

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147405.11 + phase: 0
(874 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
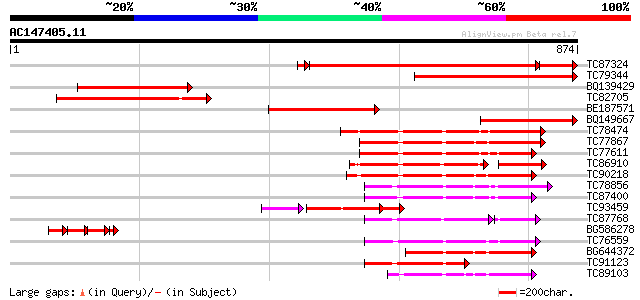
Score E
Sequences producing significant alignments: (bits) Value
TC87324 homologue to PIR|E86217|E86217 protein T27G7.10 [importe... 726 0.0
TC79344 homologue to PIR|E86217|E86217 protein T27G7.10 [importe... 508 e-144
BQ139429 homologue to PIR|E86217|E8 protein T27G7.10 [imported] ... 358 4e-99
TC82705 similar to GP|22022522|gb|AAM83219.1 AT4g03080/T4I9_4 {A... 318 4e-87
BE187571 homologue to PIR|E86217|E8 protein T27G7.10 [imported] ... 300 2e-81
BQ149667 homologue to GP|10638981|em protein phosphatase {Fagus ... 261 6e-70
TC78474 homologue to PIR|T09550|T09550 phosphoprotein phosphatas... 260 1e-69
TC77867 PIR|T09544|T09544 phosphoprotein phosphatase (EC 3.1.3.1... 260 2e-69
TC77611 SP|P48488|PP1_MEDVA Serine/threonine protein phosphatase... 256 3e-68
TC86910 homologue to PIR|T09548|T09548 phosphoprotein phosphatas... 187 1e-64
TC90218 homologue to SP|P48490|PP1_PHAVU Serine/threonine protei... 243 2e-64
TC78856 GP|7248359|dbj|BAA92697.1 type 2A protein phosphatase-1 ... 203 3e-52
TC87400 SP|Q06009|P2A_MEDSA Serine/threonine protein phosphatase... 201 1e-51
TC93459 similar to GP|22022522|gb|AAM83219.1 AT4g03080/T4I9_4 {A... 126 2e-46
TC87768 homologue to SP|P48528|PPX2_ARATH Serine/threonine prote... 153 4e-46
BG586278 homologue to PIR|E86217|E86 protein T27G7.10 [imported]... 80 8e-45
TC76559 GP|20385063|gb|AAM21172.1 serine/threonine protein phosp... 178 9e-45
BG644372 homologue to SP|O04860|P2A5 Serine/threonine protein ph... 158 7e-39
TC91123 homologue to GP|7248361|dbj|BAA92698.1 type 2A protein p... 142 7e-34
TC89103 similar to GP|16930441|gb|AAL31906.1 At2g42810/F7D19.19 ... 135 5e-32
>TC87324 homologue to PIR|E86217|E86217 protein T27G7.10 [imported] -
Arabidopsis thaliana, partial (37%)
Length = 1897
Score = 726 bits (1874), Expect(3) = 0.0
Identities = 356/356 (100%), Positives = 356/356 (100%)
Frame = +2
Query: 462 LHHRAVVIAAETGGALGGMVRQLSIDQFENEGRRVSYGTPESATAARKLLDRQMSINSVP 521
LHHRAVVIAAETGGALGGMVRQLSIDQFENEGRRVSYGTPESATAARKLLDRQMSINSVP
Sbjct: 56 LHHRAVVIAAETGGALGGMVRQLSIDQFENEGRRVSYGTPESATAARKLLDRQMSINSVP 235
Query: 522 KKVIAHLLKPRGWKPPVRRQFFLDCNEIADLCDSAERIFSSEPTVIQLRAPIKIFGDLHG 581
KKVIAHLLKPRGWKPPVRRQFFLDCNEIADLCDSAERIFSSEPTVIQLRAPIKIFGDLHG
Sbjct: 236 KKVIAHLLKPRGWKPPVRRQFFLDCNEIADLCDSAERIFSSEPTVIQLRAPIKIFGDLHG 415
Query: 582 QFGDLMRLFDEYGSPSTAGDIAYIDYLFLGDYVDRGQHSLETISLLLALKVEYPNNIHLI 641
QFGDLMRLFDEYGSPSTAGDIAYIDYLFLGDYVDRGQHSLETISLLLALKVEYPNNIHLI
Sbjct: 416 QFGDLMRLFDEYGSPSTAGDIAYIDYLFLGDYVDRGQHSLETISLLLALKVEYPNNIHLI 595
Query: 642 RGNHEAADINALFGFRIECIERMGERDGIWTWHRINRLFNWLPLAALIEKKIICMHGGIG 701
RGNHEAADINALFGFRIECIERMGERDGIWTWHRINRLFNWLPLAALIEKKIICMHGGIG
Sbjct: 596 RGNHEAADINALFGFRIECIERMGERDGIWTWHRINRLFNWLPLAALIEKKIICMHGGIG 775
Query: 702 RSINHLEQIENIQRPITMEAGSIVLMDLLWSDPTENDSVEGLRPNARGPGLVTFGPDRVM 761
RSINHLEQIENIQRPITMEAGSIVLMDLLWSDPTENDSVEGLRPNARGPGLVTFGPDRVM
Sbjct: 776 RSINHLEQIENIQRPITMEAGSIVLMDLLWSDPTENDSVEGLRPNARGPGLVTFGPDRVM 955
Query: 762 EFCNNNDLQLIVRAHECVMDGFERFAQGHLITLFSATNYCGTANNAGAILVLGRDL 817
EFCNNNDLQLIVRAHECVMDGFERFAQGHLITLFSATNYCGTANNAGAILVLGRDL
Sbjct: 956 EFCNNNDLQLIVRAHECVMDGFERFAQGHLITLFSATNYCGTANNAGAILVLGRDL 1123
Score = 126 bits (317), Expect(3) = 0.0
Identities = 58/58 (100%), Positives = 58/58 (100%)
Frame = +3
Query: 817 LVVVPKLIHPLPPALSPETSPERHIEDTWMQELNANRPPTPTRGRPPVANDRGSLAWI 874
LVVVPKLIHPLPPALSPETSPERHIEDTWMQELNANRPPTPTRGRPPVANDRGSLAWI
Sbjct: 1122 LVVVPKLIHPLPPALSPETSPERHIEDTWMQELNANRPPTPTRGRPPVANDRGSLAWI 1295
Score = 41.2 bits (95), Expect(3) = 0.0
Identities = 18/18 (100%), Positives = 18/18 (100%)
Frame = +1
Query: 444 ISSMIKPEPGGPNSGGVR 461
ISSMIKPEPGGPNSGGVR
Sbjct: 1 ISSMIKPEPGGPNSGGVR 54
>TC79344 homologue to PIR|E86217|E86217 protein T27G7.10 [imported] -
Arabidopsis thaliana, partial (22%)
Length = 1301
Score = 508 bits (1309), Expect = e-144
Identities = 244/252 (96%), Positives = 248/252 (97%), Gaps = 1/252 (0%)
Frame = +1
Query: 624 ISLLLALKVEYPNNIHLIRGNHEAADINALFGFRIECIERMGERDGIWTWHRINRLFNWL 683
I+LLLALKVEYP N+HLIRGNHEAADINALFGFRIECIERMGERDGIWTWHRINRLFNWL
Sbjct: 7 ITLLLALKVEYPTNVHLIRGNHEAADINALFGFRIECIERMGERDGIWTWHRINRLFNWL 186
Query: 684 PLAALIEKKIICMHGGIGRSINHLEQIENIQRPITMEAGSIVLMDLLWSDPTENDSVEGL 743
PLAALIEKKIICMHGGIGRSINH+EQIENIQRPI MEAGSIVLMDLLWSDPTENDSVEGL
Sbjct: 187 PLAALIEKKIICMHGGIGRSINHIEQIENIQRPIPMEAGSIVLMDLLWSDPTENDSVEGL 366
Query: 744 RPNARGPGLVTFGPDRVMEFCNNNDLQLIVRAHECVMDGFERFAQGHLITLFSATNYCGT 803
RPNARGPGLVTFGPDRVMEFCNNNDLQLIVRAHECVMDGFERFAQGHLITLFSATNYCGT
Sbjct: 367 RPNARGPGLVTFGPDRVMEFCNNNDLQLIVRAHECVMDGFERFAQGHLITLFSATNYCGT 546
Query: 804 ANNAGAILVLGRDLVVVPKLIHPLPPAL-SPETSPERHIEDTWMQELNANRPPTPTRGRP 862
ANNAGAILVLGRDLVVVPKLIHPLPPA+ SPETSPERHIEDTWMQELNANRPPTPTRGRP
Sbjct: 547 ANNAGAILVLGRDLVVVPKLIHPLPPAISSPETSPERHIEDTWMQELNANRPPTPTRGRP 726
Query: 863 PVANDRGSLAWI 874
PV NDRGSLAWI
Sbjct: 727 PVTNDRGSLAWI 762
>BQ139429 homologue to PIR|E86217|E8 protein T27G7.10 [imported] -
Arabidopsis thaliana, partial (13%)
Length = 531
Score = 358 bits (920), Expect = 4e-99
Identities = 174/177 (98%), Positives = 174/177 (98%)
Frame = +1
Query: 105 MALVGQRYLMAIGGNDGKRPLADVWALDTAAKPYEWRKLEPEGEGPPPCMYATASARSDG 164
MALVGQRYLMAIGGNDGKRPLADVWALDTAAKPYEWRKLEPEGEGPPPCMYATASARSDG
Sbjct: 1 MALVGQRYLMAIGGNDGKRPLADVWALDTAAKPYEWRKLEPEGEGPPPCMYATASARSDG 180
Query: 165 LLLLCGGRDASSVPLASAYGLAKHRDGRWEWAIAPGVSPSSRYQHAAVFVNARLHVSGGA 224
LLLLCGGRDASSVPLASAYGLAKHRDGRWEWAIAPGVSPSSRYQHAAVFVNARLHVSGGA
Sbjct: 181 LLLLCGGRDASSVPLASAYGLAKHRDGRWEWAIAPGVSPSSRYQHAAVFVNARLHVSGGA 360
Query: 225 LGGGRMVEDSSSVAVLDTAAGVWCDTKSVVTTPRTGRYSADAAGGDASVELVRRCRH 281
LGGGRMVEDSSSVAVLDTAAGVWCDTKSVVTTPRTGRYSADAAGGDAS LVRRC H
Sbjct: 361 LGGGRMVEDSSSVAVLDTAAGVWCDTKSVVTTPRTGRYSADAAGGDASXXLVRRCXH 531
Score = 40.0 bits (92), Expect = 0.004
Identities = 37/120 (30%), Positives = 48/120 (39%), Gaps = 6/120 (5%)
Frame = +1
Query: 22 ADVHCYDVLTN--KWSRITPFGEPPTPRAAHVATAVGTMVVIQGGIGPAGLSAEDLHVLD 79
ADV D +W ++ P GE P P A+A +++ G G S
Sbjct: 64 ADVWALDTAAKPYEWRKLEPEGEGPPPCMYATASARSDGLLLLCG-GRDASSVPLASAYG 240
Query: 80 LTQQRP-RWHRVGVPGPGPGPRYGHVMALVGQRYLMAIGGNDGKRPLAD---VWALDTAA 135
L + R RW PG P RY H V R ++ G G R + D V LDTAA
Sbjct: 241 LAKHRDGRWEWAIAPGVSPSSRYQHAAVFVNARLHVSGGALGGGRMVEDSSSVAVLDTAA 420
>TC82705 similar to GP|22022522|gb|AAM83219.1 AT4g03080/T4I9_4 {Arabidopsis
thaliana}, partial (26%)
Length = 849
Score = 318 bits (816), Expect = 4e-87
Identities = 153/238 (64%), Positives = 181/238 (75%)
Frame = +2
Query: 73 EDLHVLDLTQQRPRWHRVGVPGPGPGPRYGHVMALVGQRYLMAIGGNDGKRPLADVWALD 132
+DL+VLDLT + +WHRV V G GPGPRYGHVM LV QRYL+ + GNDGKR ++D W LD
Sbjct: 8 DDLYVLDLTNDKYKWHRVVVQGQGPGPRYGHVMDLVAQRYLVTVSGNDGKRVVSDAWTLD 187
Query: 133 TAAKPYEWRKLEPEGEGPPPCMYATASARSDGLLLLCGGRDASSVPLASAYGLAKHRDGR 192
TA KPY W+KL PEG+ P MYATASARSDG+ LLCGGRD+S P A AYGL HR+G
Sbjct: 188 TAQKPYAWQKLNPEGDRPSARMYATASARSDGMFLLCGGRDSSGTPQADAYGLLMHRNGT 367
Query: 193 WEWAIAPGVSPSSRYQHAAVFVNARLHVSGGALGGGRMVEDSSSVAVLDTAAGVWCDTKS 252
WEW +APGVSPS RYQHA+VFV ARLHV+GG L GGR VE S+AVLDTAAGVW D
Sbjct: 368 WEWTLAPGVSPSPRYQHASVFVGARLHVTGGVLRGGRAVEGEPSIAVLDTAAGVWLDRNG 547
Query: 253 VVTTPRTGRYSADAAGGDASVELVRRCRHAAAAVGDLIFIYGGLRGGILLDDLLVAED 310
+V++ R+ + D S+EL+RRCRHAAAAVG ++IYGGLRG +LL D LVAE+
Sbjct: 548 IVSSSRSNK----GHDYDHSLELMRRCRHAAAAVGVQVYIYGGLRGDLLLGDFLVAEN 709
>BE187571 homologue to PIR|E86217|E8 protein T27G7.10 [imported] -
Arabidopsis thaliana, partial (14%)
Length = 523
Score = 300 bits (767), Expect = 2e-81
Identities = 155/172 (90%), Positives = 160/172 (92%), Gaps = 2/172 (1%)
Frame = +2
Query: 400 VEASAAEAEAISATLAAAKARQENGGVESPDRDRGAEATPSGKQISSMIKPEPGGPN--S 457
VEA AEAEAISA LAAAKARQENG VE PDRDRGAEATPSGK SS+IKP+P N S
Sbjct: 5 VEAYPAEAEAISAALAAAKARQENGEVELPDRDRGAEATPSGKHTSSLIKPDPVANNIAS 184
Query: 458 GGVRLHHRAVVIAAETGGALGGMVRQLSIDQFENEGRRVSYGTPESATAARKLLDRQMSI 517
GGVRLHHRAVVIAAETGGALGGMVRQLSIDQFENEGRRVSYGTPE+ATAARKLLDRQMSI
Sbjct: 185 GGVRLHHRAVVIAAETGGALGGMVRQLSIDQFENEGRRVSYGTPENATAARKLLDRQMSI 364
Query: 518 NSVPKKVIAHLLKPRGWKPPVRRQFFLDCNEIADLCDSAERIFSSEPTVIQL 569
NSVPKKVIAHLLKPRGWKPPVRRQFFLDCNEIADLCDSAERIFSSEP+V+QL
Sbjct: 365 NSVPKKVIAHLLKPRGWKPPVRRQFFLDCNEIADLCDSAERIFSSEPSVLQL 520
>BQ149667 homologue to GP|10638981|em protein phosphatase {Fagus sylvatica},
partial (57%)
Length = 646
Score = 261 bits (668), Expect = 6e-70
Identities = 127/150 (84%), Positives = 135/150 (89%), Gaps = 1/150 (0%)
Frame = +3
Query: 726 LMDLLWSDPTENDSVEGLRPNARGPGLVTFGPDRVMEFCNNNDLQLIVRAHECVMDGFER 785
LMDLLWSDPTENDSVEGLRPNARGPGLVTFGPDRV EFC N LQLI+RAHECVMDGFER
Sbjct: 3 LMDLLWSDPTENDSVEGLRPNARGPGLVTFGPDRVTEFCKKNKLQLIIRAHECVMDGFER 182
Query: 786 FAQGHLITLFSATNYCGTANNAGAILVLGRDLVVVPKLIHPLPPAL-SPETSPERHIEDT 844
FAQG LITLFSATNYCGTANNAGAILV+GR LVVVPKLIHP+PP L SPE+SPER +++T
Sbjct: 183 FAQGQLITLFSATNYCGTANNAGAILVVGRGLVVVPKLIHPIPPPLQSPESSPERVMDET 362
Query: 845 WMQELNANRPPTPTRGRPPVANDRGSLAWI 874
WMQELN RPPTPTRGRP DRGSLA+I
Sbjct: 363 WMQELNIQRPPTPTRGRPQPDLDRGSLAYI 452
>TC78474 homologue to PIR|T09550|T09550 phosphoprotein phosphatase (EC
3.1.3.16) 1 catalytic epsilon chain - alfalfa, complete
Length = 1597
Score = 260 bits (665), Expect = 1e-69
Identities = 144/316 (45%), Positives = 198/316 (62%)
Frame = +3
Query: 511 LDRQMSINSVPKKVIAHLLKPRGWKPPVRRQFFLDCNEIADLCDSAERIFSSEPTVIQLR 570
L Q+ +V +I L + R +P +Q L EI LC ++ IF +P +++L
Sbjct: 165 LQAQVIDETVLDDIIRRLTEVRLSRPG--KQVQLSEAEIKQLCSASRDIFLQQPNLLELE 338
Query: 571 APIKIFGDLHGQFGDLMRLFDEYGSPSTAGDIAYIDYLFLGDYVDRGQHSLETISLLLAL 630
APIKI GD+HGQ+ DL+RLF+ G P + +YLFLGDYVDRG+ SLETI LLLA
Sbjct: 339 APIKICGDIHGQYSDLLRLFEYGGLPPQS------NYLFLGDYVDRGKQSLETICLLLAY 500
Query: 631 KVEYPNNIHLIRGNHEAADINALFGFRIECIERMGERDGIWTWHRINRLFNWLPLAALIE 690
K++YP N L+RGNHE A IN ++GF EC R R W FN LP+AALI+
Sbjct: 501 KIKYPENFFLLRGNHECASINRIYGFYDECKRRFNVR----LWKAFTESFNCLPVAALID 668
Query: 691 KKIICMHGGIGRSINHLEQIENIQRPITMEAGSIVLMDLLWSDPTENDSVEGLRPNARGP 750
+KI+CMHGG+ + +L+QI N+ RP+ + + L DLLWSDP ++ V+G N RG
Sbjct: 669 EKILCMHGGLSPDLTNLDQIRNLPRPVPIPDTGL-LCDLLWSDPGKD--VKGWGMNDRGV 839
Query: 751 GLVTFGPDRVMEFCNNNDLQLIVRAHECVMDGFERFAQGHLITLFSATNYCGTANNAGAI 810
TFGPD+V EF +DL LI RAH+ V DG+E FA L+T+FSA NYCG +NAGA+
Sbjct: 840 S-YTFGPDKVAEFLTRHDLDLICRAHQVVEDGYEFFADRQLVTIFSAPNYCGEFDNAGAM 1016
Query: 811 LVLGRDLVVVPKLIHP 826
+ + +L+ +++ P
Sbjct: 1017MSVDENLMCSFQILKP 1064
>TC77867 PIR|T09544|T09544 phosphoprotein phosphatase (EC 3.1.3.16)
catalytic beta chain - alfalfa, complete
Length = 1583
Score = 260 bits (664), Expect = 2e-69
Identities = 140/287 (48%), Positives = 186/287 (64%)
Frame = +3
Query: 540 RQFFLDCNEIADLCDSAERIFSSEPTVIQLRAPIKIFGDLHGQFGDLMRLFDEYGSPSTA 599
+Q L EI LC +A IF +P +++L APIKI GD+HGQ+ DL+RLF+ G P A
Sbjct: 294 KQVQLSETEIRQLCSTAREIFLQQPNLLELEAPIKICGDIHGQYSDLLRLFEYGGLPPQA 473
Query: 600 GDIAYIDYLFLGDYVDRGQHSLETISLLLALKVEYPNNIHLIRGNHEAADINALFGFRIE 659
+YLFLGDYVDRG+ SLETI LLLA K++YP N L+RGNHE A IN ++GF E
Sbjct: 474 ------NYLFLGDYVDRGKQSLETICLLLAYKIKYPENFFLLRGNHECASINRIYGFYDE 635
Query: 660 CIERMGERDGIWTWHRINRLFNWLPLAALIEKKIICMHGGIGRSINHLEQIENIQRPITM 719
C R R W FN LP+AALI++KI+CMHGG+ +++L+QI N+QRP T
Sbjct: 636 CKRRFNVR----IWKVFTDCFNCLPVAALIDEKILCMHGGLSPDLHNLDQIRNLQRP-TD 800
Query: 720 EAGSIVLMDLLWSDPTENDSVEGLRPNARGPGLVTFGPDRVMEFCNNNDLQLIVRAHECV 779
+ +L DLLWSDP++ V+G N RG TFG D+V EF +DL LI RAH+ V
Sbjct: 801 VPDTGLLCDLLWSDPSK--EVQGWGMNDRGVS-YTFGADKVSEFLQKHDLDLICRAHQVV 971
Query: 780 MDGFERFAQGHLITLFSATNYCGTANNAGAILVLGRDLVVVPKLIHP 826
DG+E FA L+T+FSA NYCG +NAGA++ + L+ +++ P
Sbjct: 972 EDGYEFFANRQLVTIFSAPNYCGEFDNAGAMMSVDETLMCSFQILKP 1112
>TC77611 SP|P48488|PP1_MEDVA Serine/threonine protein phosphatase PP1 (EC
3.1.3.16). {Medicago varia}, complete
Length = 1509
Score = 256 bits (653), Expect = 3e-68
Identities = 136/272 (50%), Positives = 181/272 (66%)
Frame = +2
Query: 540 RQFFLDCNEIADLCDSAERIFSSEPTVIQLRAPIKIFGDLHGQFGDLMRLFDEYGSPSTA 599
+Q L +I LC SA+ IF S+P +++L APIKI GD+HGQF DL+RLF+ G P A
Sbjct: 251 KQVHLTEADIRQLCTSAKEIFLSQPNLLELEAPIKICGDVHGQFSDLLRLFEYGGYPPEA 430
Query: 600 GDIAYIDYLFLGDYVDRGQHSLETISLLLALKVEYPNNIHLIRGNHEAADINALFGFRIE 659
+YLFLGDYVDRG+ S+ETI LLLA K++Y N L+RGNHE A IN ++GF E
Sbjct: 431 ------NYLFLGDYVDRGKQSIETICLLLAYKIKYKENFFLLRGNHECASINRIYGFYDE 592
Query: 660 CIERMGERDGIWTWHRINRLFNWLPLAALIEKKIICMHGGIGRSINHLEQIENIQRPITM 719
C R R W FN LP+AAL+++KI+CMHGG+ + +L+QI NI RPI +
Sbjct: 593 CKRRYNVR----LWKTFTDCFNCLPVAALVDEKILCMHGGLSPELKNLDQIRNIARPIDV 760
Query: 720 EAGSIVLMDLLWSDPTENDSVEGLRPNARGPGLVTFGPDRVMEFCNNNDLQLIVRAHECV 779
+ L DLLW+DP ++ +EG N RG TFG D+V+EF ++DL LI RAH+ V
Sbjct: 761 PDHGL-LCDLLWADPDKD--LEGWGENDRGVSF-TFGADKVVEFLEHHDLDLICRAHQVV 928
Query: 780 MDGFERFAQGHLITLFSATNYCGTANNAGAIL 811
DG+E FA+ L+T+FSA NYCG +NAGA++
Sbjct: 929 EDGYEFFAKRKLVTVFSAPNYCGEFDNAGAMM 1024
>TC86910 homologue to PIR|T09548|T09548 phosphoprotein phosphatase (EC
3.1.3.16) 1 catalytic chain delta - alfalfa, complete
Length = 1523
Score = 187 bits (475), Expect(2) = 1e-64
Identities = 104/215 (48%), Positives = 142/215 (65%), Gaps = 1/215 (0%)
Frame = +2
Query: 524 VIAHLLKPRGWKPPVRRQFFLDCNEIADLCDSAERIFSSEPTVIQLRAPIKIFGDLHGQF 583
+I LL+ RG +P +Q L EI LC ++ IF ++P +++L APIKI GD+HGQ+
Sbjct: 269 IINRLLEVRG-RPG--KQVQLSEAEIKQLCLVSKDIFMNQPNLLKLEAPIKICGDIHGQY 439
Query: 584 GDLMRLFDEYGSPSTAGDIAYIDYLFLGDYVDRGQHSLETISLLLALKVEYPNNIHLIRG 643
DL+RLF+ G P + +YLFLGDYVDRG+ SLETI LLLA K++YP N L+RG
Sbjct: 440 SDLLRLFEYGGFPPRS------NYLFLGDYVDRGKQSLETICLLLAYKIKYPKNFFLLRG 601
Query: 644 NHEAADINALFGFRIECIERMGERDGIWTWHRINRLFNWLPLAALIEKKIICMHGGIGRS 703
NHE A IN ++GF EC R + W FN LP+AALI++KIICMHGG+
Sbjct: 602 NHECASINRIYGFYDECKRRY----NVKLWKMFTDCFNCLPVAALIDEKIICMHGGLSPE 769
Query: 704 INHLEQIENIQRPITM-EAGSIVLMDLLWSDPTEN 737
++ L QI+N++RP + E+G +L DLLWSDP+ +
Sbjct: 770 LHDLNQIKNLRRPCEVPESG--LLCDLLWSDPSSD 868
Score = 78.6 bits (192), Expect(2) = 1e-64
Identities = 37/74 (50%), Positives = 49/74 (66%)
Frame = +3
Query: 754 TFGPDRVMEFCNNNDLQLIVRAHECVMDGFERFAQGHLITLFSATNYCGTANNAGAILVL 813
TFG DRV EF +DL LI RAH+ V DG+E FA L+T+FSA NYCG +N GA++ +
Sbjct: 909 TFGADRVKEFLQKHDLDLICRAHQVVEDGYEFFANRQLVTIFSAPNYCGDFDNVGAMMTV 1088
Query: 814 GRDLVVVPKLIHPL 827
LV +++ PL
Sbjct: 1089NESLVCSFQILKPL 1130
>TC90218 homologue to SP|P48490|PP1_PHAVU Serine/threonine protein
phosphatase PP1 (EC 3.1.3.16). [Kidney bean French
bean], partial (95%)
Length = 1024
Score = 243 bits (620), Expect = 2e-64
Identities = 139/293 (47%), Positives = 183/293 (62%)
Frame = +2
Query: 519 SVPKKVIAHLLKPRGWKPPVRRQFFLDCNEIADLCDSAERIFSSEPTVIQLRAPIKIFGD 578
+V V LL+ +G K Q L +EI LC +A +IF S+P ++ LRAPI+I GD
Sbjct: 119 TVVDDVTRRLLEGKGGK-----QVQLSESEIRQLCINARQIFLSQPILLDLRAPIRICGD 283
Query: 579 LHGQFGDLMRLFDEYGSPSTAGDIAYIDYLFLGDYVDRGQHSLETISLLLALKVEYPNNI 638
+HGQ+ DL+RLF+ G P A +YLFLGDYVDRG+ SLETI LLLA K+ YP+ +
Sbjct: 284 IHGQYQDLLRLFEYGGYPPAA------NYLFLGDYVDRGKQSLETICLLLAYKIRYPDRV 445
Query: 639 HLIRGNHEAADINALFGFRIECIERMGERDGIWTWHRINRLFNWLPLAALIEKKIICMHG 698
+L+RGNHE A IN ++GF EC R R W FN LP+AALI++KI+CMHG
Sbjct: 446 YLLRGNHEDAKINRIYGFYDECKRRFNVR----LWKIFTDCFNCLPVAALIDEKILCMHG 613
Query: 699 GIGRSINHLEQIENIQRPITMEAGSIVLMDLLWSDPTENDSVEGLRPNARGPGLVTFGPD 758
G+ + +L+QI + RP T S +L DLLWSDP + EG + RG TFG D
Sbjct: 614 GLSPELQNLDQIREVTRP-TEIPDSGLLCDLLWSDP--DPKAEGWADSDRGVS-CTFGSD 781
Query: 759 RVMEFCNNNDLQLIVRAHECVMDGFERFAQGHLITLFSATNYCGTANNAGAIL 811
V EF + NDL LI R H+ V DG+E FA+ L+T+FSA NY G +NA ++
Sbjct: 782 VVAEFLDKNDLDLICRGHQVVEDGYEFFAKRRLVTIFSAPNYGGEFDNARCVI 940
>TC78856 GP|7248359|dbj|BAA92697.1 type 2A protein phosphatase-1 {Vicia
faba}, complete
Length = 1426
Score = 203 bits (516), Expect = 3e-52
Identities = 113/289 (39%), Positives = 165/289 (56%)
Frame = +3
Query: 548 EIADLCDSAERIFSSEPTVIQLRAPIKIFGDLHGQFGDLMRLFDEYGSPSTAGDIAYIDY 607
++ LCD A I E V ++ P+ + GD+HGQF DL+ LF G+ +Y
Sbjct: 258 DVKALCDQARAILVEEWNVQPVKCPVTVCGDIHGQFYDLIELF------RIGGNAPDTNY 419
Query: 608 LFLGDYVDRGQHSLETISLLLALKVEYPNNIHLIRGNHEAADINALFGFRIECIERMGER 667
LF+GDYVDRG +S+ET+SLL+ALKV Y + I ++RGNHE+ I ++GF EC+ + G
Sbjct: 420 LFMGDYVDRGYYSVETVSLLVALKVRYRDRITILRGNHESRQITQVYGFYDECLRKYGNA 599
Query: 668 DGIWTWHRINRLFNWLPLAALIEKKIICMHGGIGRSINHLEQIENIQRPITMEAGSIVLM 727
+ W LF++LPL ALIE +I C+HGG+ S++ L+ I + R I +
Sbjct: 600 N---VWKFFTDLFDYLPLTALIESQIFCLHGGLSPSLDTLDNIRALDR-IQEVPHEGPMC 767
Query: 728 DLLWSDPTENDSVEGLRPNARGPGLVTFGPDRVMEFCNNNDLQLIVRAHECVMDGFERFA 787
DLLWSDP + G+ P G TFG D +F + N L LI RAH+ VM+G+
Sbjct: 768 DLLWSDPDDRCG-WGISPRGAG---YTFGQDIAAQFNHTNGLSLISRAHQLVMEGYNWCQ 935
Query: 788 QGHLITLFSATNYCGTANNAGAILVLGRDLVVVPKLIHPLPPALSPETS 836
+ +++T+FSA NYC N AIL +G ++ P P + P+T+
Sbjct: 936 EKNVVTVFSAPNYCYRCGNMAAILEIGENMDQNFLQFDPAPRQIEPDTT 1082
>TC87400 SP|Q06009|P2A_MEDSA Serine/threonine protein phosphatase PP2A
catalytic subunit (EC 3.1.3.16). [Alfalfa], complete
Length = 1439
Score = 201 bits (510), Expect = 1e-51
Identities = 110/265 (41%), Positives = 155/265 (57%), Gaps = 1/265 (0%)
Frame = +1
Query: 548 EIADLCDSAERIFSSEPTVIQLRAPIKIFGDLHGQFGDLMRLFDEYGS-PSTAGDIAYID 606
++ +LC+ A+ I E V +++P+ I GD+HGQF DL LF G P T +
Sbjct: 181 QVKELCEKAKEILMDESNVQPVKSPVTICGDIHGQFHDLAELFRIGGKCPDT-------N 339
Query: 607 YLFLGDYVDRGQHSLETISLLLALKVEYPNNIHLIRGNHEAADINALFGFRIECIERMGE 666
YLF+GDYVDRG +S+ET++LL+ALKV YP I ++RGNHE+ I ++GF EC+ + G
Sbjct: 340 YLFMGDYVDRGYYSVETVTLLVALKVRYPQRITILRGNHESRQITQVYGFYDECLRKYGS 519
Query: 667 RDGIWTWHRINRLFNWLPLAALIEKKIICMHGGIGRSINHLEQIENIQRPITMEAGSIVL 726
+ W LF++ PL AL+E +I C+HGG+ SI L+ I N R + +
Sbjct: 520 AN---VWKIFTDLFDFFPLTALVESEIFCLHGGLSPSIETLDNIRNFDR-VQEVPHEGPM 687
Query: 727 MDLLWSDPTENDSVEGLRPNARGPGLVTFGPDRVMEFCNNNDLQLIVRAHECVMDGFERF 786
DLLWSDP + G+ P G TFG D +F + N L+LI RAH+ VMDGF
Sbjct: 688 CDLLWSDPDDRCG-WGISPRGAG---YTFGQDISEQFNHTNSLKLIARAHQLVMDGFNWA 855
Query: 787 AQGHLITLFSATNYCGTANNAGAIL 811
+ ++T+FSA NYC N +IL
Sbjct: 856 HEQKVVTIFSAPNYCYRCGNMASIL 930
>TC93459 similar to GP|22022522|gb|AAM83219.1 AT4g03080/T4I9_4 {Arabidopsis
thaliana}, partial (20%)
Length = 676
Score = 126 bits (316), Expect(3) = 2e-46
Identities = 65/120 (54%), Positives = 84/120 (69%)
Frame = +1
Query: 458 GGVRLHHRAVVIAAETGGALGGMVRQLSIDQFENEGRRVSYGTPESATAARKLLDRQMSI 517
G VRLH RAVV+A E L G+VRQLS+DQFENE RR+ +S +K RQ S
Sbjct: 226 GDVRLHPRAVVVAKEAVSNLSGLVRQLSLDQFENESRRMLPINNDSPYPTKKFT-RQKSP 402
Query: 518 NSVPKKVIAHLLKPRGWKPPVRRQFFLDCNEIADLCDSAERIFSSEPTVIQLRAPIKIFG 577
+ KK+I++LLKPR WK PV R+ FLD E+ +LC +AE+IF EPTV+QL+AP+K+FG
Sbjct: 403 QGLHKKIISNLLKPRNWKAPVNRRVFLDSYEVGELCYAAEQIFMHEPTVLQLKAPVKVFG 582
Score = 68.2 bits (165), Expect(3) = 2e-46
Identities = 31/39 (79%), Positives = 32/39 (81%)
Frame = +2
Query: 570 RAPIKIFGDLHGQFGDLMRLFDEYGSPSTAGDIAYIDYL 608
+ P K GDLHGQFGDLMRLFDEYG PSTA DI YIDYL
Sbjct: 560 KPPSKYLGDLHGQFGDLMRLFDEYGFPSTARDITYIDYL 676
Score = 31.6 bits (70), Expect(3) = 2e-46
Identities = 23/68 (33%), Positives = 28/68 (40%), Gaps = 3/68 (4%)
Frame = +3
Query: 388 GSRRTSKGVEYLVEASAAEAEAISATLAAAKARQENGGVE---SPDRDRGAEATPSGKQI 444
GS +E L EASAAEAEA SA + +A + E S D A+ G
Sbjct: 42 GSGLDKVSLEKLREASAAEAEAASAVWQSVQAISSSPADETFVSDDNSHAADTVSDGSDT 221
Query: 445 SSMIKPEP 452
P P
Sbjct: 222 VGRCSPSP 245
>TC87768 homologue to SP|P48528|PPX2_ARATH Serine/threonine protein
phosphatase PP-X isozyme 2 (EC 3.1.3.16). [Mouse-ear
cress], complete
Length = 1430
Score = 153 bits (386), Expect(2) = 4e-46
Identities = 86/199 (43%), Positives = 116/199 (58%)
Frame = +3
Query: 547 NEIADLCDSAERIFSSEPTVIQLRAPIKIFGDLHGQFGDLMRLFDEYGSPSTAGDIAYID 606
+E+ LC A I E V ++ AP+ I GD+HGQF D+ LF GD +
Sbjct: 195 SEVKSLCLKAIEILVEESNVQRVDAPVTICGDIHGQFYDMKELF------KVGGDCPKTN 356
Query: 607 YLFLGDYVDRGQHSLETISLLLALKVEYPNNIHLIRGNHEAADINALFGFRIECIERMGE 666
YLFLGD+VDRG +S+ET LLLALKV YP+ I LIRGNHE+ I ++GF EC+ + G
Sbjct: 357 YLFLGDFVDRGFYSVETFLLLLALKVRYPDRITLIRGNHESRQITQVYGFYDECLRKYG- 533
Query: 667 RDGIWTWHRINRLFNWLPLAALIEKKIICMHGGIGRSINHLEQIENIQRPITMEAGSIVL 726
+ W +F++L L+ALIE KI +HGG+ +I+ L+QI I R + +
Sbjct: 534 --SVNVWRYCTDIFDYLSLSALIENKIFSVHGGLSPAISTLDQIRTIDRKQEVPHDG-AM 704
Query: 727 MDLLWSDPTENDSVEGLRP 745
DLLWSDP + GL P
Sbjct: 705 CDLLWSDPEDIVDGWGLSP 761
Score = 51.2 bits (121), Expect(2) = 4e-46
Identities = 27/70 (38%), Positives = 40/70 (56%)
Frame = +1
Query: 748 RGPGLVTFGPDRVMEFCNNNDLQLIVRAHECVMDGFERFAQGHLITLFSATNYCGTANNA 807
RG G + FG V F + N++ I RAH+ VM+G++ ++T++SA NYC N
Sbjct: 763 RGAGFL-FGGSVVTGFNHTNNIDYICRAHQLVMEGYKWMFNNQIVTVWSAPNYCYRCGNV 939
Query: 808 GAILVLGRDL 817
AIL L +L
Sbjct: 940 AAILELDGNL 969
>BG586278 homologue to PIR|E86217|E86 protein T27G7.10 [imported] -
Arabidopsis thaliana, partial (8%)
Length = 755
Score = 80.5 bits (197), Expect(4) = 8e-45
Identities = 34/34 (100%), Positives = 34/34 (100%)
Frame = +2
Query: 121 GKRPLADVWALDTAAKPYEWRKLEPEGEGPPPCM 154
GKRPLADVWALDTAAKPYEWRKLEPEGEGPPPCM
Sbjct: 509 GKRPLADVWALDTAAKPYEWRKLEPEGEGPPPCM 610
Score = 69.3 bits (168), Expect(4) = 8e-45
Identities = 31/35 (88%), Positives = 33/35 (93%)
Frame = +3
Query: 89 RVGVPGPGPGPRYGHVMALVGQRYLMAIGGNDGKR 123
RV V GPGPGPRYGHVMALVGQRYLMAIGGNDG++
Sbjct: 315 RVVVQGPGPGPRYGHVMALVGQRYLMAIGGNDGRQ 419
Score = 65.1 bits (157), Expect(4) = 8e-45
Identities = 28/30 (93%), Positives = 30/30 (99%)
Frame = +2
Query: 60 VIQGGIGPAGLSAEDLHVLDLTQQRPRWHR 89
++QGGIGPAGLSAEDLHVLDLTQQRPRWHR
Sbjct: 137 LVQGGIGPAGLSAEDLHVLDLTQQRPRWHR 226
Score = 25.8 bits (55), Expect(4) = 8e-45
Identities = 12/13 (92%), Positives = 12/13 (92%)
Frame = +3
Query: 155 YATASARSDGLLL 167
YA ASARSDGLLL
Sbjct: 693 YAPASARSDGLLL 731
>TC76559 GP|20385063|gb|AAM21172.1 serine/threonine protein phosphatase 2A
{Pisum sativum}, complete
Length = 1305
Score = 178 bits (451), Expect = 9e-45
Identities = 98/272 (36%), Positives = 154/272 (56%), Gaps = 1/272 (0%)
Frame = +2
Query: 547 NEIADLCDSAERIFSSEPTVIQLRAPIKIFGDLHGQFGDLMRLFDEYGSPSTAGDIAYID 606
+E+ LC+ + I E V + +P+ + GD+HGQF DLM+LF T G + +
Sbjct: 185 DELQLLCEYVKEILIEESNVQPVNSPVTVCGDIHGQFHDLMKLFQ------TGGHVPETN 346
Query: 607 YLFLGDYVDRGQHSLETISLLLALKVEYPNNIHLIRGNHEAADINALFGFRIECIERMGE 666
Y+F+GD+VDRG +SLE ++LL LK YP NI L+RGNHE+ + ++GF EC + G
Sbjct: 347 YIFMGDFVDRGYNSLEVFTILLLLKARYPANITLLRGNHESRQLTQVYGFYDECQRKYGN 526
Query: 667 RDGIWTWHRINRLFNWLPLAALIEKKIICMHGGIGRSINHLEQIENIQRPITMEAGSIVL 726
+ W +F++L L+A+I+ ++C+HGG+ I ++QI I+R +
Sbjct: 527 AN---AWRYCTDVFDYLTLSAIIDGTVLCVHGGLSPDIRTIDQIRVIERNCEIPHEG-PF 694
Query: 727 MDLLWSDPTENDSVEGLRPNARGPGLVTFGPDRVMEFCNNNDLQLIVRAHECVMDGFE-R 785
DL+WSDP + +E + RG G + FG EF + N+L L+ RAH+ V +G +
Sbjct: 695 CDLMWSDP---EDIETWAVSPRGAGWL-FGSRVTSEFNHINNLDLVCRAHQLVQEGLKYM 862
Query: 786 FAQGHLITLFSATNYCGTANNAGAILVLGRDL 817
F L+T++SA NYC N +IL ++
Sbjct: 863 FQDKGLVTVWSAPNYCYRCGNVASILSFNENM 958
>BG644372 homologue to SP|O04860|P2A5 Serine/threonine protein phosphatase
PP2A-5 catalytic subunit (EC 3.1.3.16). [Common
tobacco], partial (69%)
Length = 658
Score = 158 bits (400), Expect = 7e-39
Identities = 83/202 (41%), Positives = 122/202 (60%)
Frame = +1
Query: 610 LGDYVDRGQHSLETISLLLALKVEYPNNIHLIRGNHEAADINALFGFRIECIERMGERDG 669
+GDYVDRG +S+ET++LL+ALKV YP + ++RGNHE+ I ++GF EC+ + G +
Sbjct: 4 MGDYVDRGYYSVETVTLLVALKVRYPQRLTILRGNHESRQITQVYGFYDECLRKYGNAN- 180
Query: 670 IWTWHRINRLFNWLPLAALIEKKIICMHGGIGRSINHLEQIENIQRPITMEAGSIVLMDL 729
W LF++ PL AL+E +I C+HGG+ SI L+ + + R + + DL
Sbjct: 181 --VWKTFTDLFDYFPLTALVESEIFCLHGGLSPSIETLDNVRSFDR-VQEVPHEGAMCDL 351
Query: 730 LWSDPTENDSVEGLRPNARGPGLVTFGPDRVMEFCNNNDLQLIVRAHECVMDGFERFAQG 789
LWSDP D G + RG G TFG D +F + N+L+LI RAH+ VM+G+ +
Sbjct: 352 LWSDP---DDRCGWGMSPRGAG-YTFGQDISEQFHHTNNLKLIARAHQLVMEGYNWSHEQ 519
Query: 790 HLITLFSATNYCGTANNAGAIL 811
++T+FSA NYC N +IL
Sbjct: 520 KVVTIFSAPNYCYRCGNMASIL 585
>TC91123 homologue to GP|7248361|dbj|BAA92698.1 type 2A protein
phosphatase-2 {Vicia faba}, partial (57%)
Length = 690
Score = 142 bits (357), Expect = 7e-34
Identities = 69/161 (42%), Positives = 102/161 (62%)
Frame = +2
Query: 548 EIADLCDSAERIFSSEPTVIQLRAPIKIFGDLHGQFGDLMRLFDEYGSPSTAGDIAYIDY 607
E+ LCD A I E V ++ P+ + GD+HGQF DL+ LF G+ + +Y
Sbjct: 230 EVKALCDQARAILVEEWNVQPVKCPVTVCGDIHGQFYDLIELF------RIGGNAPHTNY 391
Query: 608 LFLGDYVDRGQHSLETISLLLALKVEYPNNIHLIRGNHEAADINALFGFRIECIERMGER 667
LF+GDYVDRG +S+ET+SLL+ALKV Y + I ++RGNHE+ I ++GF EC+ + G
Sbjct: 392 LFMGDYVDRGYYSVETVSLLVALKVRYRDRITILRGNHESRQITQVYGFYDECLRKYGNA 571
Query: 668 DGIWTWHRINRLFNWLPLAALIEKKIICMHGGIGRSINHLE 708
+ W LF++LPL ALIE ++ C+HGG+ S++ L+
Sbjct: 572 N---VWKFFTDLFDYLPLTALIESQVFCLHGGLSPSLDTLD 685
>TC89103 similar to GP|16930441|gb|AAL31906.1 At2g42810/F7D19.19
{Arabidopsis thaliana}, partial (66%)
Length = 1107
Score = 135 bits (341), Expect = 5e-32
Identities = 86/231 (37%), Positives = 123/231 (53%), Gaps = 2/231 (0%)
Frame = +1
Query: 583 FGDLMRLFDEYGSPSTAGDIAYIDYLFLGDYVDRGQHSLETISLLLALKVEYPNNIHLIR 642
F DL+ +F+ G PS YLF GD+VDRG S+E L A K P+ +++ R
Sbjct: 250 FYDLLNIFELNGIPSEENP-----YLFNGDFVDRGSFSVEVAFTLFAFKCMSPSAMYIAR 414
Query: 643 GNHEAADINALFGFRIECIERMGERDGIWTWHRINRLFNWLPLAALIEKKIICMHGGIGR 702
GNHE+ +N +FGF E ++ ++ +FN LPLA +I KK+ +HGG+
Sbjct: 415 GNHESKSMNKIFGFEGEVRAKLNDK----FVKMFAEVFNCLPLAHVINKKVFVVHGGLF- 579
Query: 703 SIN--HLEQIENIQRPITMEAGSIVLMDLLWSDPTENDSVEGLRPNARGPGLVTFGPDRV 760
S++ L QI +I R ++ DLLWSDP + G P+ RG FG D
Sbjct: 580 SVDGVKLSQIRSINR-FCEPPDEGLMHDLLWSDP---QPLPGRGPSKRGNRCFCFGEDVT 747
Query: 761 MEFCNNNDLQLIVRAHECVMDGFERFAQGHLITLFSATNYCGTANNAGAIL 811
F +N+L L+VR+H+ + G+E G LIT+FSA NYC N GA +
Sbjct: 748 KRFLKDNNLDLVVRSHKSMHKGYEIQHDGKLITVFSAPNYCDKKGNKGAFI 900
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.137 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,837,343
Number of Sequences: 36976
Number of extensions: 392436
Number of successful extensions: 2092
Number of sequences better than 10.0: 76
Number of HSP's better than 10.0 without gapping: 1924
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2009
length of query: 874
length of database: 9,014,727
effective HSP length: 105
effective length of query: 769
effective length of database: 5,132,247
effective search space: 3946697943
effective search space used: 3946697943
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC147405.11


