

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
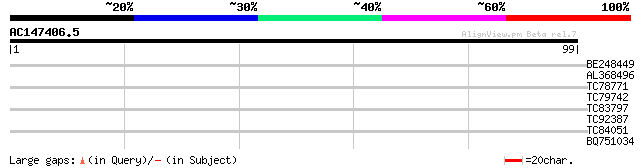
Query= AC147406.5 - phase: 0 /pseudo
(99 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BE248449 similar to GP|12330687|gb 2-isopropylmalate synthase {A... 31 0.068
AL368496 similar to GP|16612268|gb| AT3g61860/F21F14_30 {Arabido... 29 0.26
TC78771 similar to GP|16612268|gb|AAL27502.1 AT3g61860/F21F14_30... 29 0.26
TC79742 similar to GP|10177241|dbj|BAB10615. gb|AAC98455.1~gene_... 25 4.9
TC83797 similar to GP|21618159|gb|AAM67209.1 probable riboflavin... 25 6.4
TC92387 25 6.4
TC84051 24 8.3
BQ751034 similar to GP|18307453|e probable DNA-directed RNA poly... 24 8.3
>BE248449 similar to GP|12330687|gb 2-isopropylmalate synthase {Arabidopsis
thaliana}, partial (18%)
Length = 344
Score = 31.2 bits (69), Expect = 0.068
Identities = 18/50 (36%), Positives = 29/50 (58%)
Frame = +2
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVIT 82
+LGK G HAMR+RL++ G+E+ D+ +F A+++K IT
Sbjct: 56 VLGKLSGRHAMRKRLEEL-----GYELKDDQVESLFWRFKAVAEQKKRIT 190
>AL368496 similar to GP|16612268|gb| AT3g61860/F21F14_30 {Arabidopsis
thaliana}, partial (36%)
Length = 616
Score = 29.3 bits (64), Expect = 0.26
Identities = 15/55 (27%), Positives = 26/55 (47%)
Frame = -2
Query: 37 NLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWLIFDH 91
NL Y + L++ + G E +D +GF V +++ D E+ I + F H
Sbjct: 585 NLEYDTRQSELERLFSKYGRIERVDMKSGFAFVYYEDERDAEEAIRALDNIPFGH 421
>TC78771 similar to GP|16612268|gb|AAL27502.1 AT3g61860/F21F14_30
{Arabidopsis thaliana}, partial (87%)
Length = 1254
Score = 29.3 bits (64), Expect = 0.26
Identities = 15/55 (27%), Positives = 26/55 (47%)
Frame = +1
Query: 37 NLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWLIFDH 91
NL Y + L++ + G E +D +GF V +++ D E+ I + F H
Sbjct: 109 NLEYDTRQSELERLFSKYGRIERVDMKSGFAFVYYEDERDAEEAIRALDNIPFGH 273
>TC79742 similar to GP|10177241|dbj|BAB10615.
gb|AAC98455.1~gene_id:MRN17.17~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (32%)
Length = 1063
Score = 25.0 bits (53), Expect = 4.9
Identities = 13/56 (23%), Positives = 30/56 (53%), Gaps = 6/56 (10%)
Frame = +2
Query: 1 MRMEIVFYPRFI---SNNQSFNNYVRHGRMQFVVKLLGKNLGYH---AMRERLQKT 50
+ + + F+ ++ +NN + NN ++H ++ +K L ++L + + LQKT
Sbjct: 317 LSLSLYFFTSYLISNNNNHNHNNNIKHHQITSTIKSLSQSLSIKKTTSSPQTLQKT 484
>TC83797 similar to GP|21618159|gb|AAM67209.1 probable riboflavin
biosynthesis related protein {Arabidopsis thaliana},
partial (44%)
Length = 842
Score = 24.6 bits (52), Expect = 6.4
Identities = 16/37 (43%), Positives = 19/37 (51%)
Frame = +2
Query: 61 DNDNGFYMVKFDEAADKEKVITGGPWLIFDHCLAVTH 97
DND+GFYM K E A K T L+ C+ V H
Sbjct: 137 DNDDGFYMRKSVELARKGLGYTSPNPLV--GCVIVKH 241
>TC92387
Length = 713
Score = 24.6 bits (52), Expect = 6.4
Identities = 16/75 (21%), Positives = 31/75 (41%), Gaps = 1/75 (1%)
Frame = +2
Query: 13 SNNQSFNNYVRHGRMQFVVKLLGKNLG-YHAMRERLQKTWRTQGGFEIMDNDNGFYMVKF 71
S S+ + R++ V +LL + M ++++ W G +I +G Y F
Sbjct: 155 SRRISYTTIIACPRLRLVGRLLTNRPN*SYVMIKKIEAFWSLVKGVKIQKI*SGLYTF*F 334
Query: 72 DEAADKEKVITGGPW 86
D + ++ PW
Sbjct: 335 HHHLDLQSILKKCPW 379
>TC84051
Length = 818
Score = 24.3 bits (51), Expect = 8.3
Identities = 13/35 (37%), Positives = 16/35 (45%)
Frame = -1
Query: 22 VRHGRMQFVVKLLGKNLGYHAMRERLQKTWRTQGG 56
VR QFVV G+ Y R R + W+ Q G
Sbjct: 764 VRPHLRQFVVPASGRGTMYQQRRPRPRCAWKPQRG 660
>BQ751034 similar to GP|18307453|e probable DNA-directed RNA polymerase I
{Neurospora crassa}, partial (13%)
Length = 791
Score = 24.3 bits (51), Expect = 8.3
Identities = 10/20 (50%), Positives = 12/20 (60%)
Frame = +3
Query: 23 RHGRMQFVVKLLGKNLGYHA 42
RHGR VV+ G G+HA
Sbjct: 504 RHGRADCVVRFHGPRRGWHA 563
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.327 0.141 0.460
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,165,918
Number of Sequences: 36976
Number of extensions: 33235
Number of successful extensions: 222
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 222
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 222
length of query: 99
length of database: 9,014,727
effective HSP length: 75
effective length of query: 24
effective length of database: 6,241,527
effective search space: 149796648
effective search space used: 149796648
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 50 (23.9 bits)
Medicago: description of AC147406.5


