

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
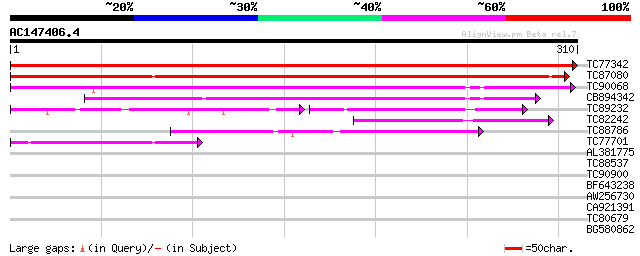
Query= AC147406.4 - phase: 0
(310 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC77342 similar to GP|13785211|emb|CAC37357. putative membrane p... 615 e-177
TC87080 similar to GP|13785209|emb|CAC37356. putative membrane p... 433 e-122
TC90068 similar to GP|20197988|gb|AAD22321.2 expressed protein {... 191 2e-49
CB894342 similar to GP|20197988|gb expressed protein {Arabidopsi... 170 7e-43
TC89232 similar to PIR|T47967|T47967 hypothetical protein F15G16... 65 6e-24
TC82242 weakly similar to GP|21592781|gb|AAM64730.1 putative mem... 84 9e-17
TC88786 weakly similar to GP|7268619|emb|CAB78828.1 unknown prot... 59 2e-09
TC77701 similar to PIR|T47789|T47789 hypothetical protein F17J16... 54 6e-08
AL381775 32 0.41
TC88537 weakly similar to GP|21536818|gb|AAM61150.1 F-box protei... 30 1.5
TC90900 weakly similar to PIR|D96753|D96753 Similar to disease r... 29 2.0
BF643238 similar to PIR|T51383|T51 receptor protein kinase-like ... 28 3.4
AW256730 weakly similar to GP|9757922|dbj respiratory burst oxid... 28 5.9
CA921391 similar to GP|1669591|dbj O-methyltransferase {Glycyrrh... 27 7.7
TC80679 phosphatase-like protein Mtc923 [Medicago truncatula] 27 7.7
BG580862 27 7.7
>TC77342 similar to GP|13785211|emb|CAC37357. putative membrane protein
{Solanum tuberosum}, partial (47%)
Length = 1581
Score = 615 bits (1587), Expect = e-177
Identities = 305/310 (98%), Positives = 308/310 (98%)
Frame = +2
Query: 1 MIGAQALVAIPQASGSPKAYTSNIVDTSTRLQEGTISYPVSGLSATYQNNKVTIFATLTL 60
MIGAQALVAIPQASGSPKAYTSNI+DTSTRLQEGTISYPVSGLSATYQNN+VTIFATLTL
Sbjct: 380 MIGAQALVAIPQASGSPKAYTSNIIDTSTRLQEGTISYPVSGLSATYQNNEVTIFATLTL 559
Query: 61 PNGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIGSRQRRRNT 120
PNGTTS VHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIGSRQRRRNT
Sbjct: 560 PNGTTSFVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIGSRQRRRNT 739
Query: 121 HGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSAYIVGLSGFGTGLKLGS 180
HGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSAYIVGLSGFGTGLKLGS
Sbjct: 740 HGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSAYIVGLSGFGTGLKLGS 919
Query: 181 DSEGITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNIYHHVVGYVTISISIVN 240
DS GITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNIYHHVVGYVTISISIVN
Sbjct: 920 DSVGITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNIYHHVVGYVTISISIVN 1099
Query: 241 VFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLEAYTWMVCMKRKKADNKTSDGVN 300
VFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLEAYTWMVCMKRKKA+NKTSDGVN
Sbjct: 1100VFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLEAYTWMVCMKRKKAENKTSDGVN 1279
Query: 301 GANGHGSSTL 310
GANGHGSSTL
Sbjct: 1280GANGHGSSTL 1309
>TC87080 similar to GP|13785209|emb|CAC37356. putative membrane protein
{Solanum tuberosum}, partial (47%)
Length = 1483
Score = 433 bits (1113), Expect = e-122
Identities = 214/306 (69%), Positives = 250/306 (80%)
Frame = +1
Query: 1 MIGAQALVAIPQASGSPKAYTSNIVDTSTRLQEGTISYPVSGLSATYQNNKVTIFATLTL 60
M GAQALVAI Q+SG+PKAYTS+I ++ T+L E ISYP SGL AT++NN+VTI+A++TL
Sbjct: 334 MNGAQALVAILQSSGTPKAYTSSIANSRTQLAESNISYPHSGLIATHENNEVTIYASITL 513
Query: 61 PNGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIGSRQRRRNT 120
P GT SLVH+WQDG +S STPQ H S++ SKE LDL SG S+ SG GS RRRNT
Sbjct: 514 PVGTPSLVHLWQDGAMSG-STPQMHDMTSANTQSKESLDLRSGASEQGSGGGSLSRRRNT 690
Query: 121 HGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSAYIVGLSGFGTGLKLGS 180
HGVLNAISWGILMP GAVIARYLKVFKSADPAWFYLH+TCQ +AYIVG++G+GTGLKLGS
Sbjct: 691 HGVLNAISWGILMPLGAVIARYLKVFKSADPAWFYLHVTCQSAAYIVGVAGWGTGLKLGS 870
Query: 181 DSEGITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNIYHHVVGYVTISISIVN 240
DS G+TY THR L IV+ L TLQVFAL LRP KDHK+RFYWN+YH VGY TI ISI+N
Sbjct: 871 DSAGVTYSTHRTLGIVIFCLGTLQVFALLLRPKKDHKIRFYWNLYHWGVGYATIIISIIN 1050
Query: 241 VFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLEAYTWMVCMKRKKADNKTSDGVN 300
+FKGFEAL DRY NWKHAY GII ALGG+AVLLEAYTW++ +KRKK++NK G+N
Sbjct: 1051IFKGFEALEVSAADRYDNWKHAYTGIIAALGGVAVLLEAYTWIIVIKRKKSENKL-QGMN 1227
Query: 301 GANGHG 306
G NG+G
Sbjct: 1228GTNGNG 1245
>TC90068 similar to GP|20197988|gb|AAD22321.2 expressed protein {Arabidopsis
thaliana}, partial (88%)
Length = 1352
Score = 191 bits (486), Expect = 2e-49
Identities = 111/324 (34%), Positives = 167/324 (51%), Gaps = 15/324 (4%)
Frame = +1
Query: 1 MIGAQALVAIPQASGSPKAYTSNIVDTSTRLQEGTI-SYPVSGLS-------------AT 46
M G AL++ P + I+DT+ +LQ+ + S P ++ AT
Sbjct: 238 MTGTNALISFPDPNTGQIVLLPFILDTTVKLQKSPLLSQPFDNINLLSSSAAMYGGKMAT 417
Query: 47 YQNNK-VTIFATLTLPNGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTS 105
N + I+A L L + T + VW G+ +P H S +S D++SG+S
Sbjct: 418 IHNGAPIQIYAKLKLESNKTKIHLVWNRGLYVQGYSPTIHPTTSIDLSSIATFDVLSGSS 597
Query: 106 QAASGIGSRQRRRNTHGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSAY 165
++S R HGVLNAISWGIL+PTGA+ ARYL+ F++ P+WFY H Q+ +
Sbjct: 598 SSSSQHTDLTMLRVIHGVLNAISWGILLPTGAITARYLRHFQTLGPSWFYAHAGIQMFGF 777
Query: 166 IVGLSGFGTGLKLGSDSEGITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNIY 225
I+G GFG G++L + G+ Y HR L I + L LQ AL RPN +K R YW Y
Sbjct: 778 ILGTVGFGIGIQLRKMTPGVEYGLHRKLGIAVFCLGALQTLALLFRPNTTNKFRKYWKSY 957
Query: 226 HHVVGYVTISISIVNVFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLEAYTWMVC 285
HH VGY + + VNVF+GFE +G Y K +Y + L G+++ LE +W++
Sbjct: 958 HHFVGYSCVVLGFVNVFQGFEVIG--ASRSYA--KLSYCLGLSTLIGVSIALEVNSWVIF 1125
Query: 286 MKRKKADNKTSDGVNGANGHGSST 309
++ K + +G+ G + S+
Sbjct: 1126CRKSKEEKMRKEGLIGTSSDKGSS 1197
>CB894342 similar to GP|20197988|gb expressed protein {Arabidopsis thaliana},
partial (64%)
Length = 838
Score = 170 bits (430), Expect = 7e-43
Identities = 99/251 (39%), Positives = 133/251 (52%), Gaps = 2/251 (0%)
Frame = +3
Query: 42 GLSATYQNNK-VTIFATLTLPNGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDL 100
G AT N + I+A L + T + VW G+ +P H S+ +S LD+
Sbjct: 90 GKRATIHNGAPIQIYAKFKLESNKTKIHLVWNRGLYVQGYSPTIHPTTSTDLSSIATLDV 269
Query: 101 VSGTSQAASGIGSRQRRRNTHGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITC 160
+SG+S A R HG LNAISWGIL+P GA+ ARY + +S PAWFY H
Sbjct: 270 LSGSS--ARQHTDLTMLRVIHGTLNAISWGILLPMGAITARYFRHIQSLGPAWFYAHAGI 443
Query: 161 QVSAYIVGLSGFGTGLKLGSDSEGITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRF 220
Q+ A+I+G GF G+ LG S G+ Y HR L + + L LQ AL RPN +K R
Sbjct: 444 QLFAFILGTVGFAIGIHLGQLSPGVEYSLHRKLGVAVFCLGALQTLALLFRPNARNKFRK 623
Query: 221 YWNIYHHVVGYVTISISIVNVFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLEAY 280
YW YHH VGY + + VNVF+GFE +G Y K +Y + L G+++ LE
Sbjct: 624 YWKSYHHFVGYSCVVLGFVNVFQGFEVMG--ASRSYA--KLSYCLGLSTLIGVSIALEVN 791
Query: 281 TW-MVCMKRKK 290
+W M C K K+
Sbjct: 792 SWVMFCRKSKE 824
>TC89232 similar to PIR|T47967|T47967 hypothetical protein F15G16.140 -
Arabidopsis thaliana, partial (38%)
Length = 1224
Score = 65.5 bits (158), Expect(2) = 6e-24
Identities = 36/119 (30%), Positives = 61/119 (51%)
Frame = +1
Query: 165 YIVGLSGFGTGLKLGSDSEGITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNI 224
+ GL+ GL+L S + HR + I ++ L+ LQ+ A FLRPN+D K R WN+
Sbjct: 814 FAFGLATVLLGLQLYSQMH-VHIPAHRGIGIFVLVLSILQILAFFLRPNRDSKFRKMWNL 990
Query: 225 YHHVVGYVTISISIVNVFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLEAYTWM 283
YH G + + + +N+ G +A G WK +Y + G + + ++LE ++
Sbjct: 991 YHSWFGRMALFFAALNIVLGMQAAG-----ATNAWKTSYGFLAGIIIVVVIVLEILAYL 1152
Score = 62.8 bits (151), Expect(2) = 6e-24
Identities = 44/168 (26%), Positives = 71/168 (42%), Gaps = 7/168 (4%)
Frame = +3
Query: 1 MIGAQALVAIPQASGSPKA---YTSNIVDTSTRLQEGTISYPVSGLSATYQNNKVTIFAT 57
M+G+ A+V G K Y + +G + P++ + A N I+
Sbjct: 327 MVGSSAMVGWINKHGHAKIKQYYLRGNKSSEVIANKGEL--PLNNVPAAVATNGAEIYLA 500
Query: 58 LTLPNGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKE--VLDLVSGTSQAASGIGSRQ 115
L + + + +L + ST H H S + K + D SG + S
Sbjct: 501 FQLQ----TTIPFGKQPILLAFSTKHPHEHHLSKHDDKTTIIFDFSSGAKSGSMDPASNS 668
Query: 116 --RRRNTHGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQ 161
+ R HG++ I WG+++P GA++ARY FK + WFYLH Q
Sbjct: 669 LIQMRKNHGIIGIIGWGLILPVGAIVARY---FKHKETLWFYLHSIIQ 803
>TC82242 weakly similar to GP|21592781|gb|AAM64730.1 putative membrane
protein {Arabidopsis thaliana}, partial (9%)
Length = 580
Score = 83.6 bits (205), Expect = 9e-17
Identities = 41/109 (37%), Positives = 62/109 (56%)
Frame = +2
Query: 189 THRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNIYHHVVGYVTISISIVNVFKGFEAL 248
THR I++ T +T+Q+ A L+P R YWN+YHH +GY ++I ++N+FKG L
Sbjct: 14 THRDFGILIFTFSTIQMLAFRLKPKSTDDYRKYWNMYHHFLGYGLLAIIVINIFKGINIL 193
Query: 249 GDFVGDRYKNWKHAYIGIIGALGGIAVLLEAYTWMVCMKRKKADNKTSD 297
+W+ +YIGI+ LG IA LE +TW+ + K +K D
Sbjct: 194 -----HGGSSWRWSYIGILIGLGTIAFALEIFTWIKFIMDKWKKDKHHD 325
>TC88786 weakly similar to GP|7268619|emb|CAB78828.1 unknown protein
{Arabidopsis thaliana}, partial (22%)
Length = 1166
Score = 59.3 bits (142), Expect = 2e-09
Identities = 46/176 (26%), Positives = 74/176 (41%), Gaps = 5/176 (2%)
Frame = +1
Query: 89 SSHQNSKEVLDLVSGTSQAASGIGSRQRRRNT-HGVLNAISWGILMPTGAVIARYLKVFK 147
SS Q +E+L S + + R + T HG L S G LMP G + R +
Sbjct: 343 SSSQEHQEILGANSTNNDNHIKLSPRLQFEITLHGFLLWASMGFLMPIGILAIRLSN--R 516
Query: 148 SADPAW----FYLHITCQVSAYIVGLSGFGTGLKLGSDSEGITYDTHRALAIVLVTLATL 203
+P W FY+H QV A ++ +G +K + + + H+ L + L + L
Sbjct: 517 EENPRWLRILFYVHTIFQVIAVLLATAGAIMSIK---NFNNLFNNNHQRLGVALYGVIWL 687
Query: 204 QVFALFLRPNKDHKLRFYWNIYHHVVGYVTISISIVNVFKGFEALGDFVGDRYKNW 259
QV RP + K R W H ++G + ++NV+ G A + + W
Sbjct: 688 QVLVGIFRPQRGSKRRSVWFFAHWILGTAVTFLGVLNVYIGLAAYHEKTSKGIRIW 855
>TC77701 similar to PIR|T47789|T47789 hypothetical protein F17J16.120 -
Arabidopsis thaliana, partial (4%)
Length = 961
Score = 54.3 bits (129), Expect = 6e-08
Identities = 34/105 (32%), Positives = 55/105 (52%)
Frame = +1
Query: 1 MIGAQALVAIPQASGSPKAYTSNIVDTSTRLQEGTISYPVSGLSATYQNNKVTIFATLTL 60
M+GAQAL+A + +G+ YT N+ + ++S GLSA N +TIFA + L
Sbjct: 295 MVGAQALIAY-KTNGNVGVYTYNLTSFGGINEVKSLSVETWGLSAEESNGVITIFAGVKL 471
Query: 61 PNGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTS 105
P + ++ VWQ G + + P +H E + N+ L +V T+
Sbjct: 472 PEKSDNVTQVWQVGPVVA-GKPGKHLFEKENLNAFTALSVVGSTT 603
>AL381775
Length = 487
Score = 31.6 bits (70), Expect = 0.41
Identities = 31/151 (20%), Positives = 61/151 (39%), Gaps = 7/151 (4%)
Frame = +2
Query: 1 MIGAQALVAIPQASGSPKAYTSNIVDTST--RLQEGTISYPVSGLSATYQNNKVTIFATL 58
MIG+ ++A P +GS T I + + R+ + S+ +N K T
Sbjct: 26 MIGSYIIMAWPSTNGSA-IITQRIAERYSVPRVTSQQSDLILDTTSSGVKNGKFIAKFTR 202
Query: 59 TLPNGTTSLVHVWQDGV--LSSDSTPQEHSHESS---HQNSKEVLDLVSGTSQAASGIGS 113
+ G +++ QD V L ++ P + + H ++ + +S +
Sbjct: 203 AVSVGGSTIESSGQDFVWALQTEERPPDDASTRDIFIHNGKGRYTYVMEENTVGSSSLSK 382
Query: 114 RQRRRNTHGVLNAISWGILMPTGAVIARYLK 144
+ HG++ WGI++P IAR+ +
Sbjct: 383 YDKYIMVHGIIMFSVWGIIVPGAVFIARFTR 475
>TC88537 weakly similar to GP|21536818|gb|AAM61150.1 F-box protein family
AtFBW2 {Arabidopsis thaliana}, partial (52%)
Length = 1395
Score = 29.6 bits (65), Expect = 1.5
Identities = 21/78 (26%), Positives = 37/78 (46%)
Frame = -3
Query: 35 TISYPVSGLSATYQNNKVTIFATLTLPNGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNS 94
T ++ +S L + +++ +I L P T L+H LSS P HSH+ Q +
Sbjct: 1183 TQAHHLSFLHNHHYHHQHSIHYQLQNPKLNTHLIH------LSSHHLPNHHSHKHQEQKA 1022
Query: 95 KEVLDLVSGTSQAASGIG 112
++ L + S+ S +G
Sbjct: 1021 PKL*VLENSASRTCSRLG 968
>TC90900 weakly similar to PIR|D96753|D96753 Similar to disease resistance
proteins [imported] - Arabidopsis thaliana, partial (7%)
Length = 826
Score = 29.3 bits (64), Expect = 2.0
Identities = 18/50 (36%), Positives = 25/50 (50%)
Frame = +3
Query: 120 THGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSAYIVGL 169
+HG + +L A Y K FKS DPA L++T +V Y+ GL
Sbjct: 615 SHGTCVSYEVPLLNNNDARELFYRKAFKSQDPASECLNLTPEVLKYVEGL 764
>BF643238 similar to PIR|T51383|T51 receptor protein kinase-like protein -
Arabidopsis thaliana, partial (29%)
Length = 585
Score = 28.5 bits (62), Expect = 3.4
Identities = 23/71 (32%), Positives = 35/71 (48%), Gaps = 6/71 (8%)
Frame = -2
Query: 184 GITYDTHRALAIVLVTLATLQVFALFLRPN------KDHKLRFYWNIYHHVVGYVTISIS 237
G++ D H A ++V L L FA L PN D K RF+W + ++ +S
Sbjct: 380 GLSLD-HIAHEGIIVLLLLLMQFAS-LSPNPTLV*PSDAKRRFHWTCRGKLKVFLMLSYK 207
Query: 238 IVNVFKGFEAL 248
I + +GF+AL
Sbjct: 206 IHTI*EGFDAL 174
>AW256730 weakly similar to GP|9757922|dbj respiratory burst oxidase protein
{Arabidopsis thaliana}, partial (11%)
Length = 681
Score = 27.7 bits (60), Expect = 5.9
Identities = 31/128 (24%), Positives = 54/128 (41%), Gaps = 10/128 (7%)
Frame = +2
Query: 77 SSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIGSRQRRRNTHGVLNAISWGILMPTG 136
S D P + +N +V+ + ++A S + + + R N + + N S M +G
Sbjct: 182 SIDIVPNVEVPKKDEENESKVIK--ASDNEAFSNVDNLRSRNNNNNINNNGSKMRRMESG 355
Query: 137 AVIARYLKVFKSAD--------PAWFYLH--ITCQVSAYIVGLSGFGTGLKLGSDSEGIT 186
A AR LK + D AW + T ++ FGT + +G+DS+
Sbjct: 356 A--ARGLKGLRVLDRTVTGKEADAWKSIEKRFTQHAVDGMLSKDKFGTCMGMGADSKDFA 529
Query: 187 YDTHRALA 194
+ + ALA
Sbjct: 530 GELYEALA 553
>CA921391 similar to GP|1669591|dbj O-methyltransferase {Glycyrrhiza
echinata}, partial (42%)
Length = 690
Score = 27.3 bits (59), Expect = 7.7
Identities = 11/26 (42%), Positives = 15/26 (57%), Gaps = 4/26 (15%)
Frame = +2
Query: 207 ALFLRPNKDHKLRFY----WNIYHHV 228
A F+RP +HKL Y WN + H+
Sbjct: 479 AFFIRPIMEHKLNHYSISLWNTFKHI 556
>TC80679 phosphatase-like protein Mtc923 [Medicago truncatula]
Length = 1390
Score = 27.3 bits (59), Expect = 7.7
Identities = 15/49 (30%), Positives = 23/49 (46%)
Frame = -1
Query: 69 HVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIGSRQRR 117
H W D + S+ +H+ HQ + LD VS +++ S G Q R
Sbjct: 502 HPW*DLGQAYRSSDHQHNTPEVHQTQNQNLDRVSHHAKSESHFGDYQNR 356
>BG580862
Length = 596
Score = 27.3 bits (59), Expect = 7.7
Identities = 24/96 (25%), Positives = 41/96 (42%)
Frame = +2
Query: 77 SSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIGSRQRRRNTHGVLNAISWGILMPTG 136
+++ E S +SH+ S E + G S + + I + +R +GV +S+G+L
Sbjct: 236 NTNDNAYELSDSTSHEGSAESEESKKGDSLSINEIMKKLKR---YGVSGILSYGLLN--- 397
Query: 137 AVIARYLKVFKSADPAWFYLHITCQVSAYIVGLSGF 172
A YL F WFY+ Y+ + F
Sbjct: 398 --TAYYLTTFLF---VWFYIAPAPGKMGYLAAVERF 490
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,424,308
Number of Sequences: 36976
Number of extensions: 131204
Number of successful extensions: 795
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 785
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 789
length of query: 310
length of database: 9,014,727
effective HSP length: 96
effective length of query: 214
effective length of database: 5,465,031
effective search space: 1169516634
effective search space used: 1169516634
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC147406.4


