Sbjct: 128 GNVSFGHLNHGYQYLTIHASHISIKN*SEPNRYEPLVCIYLDMNLI 265
>TC81945 weakly similar to GP|10177640|dbj|BAB10787. contains similarity to
heparanase~gene_id:MGG23.2 {Arabidopsis thaliana},
partial (38%)
Length = 1031
Score = 26.9 bits (58), Expect = 3.1
Identities = 10/26 (38%), Positives = 16/26 (61%)
Frame = +1
Query: 95 IKGGQYARDLDSTKSVIEDYFNGVKP 120
+ QYA D+ + +I+D + GVKP
Sbjct: 856 VAADQYASDVIVLRKIIQDVYRGVKP 933
>BG648453 similar to GP|15293215|gb putative oligopeptide transporter protein
{Arabidopsis thaliana}, partial (29%)
Length = 701
Score = 26.2 bits (56), Expect = 5.3
Identities = 12/36 (33%), Positives = 19/36 (52%)
Frame = +1
Query: 27 ALGLLLVTLLVSMAMMLQSCQNSNGGIIELRNINND 62
+ G L +LLVS+ + S + GG + N+N D
Sbjct: 334 SFGFYLSSLLVSLVNKITSTSSGGGGWLHDNNLNKD 441
>TC83692 weakly similar to GP|8809708|dbj|BAA97249.1 gene_id:MQM1.22~unknown
protein {Arabidopsis thaliana}, partial (25%)
Length = 819
Score = 26.2 bits (56), Expect = 5.3
Identities = 12/31 (38%), Positives = 19/31 (60%), Gaps = 2/31 (6%)
Frame = +3
Query: 53 IIELRNINNDYSYCKIHSLHA--KLNNLEEH 81
I+ ++N +ND S +H LHA + L+EH
Sbjct: 255 ILSIKNSDNDISRLDLHGLHAXXAVQALQEH 347
>TC89185 weakly similar to GP|13605746|gb|AAK32866.1 AT4g31930/F10N7_260
{Arabidopsis thaliana}, partial (63%)
Length = 1181
Score = 26.2 bits (56), Expect = 5.3
Identities = 15/33 (45%), Positives = 18/33 (54%)
Frame = -2
Query: 49 SNGGIIELRNINNDYSYCKIHSLHAKLNNLEEH 81
SNG IE N ND ++C L AKL +L H
Sbjct: 973 SNGFSIEETNSYNDKTWCSHQLLVAKLTSLFFH 875
>TC91501 N7 protein
Length = 1199
Score = 25.8 bits (55), Expect = 6.9
Identities = 11/27 (40%), Positives = 16/27 (58%)
Frame = +2
Query: 69 HSLHAKLNNLEEHNVPNICKDLALQYI 95
++L NLEE N+ +C D L+YI
Sbjct: 224 YALERSCGNLEEINIECLCSDGLLKYI 304
>TC77685 apyrase-like protein [Medicago truncatula]
Length = 2172
Score = 25.8 bits (55), Expect = 6.9
Identities = 10/28 (35%), Positives = 16/28 (56%)
Frame = +1
Query: 67 KIHSLHAKLNNLEEHNVPNICKDLALQY 94
K+ + + +EE N+P +C DL QY
Sbjct: 1531 KLENAKSTYPRVEEGNLPYLCMDLVYQY 1614
>BF637067 weakly similar to GP|21618058|gb| unknown {Arabidopsis thaliana},
partial (25%)
Length = 610
Score = 25.4 bits (54), Expect = 9.0
Identities = 13/34 (38%), Positives = 18/34 (52%)
Frame = +3
Query: 30 LLLVTLLVSMAMMLQSCQNSNGGIIELRNINNDY 63
LL VT + MLQ C++S +RN N+ Y
Sbjct: 150 LLYVTSAAAGYNMLQLCKHSFSACSTIRNFNDSY 251
>TC79284 similar to GP|10178077|dbj|BAB11496. gene_id:K19B1.7~unknown
protein {Arabidopsis thaliana}, partial (56%)
Length = 1009
Score = 25.4 bits (54), Expect = 9.0
Identities = 16/59 (27%), Positives = 31/59 (52%)
Frame = -2
Query: 67 KIHSLHAKLNNLEEHNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSEDGF 125
+I L +L +E+H P +C+ ++ ++ + +D+D V+E Y N V + D F
Sbjct: 561 RIDVLPLQLPTVEDHASPKVCQRVSSLHLYRWWFLQDVD---EVVEQY-NQVHRTYDKF 397
>BF633404 similar to GP|11994779|dbj gene_id:T19N8.1~pir||B71428~similar to
unknown protein {Arabidopsis thaliana}, partial (8%)
Length = 325
Score = 25.4 bits (54), Expect = 9.0
Identities = 9/27 (33%), Positives = 18/27 (66%)
Frame = +1
Query: 106 STKSVIEDYFNGVKPSEDGFDVVLIDI 132
S +I+++FN + P +GF + LI++
Sbjct: 13 SIDKIIQNHFNTLSPEPNGFHLYLINL 93
>TC92445 weakly similar to PIR|T01004|T01004 hypothetical protein At2g39740
[imported] - Arabidopsis thaliana, partial (16%)
Length = 1141
Score = 25.4 bits (54), Expect = 9.0
Identities = 13/32 (40%), Positives = 19/32 (58%), Gaps = 2/32 (6%)
Frame = +1
Query: 85 NICKDLALQYIKGGQ--YARDLDSTKSVIEDY 114
+IC+DL L YIK + + D ST + +E Y
Sbjct: 439 SICRDLQLTYIKKSESKLSSDTVSTTTCVEPY 534
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.138 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,846,004
Number of Sequences: 36976
Number of extensions: 62919
Number of successful extensions: 379
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 378
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 379
length of query: 149
length of database: 9,014,727
effective HSP length: 87
effective length of query: 62
effective length of database: 5,797,815
effective search space: 359464530
effective search space used: 359464530
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC147429.11


