

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
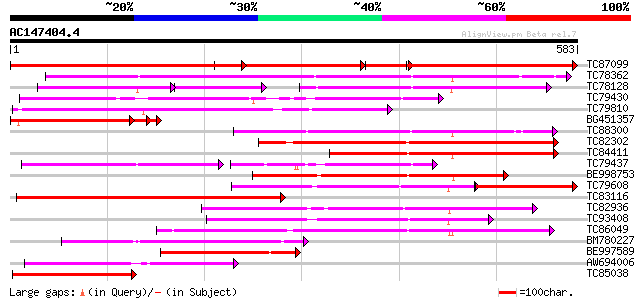
Query= AC147404.4 - phase: 0
(583 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC87099 similar to PIR|G86449|G86449 hypothetical protein AAF813... 496 0.0
TC78362 similar to GP|15391731|emb|CAA93316. nitrite transporter... 304 5e-83
TC78128 weakly similar to GP|21928025|gb|AAM78041.1 At1g69870/T1... 177 5e-75
TC79430 similar to PIR|F86358|F86358 Similar to peptide transpor... 255 4e-68
TC79810 similar to GP|11994403|dbj|BAB02362. nitrate transporter... 229 2e-60
BG451357 similar to PIR|G86449|G86 hypothetical protein AAF81343... 211 3e-60
TC88300 similar to GP|9581817|emb|CAC00544.1 putative low-affini... 225 4e-59
TC82302 similar to PIR|F84663|F84663 probable nitrate transporte... 216 1e-56
TC84411 similar to GP|4102839|gb|AAD01600.1| LeOPT1 {Lycopersico... 214 1e-55
TC79437 similar to PIR|G96720|G96720 nitrate transporter (NTL1) ... 120 1e-55
BE998753 similar to PIR|T45958|T45 oligopeptide transporter-like... 213 1e-55
TC79608 similar to GP|20147231|gb|AAM10330.1 At1g68570/F24J5_7 {... 149 2e-53
TC83116 similar to GP|11933407|dbj|BAB19758. putative nitrate tr... 201 5e-52
TC82936 similar to PIR|G84829|G84829 probable PTR2 family peptid... 200 1e-51
TC93408 similar to GP|11933407|dbj|BAB19758. putative nitrate tr... 198 4e-51
TC86049 weakly similar to GP|20466248|gb|AAM20441.1 putative tra... 195 4e-50
BM780227 similar to PIR|E84798|E84 probable peptide/amino acid t... 166 2e-41
BE997589 weakly similar to PIR|G86449|G86 hypothetical protein A... 152 4e-37
AW694006 similar to PIR|F86358|F86 Similar to peptide transporte... 148 5e-36
TC85038 weakly similar to PIR|G86449|G86449 hypothetical protein... 146 2e-35
>TC87099 similar to PIR|G86449|G86449 hypothetical protein AAF81343.1
[imported] - Arabidopsis thaliana, partial (86%)
Length = 2205
Score = 496 bits (1276), Expect = e-140
Identities = 243/243 (100%), Positives = 243/243 (100%)
Frame = +2
Query: 1 GKFKEETEELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLT 60
GKFKEETEELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLT
Sbjct: 113 GKFKEETEELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLT 292
Query: 61 RVLGQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTT 120
RVLGQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTT
Sbjct: 293 RVLGQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTT 472
Query: 121 YLALLRPKGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSK 180
YLALLRPKGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSK
Sbjct: 473 YLALLRPKGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSK 652
Query: 181 ESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPK 240
ESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPK
Sbjct: 653 ESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPK 832
Query: 241 YRH 243
YRH
Sbjct: 833 YRH 841
Score = 347 bits (889), Expect(3) = 0.0
Identities = 172/175 (98%), Positives = 172/175 (98%)
Frame = +2
Query: 409 KKSSSKGLTELQRMGIGLVIAIIAMVTAGIVECYRLKYAKQGDTSSLSIFWQIPQYALIG 468
K SKGLTELQRMGIGLVIAIIAMVTAGIVECYRLKYAKQGDTSSLSIFWQIPQYALIG
Sbjct: 1340 KNQVSKGLTELQRMGIGLVIAIIAMVTAGIVECYRLKYAKQGDTSSLSIFWQIPQYALIG 1519
Query: 469 ASEVFMYVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGW 528
ASEVFMYVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGW
Sbjct: 1520 ASEVFMYVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGW 1699
Query: 529 IPGNLNRGHLDRFFFLLAVLTSLDLIAYIACAKWFQNIQMACKYDNNDEPSSCKV 583
IPGNLNRGHLDRFFFLLAVLTSLDLIAYIACAKWFQNIQMACKYDNNDEPSSCKV
Sbjct: 1700 IPGNLNRGHLDRFFFLLAVLTSLDLIAYIACAKWFQNIQMACKYDNNDEPSSCKV 1864
Score = 263 bits (673), Expect(3) = 0.0
Identities = 133/156 (85%), Positives = 135/156 (86%)
Frame = +3
Query: 211 EDEGLWALGFWASAGSAFLALVLFLVGTPKYRHFKPCGNPLSRFCQVFFAASRKLGVQMT 270
+D GL LGF G F L FL+ FKPCGNPLSRFCQVFFAASRKLGVQMT
Sbjct: 750 KDYGL--LGFGLLLGLPFWLLYFFLLAPQNIDTFKPCGNPLSRFCQVFFAASRKLGVQMT 923
Query: 271 SNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRAAYITSRDLEVQKGGQHNPWYLCPITQ 330
SNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRAAYITSRDLEVQKGGQHNPW LCPITQ
Sbjct: 924 SNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRAAYITSRDLEVQKGGQHNPWXLCPITQ 1103
Query: 331 VEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGA 366
VEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGA
Sbjct: 1104VEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGA 1211
Score = 92.4 bits (228), Expect(3) = 0.0
Identities = 47/49 (95%), Positives = 48/49 (97%)
Frame = +1
Query: 367 AMKTTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKG 415
AMKTTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSS +G
Sbjct: 1213 AMKTTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSFQG 1359
>TC78362 similar to GP|15391731|emb|CAA93316. nitrite transporter {Cucumis
sativus}, partial (88%)
Length = 1871
Score = 304 bits (779), Expect = 5e-83
Identities = 176/550 (32%), Positives = 304/550 (55%), Gaps = 9/550 (1%)
Frame = +3
Query: 37 ILLNQGLATLAFFGVGVNLVLFLTRVLGQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYW 96
IL N+ A G N++ +LT+ L +A+N +S ++G + L+GAF++DS+
Sbjct: 111 ILANEVCDRFASAGFHANMITYLTQQLNMPLVSASNTLSNFSGLSSLTPLLGAFIADSFA 290
Query: 97 GRYKTCAIFQGIFVTGLVSLSVTTYLALLRPKGCGNGKLECGEHSSLEMGMFYLSIYLIA 156
GR+ T I+ GL++++ + + RP C ++ C E S ++ + YL+++L +
Sbjct: 291 GRFWTIVFATLIYELGLITITTSAIVPHFRPPPCPT-QVNCQEAKSSQLWILYLALFLTS 467
Query: 157 LGNGGYQPNIATFGADQFDEDHSKESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLW 216
LG+GG + + F DQFD K F++++ + SL + TI+ Y +D W
Sbjct: 468 LGSGGIRSCVVPFSGDQFDMTKKGVESRKWNLFNWYFFCMGFASLSALTIVVYIQDHMGW 647
Query: 217 ALGFWASAGSAFLALVLFLVGTPKYRHFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDL 276
G + F+A++ FL+G+ Y+ KP G+P R QV AA RK + ++ L
Sbjct: 648 GWGLGIPTIAMFIAIIAFLLGSRLYKTLKPSGSPFLRLAQVIVAAFRKRNDALPNDPKLL 827
Query: 277 YVIDEKESSNNSNRKILHTHGFKFLDRAAYITSRDLEVQKGGQHNPWYLCPITQVEEVKC 336
Y E +SS + ++ HT +K+LD+AA +T + + N W L + +VEE+KC
Sbjct: 828 YQNLELDSSISLEGRLSHTDQYKWLDKAAIVTDEEAK-NLNKLPNLWNLATVHRVEELKC 1004
Query: 337 ILRLLPIWLCTIIYSVVFTQMASLFVEQGAAMKTTIS-SFKIPPASMSSFDILSVAIFIF 395
++R+LPIW I+ + S + Q M +S +F+I PASM+ F +L++ +
Sbjct: 1005LVRMLPIWASGILLITASSSQHSFVIVQARTMDRHLSHTFEISPASMAIFSVLTMMTGVI 1184
Query: 396 FYRRVLDPLVGKLKKSSSKGLTELQRMGIGLVIAIIAMVTAGIVECYRLKYAKQGDTSS- 454
Y R+ P + + K+ + G+T LQRMGIG VI IIA + + ++E R K A +
Sbjct: 1185LYERLFVPFIRRFTKNPA-GITCLQRMGIGFVINIIATIVSALLEIKRKKVASKYHLLDS 1361
Query: 455 ------LSIFWQIPQYALIGASEVFMYVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYV 508
+S+FW +PQY L G +EVFM VG LEF Q+P+ ++S +AL +I++G+++
Sbjct: 1362PKAIIPISVFWLVPQYFLHGVAEVFMNVGHLEFLYDQSPESMRSSATALYCIAIAIGHFI 1541
Query: 509 SSLIVSIVMKISTEDHMPGWIPG-NLNRGHLDRFFFLLAVLTSLDLIAYIACAKWFQNIQ 567
+L+V++V K + ++ W+P NLNRG L+ ++FL+ + ++ I Y+ CA W N +
Sbjct: 1542GTLLVTLVHKYTGKER--NWLPDRNLNRGRLEYYYFLVCGIQVINFIYYVICA-WIYNYK 1712
Query: 568 MACKYDNNDE 577
+ + N++
Sbjct: 1713PLEEINENNQ 1742
>TC78128 weakly similar to GP|21928025|gb|AAM78041.1 At1g69870/T17F3_10
{Arabidopsis thaliana}, partial (77%)
Length = 2022
Score = 177 bits (449), Expect(3) = 5e-75
Identities = 98/264 (37%), Positives = 148/264 (55%), Gaps = 5/264 (1%)
Frame = +2
Query: 299 KFLDRAAYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMA 358
+ LD+AA + DL G N W L I QVEEVKC+ R LPIW I+ Q
Sbjct: 1058 RVLDKAALVMEGDLNPD-GTIVNQWNLVSIQQVEEVKCLARTLPIWAAGILGFTAMAQQG 1234
Query: 359 SLFVEQGAAMKTTIS-SFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLT 417
+ V Q M + F+IP S+ ++ + +++ FY R+ P + K+ K+ G+T
Sbjct: 1235 TFIVSQAMKMDRHLGPKFQIPAGSLGVISLIVIGLWVPFYDRICVPSLRKITKNEG-GIT 1411
Query: 418 ELQRMGIGLVIAIIAMVTAGIVECYRLKYAKQGDT----SSLSIFWQIPQYALIGASEVF 473
LQR+GIG+V +II+M+ AG+VE R A T + +S+ W PQ L+G E F
Sbjct: 1412 LLQRIGIGMVFSIISMIVAGLVEKVRRDVANSNPTPQGIAPMSVMWLFPQLVLMGFCEAF 1591
Query: 474 MYVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNL 533
+G +EFFN Q PD ++S +AL S +L NYVSS++V V ++ + P W+ N+
Sbjct: 1592 NIIGLIEFFNRQFPDHMRSIANALFSCSFALANYVSSILVITVHSVTKTHNHPDWLTNNI 1771
Query: 534 NRGHLDRFFFLLAVLTSLDLIAYI 557
N G LD F++LLA + L+L+ ++
Sbjct: 1772 NEGRLDYFYYLLAGVGVLNLVYFL 1843
Score = 82.0 bits (201), Expect(3) = 5e-75
Identities = 44/145 (30%), Positives = 77/145 (52%), Gaps = 3/145 (2%)
Frame = +2
Query: 29 GRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDNAAAANNVSKWTGTVYIFSLVG 88
G W A IL N+ LA FG+ N +++LTR + A+N ++ W G L+G
Sbjct: 233 GGWKAMPFILGNESFERLAAFGLLANFMVYLTREFHLEQVHASNILNIWGGISNFAPLLG 412
Query: 89 AFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRPKGC---GNGKLECGEHSSLEM 145
AF+SD+Y GR+KT A + G+ ++++T +L L+P C +C +S ++
Sbjct: 413 AFISDTYTGRFKTIAFASFFSLLGMTAVTLTAWLPKLQPPSCTPQQQALNQCVTANSSQV 592
Query: 146 GMFYLSIYLIALGNGGYQPNIATFG 170
G ++ + +++G+ G +P FG
Sbjct: 593 GFLFMGLIFLSIGSSGIRPCSIPFG 667
Score = 62.0 bits (149), Expect(3) = 5e-75
Identities = 31/95 (32%), Positives = 48/95 (49%)
Frame = +3
Query: 170 GADQFDEDHSKESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFL 229
G DQFD K +FF+++Y + + LF+ T++ Y +D W GF
Sbjct: 666 GVDQFDPTTEKGKKGINSFFNWYYTSFTVVLLFTQTVIVYIQDSISWKFGFAIPTLCMLX 845
Query: 230 ALVLFLVGTPKYRHFKPCGNPLSRFCQVFFAASRK 264
+++LF +GT Y H KP G+ S QV A+ +K
Sbjct: 846 SIILFFIGTKIYVHVKPEGSIFSSIAQVLVASFKK 950
>TC79430 similar to PIR|F86358|F86358 Similar to peptide transporter
[imported] - Arabidopsis thaliana, partial (55%)
Length = 1303
Score = 255 bits (651), Expect = 4e-68
Identities = 156/443 (35%), Positives = 243/443 (54%), Gaps = 7/443 (1%)
Frame = +2
Query: 11 TLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDNAAA 70
T+DG VD+ GRP +R++SG W A I+ + A++G+ NL+ +LT LGQ A A
Sbjct: 80 TVDGVVDFKGRPVLRSSSGGWRAAAFIISVEVAERFAYYGINSNLINYLTGPLGQSTATA 259
Query: 71 ANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRPKGC 130
A NV+ W+GT + L+GAFL+DS+ GRY+T I +++ ++LS+ T A L G
Sbjct: 260 AENVNIWSGTASLLPLLGAFLADSFLGRYRTIIIASLVYI---LALSLLTLSATLPSDG- 427
Query: 131 GNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYSKVAFFS 190
+ +F+ S+YL+A GG++P + FGADQFD +H +E S+ +FF+
Sbjct: 428 -----------DAQAILFFFSLYLVAFAQGGHKPCVQAFGADQFDINHPQERRSRSSFFN 574
Query: 191 YFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRHFKPCGN- 249
++Y G L + +IL Y +D W LGF + +AL LF +GT YR P GN
Sbjct: 575 WWYFTFTAGLLVTVSILNYVQDNVGWVLGFGIPWIAMIIALSLFSLGTWTYRFSIP-GNQ 751
Query: 250 ---PLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILH--THGFKFLDRA 304
P SR +VF A R + E+E S +LH + F FL++A
Sbjct: 752 QRGPFSRIGRVFITALRN------------FRTTEEEEPRPS---LLHQPSQQFSFLNKA 886
Query: 305 AYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQ 364
+ G +C +++VEE K ILRL+PIW ++I+++VF+Q ++ F +Q
Sbjct: 887 L--------IASDGSKENGKVCSVSEVEEAKAILRLVPIWATSLIFAIVFSQSSTFFTKQ 1042
Query: 365 GAAM-KTTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMG 423
G + + + F +PPAS+ SF LS+ +FI Y R++ P + + G+T LQR+G
Sbjct: 1043GVTLDRKILPGFYVPPASLQSFISLSIVLFIPVYDRIIVP-IARTFTGKPSGITMLQRIG 1219
Query: 424 IGLVIAIIAMVTAGIVECYRLKY 446
G++ ++I+MV A VE R K+
Sbjct: 1220AGILFSVISMVIAAFVEMKRSKW 1288
>TC79810 similar to GP|11994403|dbj|BAB02362. nitrate transporter
{Arabidopsis thaliana}, partial (60%)
Length = 1236
Score = 229 bits (584), Expect = 2e-60
Identities = 130/393 (33%), Positives = 219/393 (55%), Gaps = 3/393 (0%)
Frame = +3
Query: 4 KEETEELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVL 63
K+ TE+ + +VD+ G P+ + SG W A +IL + + G+ +NLV +L L
Sbjct: 78 KKGTED---NAAVDFRGHPADKTKSGGWLAAGLILGTELAERICVMGISMNLVTYLVGDL 248
Query: 64 GQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLA 123
+A++A V+ + GT+ + L+G FL+D+ GRY T AI I G+ L++ T +
Sbjct: 249 HLHSASSATIVTNFMGTLNLLGLLGGFLADAKLGRYLTVAISATIATVGVCMLTLATTIP 428
Query: 124 LLRPKGCGNGKLE---CGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSK 180
+ P C + + C + S ++ + + ++Y IALG GG + N++ FG+DQFD + K
Sbjct: 429 SMTPPPCSEVRRQHHQCIQASGKQLSLLFAALYTIALGGGGIKSNVSGFGSDQFDTNDPK 608
Query: 181 ESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPK 240
E + + FF+ FY +++GSLFS +L Y +D G+ SAG+ +A+ + L GTP
Sbjct: 609 EEKNMIFFFNRFYFFISIGSLFSVVVLVYVQDNIGRGWGYGISAGTMLVAVGILLCGTPL 788
Query: 241 YRHFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKF 300
YR KP G+PL+ +V F A +K + + S D + K+ +T F+
Sbjct: 789 YRFKKPQGSPLTIIWRVLFLAWKKRTLPIPS--------DPTLLNGYLEAKVTYTDKFRS 944
Query: 301 LDRAAYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASL 360
LD+AA + + + + G NPW + +TQVEEVK +++LLPIW I++ V++QM +
Sbjct: 945 LDKAAILD--ETKSKDGNNENPWLVSTMTQVEEVKMVIKLLPIWSTCILFWTVYSQMNTF 1118
Query: 361 FVEQGAAMKTTISSFKIPPASMSSFDILSVAIF 393
+EQ M + S +IP S+S+F +++ +F
Sbjct: 1119TIEQATFMNRKVGSLEIPAGSLSAFLFITILLF 1217
>BG451357 similar to PIR|G86449|G86 hypothetical protein AAF81343.1
[imported] - Arabidopsis thaliana, partial (20%)
Length = 606
Score = 211 bits (537), Expect(3) = 3e-60
Identities = 106/133 (79%), Positives = 120/133 (89%), Gaps = 6/133 (4%)
Frame = +1
Query: 2 KFKEET------EELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNL 55
KFKEET EELTLDGSVD++GRP+IRA SGRW AGTIILLNQ LATLAFFG+GVNL
Sbjct: 121 KFKEETDFESESEELTLDGSVDFNGRPAIRAKSGRWSAGTIILLNQCLATLAFFGIGVNL 300
Query: 56 VLFLTRVLGQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVS 115
V+FLTRV+GQ+NA AANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCA+FQ IFV GL+S
Sbjct: 301 VVFLTRVVGQNNAEAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAVFQVIFVIGLMS 480
Query: 116 LSVTTYLALLRPK 128
LS+++YL L++PK
Sbjct: 481 LSLSSYLFLIKPK 519
Score = 32.0 bits (71), Expect(3) = 3e-60
Identities = 15/16 (93%), Positives = 15/16 (93%)
Frame = +3
Query: 141 SSLEMGMFYLSIYLIA 156
SS EMGMFYLSIYLIA
Sbjct: 558 SSWEMGMFYLSIYLIA 605
Score = 28.5 bits (62), Expect(3) = 3e-60
Identities = 11/22 (50%), Positives = 14/22 (63%)
Frame = +2
Query: 125 LRPKGCGNGKLECGEHSSLEMG 146
L KGCGN ++CG+H L G
Sbjct: 509 LNLKGCGNESIQCGKHFKLGDG 574
>TC88300 similar to GP|9581817|emb|CAC00544.1 putative low-affinity nitrate
transporter {Nicotiana plumbaginifolia}, partial (56%)
Length = 1169
Score = 225 bits (573), Expect = 4e-59
Identities = 130/345 (37%), Positives = 207/345 (59%), Gaps = 12/345 (3%)
Frame = +1
Query: 231 LVLFLVG--TPKYRHFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNS 288
L+LF+ + R K G+PL++ V+ AA RK +++ + L+ +D+ E
Sbjct: 1 LLLFVCSYWNKEVRFKKLAGSPLTQIAVVYVAAWRKRNMELPYDSSLLFNVDDIEDEMLR 180
Query: 289 NRK--ILHTHGFKFLDRAAYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLC 346
+K + H+ F+FLD+AA I + + WYL +T VEEVK +LR+LPIW
Sbjct: 181 KKKQVLPHSKQFRFLDKAA-IKDPKTDGNEINVVRKWYLSTLTDVEEVKLVLRMLPIWAT 357
Query: 347 TIIYSVVFTQMASLFVEQGAAMKTTIS-SFKIPPASMSSFDILSVAIFIFFYRRVLDPLV 405
TI++ V+ QM + V Q + I SF+IPPAS+++F I S+ + I Y RV+ P+
Sbjct: 358 TIMFWTVYAQMTTFSVSQATTLNRHIGKSFQIPPASLTAFFIGSILLTIPIYDRVIVPIT 537
Query: 406 GKLKKSSSKGLTELQRMGIGLVIAIIAMVTAGIVECYRLKYAKQGDTSS-------LSIF 458
K+ K+ +GLT LQR+G+GLV +I AMV A + E R++ A + + +S+F
Sbjct: 538 RKIFKNP-QGLTPLQRIGVGLVFSIFAMVAAALTELKRMRMAHLHNLTHNPNSEIPMSVF 714
Query: 459 WQIPQYALIGASEVFMYVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMK 518
W IPQ+ +G+ E F Y+GQL+FF + P G+K+ + L ++++SLG + SSL+V++V K
Sbjct: 715 WLIPQFFFVGSGEAFTYIGQLDFFLRECPKGMKTMSTGLFLSTLSLGFFFSSLLVTLVHK 894
Query: 519 ISTEDHMPGWIPGNLNRGHLDRFFFLLAVLTSLDLIAYIACAKWF 563
+ T H P W+ NLN L F++LLA+L+ L+L+ Y+ CA W+
Sbjct: 895 V-TSQHKP-WLADNLNEAKLYNFYWLLALLSVLNLVIYLLCANWY 1023
>TC82302 similar to PIR|F84663|F84663 probable nitrate transporter
[imported] - Arabidopsis thaliana, partial (51%)
Length = 1133
Score = 216 bits (551), Expect = 1e-56
Identities = 114/314 (36%), Positives = 193/314 (61%), Gaps = 5/314 (1%)
Frame = +3
Query: 256 QVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRAAYITSRDLEVQ 315
QV A+ +K +++ N LY ++S +I + F+FL++AA + D +
Sbjct: 9 QVIVASIKKRKMELPYNVGSLYEDTPEDS------RIEQSEQFRFLEKAAIVVEGDFDKD 170
Query: 316 K-GGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGAAMKTTISS 374
G NPW LC +T+VEEVK ++RLLPIW TII+ + QM + VEQ A M+ + +
Sbjct: 171 LYGSGPNPWKLCSLTRVEEVKMMVRLLPIWATTIIFWTTYAQMITFSVEQAATMERNVGN 350
Query: 375 FKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMGIGLVIAIIAMV 434
F+IP S++ F ++++ + + R++ PL KL + G + LQR IGL+++ I M
Sbjct: 351 FQIPAGSLTVFFVVAILLTLAVNDRIIMPLWKKL--NGKPGFSNLQRNAIGLLLSTIGMA 524
Query: 435 TAGIVECYRLKYAK--QGDTSSL--SIFWQIPQYALIGASEVFMYVGQLEFFNAQTPDGL 490
A ++E RL AK +G+ ++L S+F +PQ+ L+G+ E F+Y GQL+FF Q+P G+
Sbjct: 525 AASLIEVKRLSVAKGVKGNQTTLPISVFLLVPQFFLVGSGEAFIYTGQLDFFITQSPKGM 704
Query: 491 KSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGHLDRFFFLLAVLTS 550
K+ + L +T++SLG +VSS +VS+V K++ GW+ ++N+G LD F+ LL +L+
Sbjct: 705 KTMSTGLFLTTLSLGFFVSSFLVSVVKKVTGTRDGQGWLADHINKGRLDLFYALLTILSF 884
Query: 551 LDLIAYIACAKWFQ 564
++ +A++ CA W++
Sbjct: 885 INFVAFLVCAFWYK 926
>TC84411 similar to GP|4102839|gb|AAD01600.1| LeOPT1 {Lycopersicon
esculentum}, partial (38%)
Length = 804
Score = 214 bits (544), Expect = 1e-55
Identities = 107/241 (44%), Positives = 165/241 (68%), Gaps = 6/241 (2%)
Frame = +1
Query: 330 QVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGAAMKTTISSFKIPPASMSSFDILS 389
QVEE+K ++R+ PIW II+S V+ QM++LFVEQG M T+I SFK+ PAS+S+F++ S
Sbjct: 1 QVEELKILIRMFPIWATGIIFSSVYAQMSTLFVEQGTMMNTSIGSFKLSPASLSTFEVAS 180
Query: 390 VAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMGIGLVIAIIAMVTAGIVECYRLKYAKQ 449
V +++ Y ++L P+V K +G++ QR+GIG I+ + M+ A VE RL+ A++
Sbjct: 181 VVMWVPVYDKILVPIVKKFT-GKKRGISVFQRIGIGPFISGLCMLAAAAVEIKRLQLARE 357
Query: 450 GDTSS------LSIFWQIPQYALIGASEVFMYVGQLEFFNAQTPDGLKSFGSALCMTSIS 503
D LS+ WQIPQY ++GA+E+F +VGQLEFF ++PD +++ AL + + S
Sbjct: 358 LDLVDKPVGVPLSVLWQIPQYLILGAAEIFTFVGQLEFFYEESPDAMRTICGALPLLNFS 537
Query: 504 LGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGHLDRFFFLLAVLTSLDLIAYIACAKWF 563
LGNY+SS I++IV +T+ GWIP NLN+GHLD +F LL+ L+ L+++ +I AK +
Sbjct: 538 LGNYLSSFILTIVTYFTTKGGRLGWIPDNLNKGHLDYYFLLLSGLSLLNMLVFIVAAKIY 717
Query: 564 Q 564
+
Sbjct: 718 K 720
>TC79437 similar to PIR|G96720|G96720 nitrate transporter (NTL1)
53025-56402 [imported] - Arabidopsis thaliana, partial
(61%)
Length = 1424
Score = 120 bits (302), Expect(2) = 1e-55
Identities = 67/210 (31%), Positives = 110/210 (51%), Gaps = 2/210 (0%)
Frame = +3
Query: 13 DGSV--DWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDNAAA 70
DG V W +P++R G A + +L+ + L LA+ NLVL+L + + +
Sbjct: 60 DGKVMLTWRNKPALRGRHGGMVAASFVLVVEILENLAYLANASNLVLYLKEYMHMSPSKS 239
Query: 71 ANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRPKGC 130
ANNV+ + GT ++ +L+G FLSD++ Y I I + GL+ L++ ++ L+P C
Sbjct: 240 ANNVTNFMGTSFLLALLGGFLSDAFLTTYHVYLISAVIELIGLIILTIQAHVPSLKPTKC 419
Query: 131 GNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYSKVAFFS 190
N C E + + M + +YL+ALG GG + ++ G +QFDE + FF+
Sbjct: 420 -NESTPCEEVNGGKAAMLFAGLYLVALGVGGIKGSLPVHGGEQFDEATPNGRKQRSTFFN 596
Query: 191 YFYLALNLGSLFSNTILGYFEDEGLWALGF 220
YF L+ G+L + T + + ED W GF
Sbjct: 597 YFVFCLSCGALIAVTFVVWVEDNKGWEWGF 686
Score = 114 bits (286), Expect(2) = 1e-55
Identities = 77/230 (33%), Positives = 124/230 (53%), Gaps = 17/230 (7%)
Frame = +1
Query: 228 FLALVLFLVGTPKYRHFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNN 287
F ++ +FL G+ YR+ P G+PL+ +V AA+ L T+ V++ S +N
Sbjct: 709 FGSIPVFLAGSTTYRNKVPSGSPLTTILKVLIAAT--LNSCCTNKNSCSAVVNMSSSPSN 882
Query: 288 SNRKI----LH------------THGFKFLDRAAYITSRDLEVQKGGQHNPWYLCPITQV 331
+ + LH + KFL++A IT++ + C I Q+
Sbjct: 883 PSTQAKEQELHQTLKPTTPSQIPSESLKFLNKA--ITNKPIHSS--------LQCTIQQL 1032
Query: 332 EEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGAAMKTTI-SSFKIPPASMSSFDILSV 390
E+VK + ++ PI+ CTI+ + Q+++ VEQ A M TT+ SSFK+PPAS+ F +L +
Sbjct: 1033EDVKLVFKIFPIFACTIMLNACLAQLSTFSVEQAATMNTTLFSSFKVPPASLPVFPVLFL 1212
Query: 391 AIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMGIGLVIAIIAMVTAGIVE 440
I Y ++ P K+ KS + G+T LQR+GIGL+++I+AM A IVE
Sbjct: 1213MILAPIYDHIIIPYARKVTKSEA-GITHLQRIGIGLILSIVAMAIAAIVE 1359
>BE998753 similar to PIR|T45958|T45 oligopeptide transporter-like protein -
Arabidopsis thaliana, partial (44%)
Length = 809
Score = 213 bits (543), Expect = 1e-55
Identities = 119/271 (43%), Positives = 169/271 (61%), Gaps = 7/271 (2%)
Frame = +3
Query: 250 PLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRAAYITS 309
P + V A+ RK V ++ LY I + ES+ +RK+ HT+ +F D+AA
Sbjct: 9 PPYSYRSVIVASIRKYRVDAPTDKSLLYEIADTESAIKGSRKLDHTNELRFFDKAAVQGE 188
Query: 310 RDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGAAMK 369
D + NPW LC +TQVEE+K ILRLLP+W II++ V+ QM++LFV QG M
Sbjct: 189 SDNLKES---INPWRLCTVTQVEELKSILRLLPVWATGIIFATVYGQMSTLFVLQGQTMN 359
Query: 370 TTI--SSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMGIGLV 427
T + SSFKIPPAS+S FD +SV ++ Y R++ P+ K GLT+LQRMG+GL
Sbjct: 360 THVGNSSFKIPPASLSIFDTISVIFWVPVYDRIIVPIARKFT-GHKNGLTQLQRMGVGLF 536
Query: 428 IAIIAMVTAGIVECYRLKYAKQGDTSSL-----SIFWQIPQYALIGASEVFMYVGQLEFF 482
I+I +MV A +E RL+ ++ + L +IFWQ+PQY LIG +EVF ++GQLEFF
Sbjct: 537 ISIFSMVAAAFLELVRLRTVRRNNYYELEEIPMTIFWQVPQYFLIGCAEVFTFIGQLEFF 716
Query: 483 NAQTPDGLKSFGSALCMTSISLGNYVSSLIV 513
Q PD ++S SAL + +++ G Y+SSL+V
Sbjct: 717 YEQAPDAMRSLCSALSLLTVAFGQYLSSLLV 809
>TC79608 similar to GP|20147231|gb|AAM10330.1 At1g68570/F24J5_7 {Arabidopsis
thaliana}, partial (59%)
Length = 1489
Score = 149 bits (376), Expect(2) = 2e-53
Identities = 92/263 (34%), Positives = 139/263 (51%), Gaps = 8/263 (3%)
Frame = +2
Query: 229 LALVLFLVGTPKYRHFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNS 288
++++ F+ G P YR+ P G+P +R +V AA RK + N LY DE ++S
Sbjct: 29 ISIIAFIGGYPLYRNLNPEGSPFTRLVKVGVAAFRKKKIPKVPNSTLLYQNDELDASITL 208
Query: 289 NRKILHTHGFKFLDRAAYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTI 348
K+LH+ KFLD+AA +T D + W L + +VEE+K I+R+ PIW I
Sbjct: 209 GGKLLHSDQLKFLDKAAIVTEED----NTKTPDLWRLSTVHRVEELKSIIRMGPIWASGI 376
Query: 349 IYSVVFTQMASLFVEQGAAMKTTIS-SFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGK 407
+ + Q + ++Q M ++ SF+IP SMS F IL++ Y RVL V +
Sbjct: 377 LLITAYAQQGTFSLQQAKTMNRHLTKSFEIPAGSMSVFTILTMLFTTALYDRVL-IRVAR 553
Query: 408 LKKSSSKGLTELQRMGIGLVIAIIAMVTAGIVECYRLKYAKQ-------GDTSSLSIFWQ 460
+G+T L RMGIG VI++ A AG VE R K A + + +S+FW
Sbjct: 554 RFTGLDRGITFLHRMGIGFVISLFATFVAGFVEMKRKKVAMEHGLIEHSSEIIPISVFWL 733
Query: 461 IPQYALIGASEVFMYVGQLEFFN 483
+PQY+L G +E FM +G + F+
Sbjct: 734 VPQYSLHGMAEAFMSIGAFKSFS 802
Score = 79.0 bits (193), Expect(2) = 2e-53
Identities = 41/107 (38%), Positives = 69/107 (64%), Gaps = 3/107 (2%)
Frame = +1
Query: 480 EFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIP-GNLNRGHL 538
EFF Q P+ + S A TSISLGNY+S+L+V++V K + + W+P NLN+G L
Sbjct: 793 EFFYDQAPESMTSTAMAFFWTSISLGNYISTLLVTLVHKFTKGPNGTNWLPDNNLNKGKL 972
Query: 539 DRFFFLLAVLTSLDLIAYIACAKW--FQNIQMACKYDNNDEPSSCKV 583
+ F++L+ +L ++LI Y+ CAK ++ IQ+ K +N+ E ++ ++
Sbjct: 973 EYFYWLITLLQFINLIYYLICAKMYTYKQIQVHHKGENSSEDNNIEL 1113
>TC83116 similar to GP|11933407|dbj|BAB19758. putative nitrate transporter
NRT1-3 {Glycine max}, partial (50%)
Length = 1192
Score = 201 bits (512), Expect = 5e-52
Identities = 97/277 (35%), Positives = 172/277 (62%), Gaps = 1/277 (0%)
Frame = +3
Query: 8 EELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDN 67
EE T DG+V+ G+P +R+ +G W A + +++ + +A++G+ NL+L T+ L Q
Sbjct: 72 EEYTEDGTVNLKGKPVLRSKTGGWKACSFVVVYEVFERMAYYGISSNLILLFTKKLHQGT 251
Query: 68 AAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRP 127
A NNV+ W GT+++ ++GA+++D++ GR+ T I I+++G+ L++ L L+P
Sbjct: 252 XTAXNNVTNWVGTIWMTPILGAYVADAHLGRFWTFLIASFIYLSGMSLLTLAVSLPTLKP 431
Query: 128 KGCGNGKL-ECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYSKV 186
C + +C + S+L++ +FY ++Y + +G GG +PNI+T GADQFD+ H KE K+
Sbjct: 432 PECHELXVTKCKKLSTLQLAVFYGALYTLXIGTGGTKPNISTIGADQFDDFHPKEKSHKL 611
Query: 187 AFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRHFKP 246
+FF+++ ++ G+LF+NT+L Y +D W LG+ +++++FL GTP YRH P
Sbjct: 612 SFFNWWMFSIFFGTLFANTVLVYIQDNVGWTLGYALPTLGLAISIMIFLGGTPFYRHKLP 791
Query: 247 CGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKE 283
G+ +R +V A+ RK V + + LY +D +E
Sbjct: 792 AGSTFTRMARVIVASLRKWKVPVPDDTKKLYELDMEE 902
>TC82936 similar to PIR|G84829|G84829 probable PTR2 family peptide
transporter [imported] - Arabidopsis thaliana, partial
(53%)
Length = 1112
Score = 200 bits (508), Expect = 1e-51
Identities = 121/353 (34%), Positives = 189/353 (53%), Gaps = 8/353 (2%)
Frame = +3
Query: 198 LGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRH-FKPCGNPLSRFCQ 256
LG+L + L Y ++ W LG+ L+LV+F +GTP YRH + +P +
Sbjct: 75 LGALIATLGLVYIQENLGWGLGYGIPTAGLILSLVIFYIGTPIYRHKVRTSKSPAKDIIR 254
Query: 257 VFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRAAYITSRDLEVQK 316
VF A + +Q+ SN +L+ + R++ HT +FLD+AA ++
Sbjct: 255 VFIVAFKSRKLQLPSNPSELHEFQMEHCVIRGKRQVYHTPTLRFLDKAAI---KEDPTTG 425
Query: 317 GGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGAAMKTTIS-SF 375
+ P + QVE K IL +L IWL T+I S ++ Q+ +LFV+QG + + F
Sbjct: 426 SSRRVPM---TVNQVEGAKLILGMLLIWLVTLIPSTIWAQINTLFVKQGTTLDRNLGPDF 596
Query: 376 KIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMGIGLVIAIIAMVT 435
KIP AS+ SF LS+ + + Y R+ P + + K +G+T LQR+GIG I IIA+
Sbjct: 597 KIPAASLGSFVTLSMLLSVPMYDRLFVPFM-RQKTGHPRGITLLQRLGIGFSIQIIAIAI 773
Query: 436 AGIVECYRLKYAKQG------DTSSLSIFWQIPQYALIGASEVFMYVGQLEFFNAQTPDG 489
A VE R+ K+ D +SIFW +PQY LIG ++VF +G LEFF Q+P+
Sbjct: 774 AYAVEVRRMHVIKENHIFGPKDIVPMSIFWLLPQYVLIGIADVFNAIGLLEFFYDQSPED 953
Query: 490 LKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGHLDRFF 542
++S G+ + I +GN+++S +V++ KI+ WI NLN HL ++
Sbjct: 954 MQSLGTTFFTSGIGVGNFLNSFLVTMTDKITGRGDRKSWIADNLNDSHLXYYY 1112
>TC93408 similar to GP|11933407|dbj|BAB19758. putative nitrate transporter
NRT1-3 {Glycine max}, partial (44%)
Length = 909
Score = 198 bits (504), Expect = 4e-51
Identities = 113/303 (37%), Positives = 176/303 (57%), Gaps = 8/303 (2%)
Frame = +1
Query: 203 SNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRHFKPCGNPLSRFCQVFFAAS 262
+NT+L Y +D W LG+ +++++FL GTP YRH P G+ +R +V A+
Sbjct: 1 ANTVLVYIQDNVGWTLGYALPTLGLAISIMIFLGGTPFYRHKLPAGSTFTRMARVIVASL 180
Query: 263 RKLGVQMTSNGDDLYVIDEKESSNNSNR--KILHTHGFKFLDRAAYITSRDLEVQKGGQH 320
RK V + + LY +D +E + + +I T +FLD+A+ K G
Sbjct: 181 RKWKVPVPDDTKKLYELDMEEYAKKGSNTYRIDSTPTLRFLDKASV---------KTGST 333
Query: 321 NPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGAAMKTTISSFKIPPA 380
+PW LC +T VEE K +LR++PI + T + S + Q+ +LFV+QG + I SFKIPPA
Sbjct: 334 SPWMLCTVTHVEETKQMLRMIPILVATFVPSTMMAQVNTLFVKQGTTLDRHIGSFKIPPA 513
Query: 381 SMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMGIGLVIAIIAMVTAGIVE 440
S+++F LS+ + + Y R ++ +L K + +G+T LQRMGIGLV+ I MV A + E
Sbjct: 514 SLAAFVTLSLLVCVVLYDRFFVRIMQRLTK-NPRGITLLQRMGIGLVLHTIIMVVASVTE 690
Query: 441 CYRLKYAKQ------GDTSSLSIFWQIPQYALIGASEVFMYVGQLEFFNAQTPDGLKSFG 494
YRL+ AK+ G LSIF +PQ+ L+G ++ F+ V ++EFF Q P +KS G
Sbjct: 691 NYRLRVAKEHGLVESGGQVPLSIFILLPQFILMGTADAFLEVAKIEFFYDQAPXSMKSIG 870
Query: 495 SAL 497
+++
Sbjct: 871 TSI 879
>TC86049 weakly similar to GP|20466248|gb|AAM20441.1 putative transport
protein {Arabidopsis thaliana}, partial (43%)
Length = 1836
Score = 195 bits (496), Expect = 4e-50
Identities = 124/419 (29%), Positives = 216/419 (50%), Gaps = 10/419 (2%)
Frame = +2
Query: 152 IYLIALGNGGYQPNIATFGADQFDEDHSKESYSKVA-FFSYFYLALNLGSLFSNTILGYF 210
+ LIA+GNGG +++ FGA Q + + + Y + FFS++Y + + + T + Y
Sbjct: 2 LILIAIGNGGITCSLS-FGAYQVNRKDNPDGYRVLEIFFSWYYAFTTIAVIIALTGIVYI 178
Query: 211 EDEGLWALGFWASAGSAFLALVLFLVGTPKYRHFKPCGNPLSRFCQVFFAASRKLGVQMT 270
+D W +GF A ++ VLF + +P Y K + + F QV AA + + +
Sbjct: 179 QDHLGWKVGFGVPAILMLISTVLFFLASPLYVKIKQKTSLFTGFAQVSVAAYKNRKLPLP 358
Query: 271 SNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRAAYITSRDLEVQKGGQH-NPWYLCPIT 329
+ +K+S ++ T +FL++A I + ++ G N W LC +
Sbjct: 359 PKTSPEFYHQQKDSE-----LVVPTDKLRFLNKACVIKDHEQDIASDGSAINRWSLCTVD 523
Query: 330 QVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGAAMKTTISS-FKIPPASMSSFDIL 388
QVEE+K I++++P+W I S+ L + SS F++P S S I+
Sbjct: 524 QVEELKAIIKVIPLWSTAITMSINIGGSFGLLQAKSLDRHIISSSNFEVPAGSFSVILIV 703
Query: 389 SVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMGIGLVIAIIAMVTAGIVECYRLKYA- 447
++ I+I Y RVL PL K++ ++ +RMGIGL + ++TA I E R K A
Sbjct: 704 AILIWIIIYDRVLIPLASKIR-GKPVIISPKKRMGIGLFFNFLHLITAAIFETVRRKEAI 880
Query: 448 KQG---DTSS---LSIFWQIPQYALIGASEVFMYVGQLEFFNAQTPDGLKSFGSALCMTS 501
K+G DT +S W PQ L G +E+F +GQ EF+ + P + S ++L +
Sbjct: 881 KEGYLNDTHGVLKMSAMWLAPQLCLAGIAEMFNVIGQNEFYYKEFPKSMSSVAASLSGLA 1060
Query: 502 ISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGHLDRFFFLLAVLTSLDLIAYIACA 560
+ +GN VSSL++SI+ + GW+ N+N+GH D++++++ + +L+L+ Y+ C+
Sbjct: 1061MGVGNLVSSLVLSIIESTTPSGGNEGWVSDNINKGHFDKYYWVIVGINALNLLYYLVCS 1237
>BM780227 similar to PIR|E84798|E84 probable peptide/amino acid transporter
[imported] - Arabidopsis thaliana, partial (36%)
Length = 750
Score = 166 bits (420), Expect = 2e-41
Identities = 91/254 (35%), Positives = 145/254 (56%)
Frame = +3
Query: 54 NLVLFLTRVLGQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGL 113
+LVL+L V+ QD AA NV+ W G + L G F++D+Y GRY +++ GL
Sbjct: 3 SLVLYLILVIHQDLKTAARNVNYWAGVTTLMPLFGGFIADAYLGRYSAVVASSIVYLMGL 182
Query: 114 VSLSVTTYLALLRPKGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQ 173
+ L+++ +L L+P C + C E + +F+L+IYLI++ GG++P++ +FGADQ
Sbjct: 183 ILLTLSWFLPSLKP--CDH-TTTCNEPRKIHEVVFFLAIYLISIATGGHKPSLESFGADQ 353
Query: 174 FDEDHSKESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVL 233
FDEDH +E K++FF+++ AL G + T++ Y +D W G L+L++
Sbjct: 354 FDEDHVEERKQKMSFFNWWNCALCSGLILGVTLIVYIQDNINWGAADIIFTGVMALSLLI 533
Query: 234 FLVGTPKYRHFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKIL 293
F++G P YR+ P G+PL+ QV AA K + SN D LY + +S N + +
Sbjct: 534 FIIGRPFYRYRVPSGSPLTPMLQVLVAAFSKRKLPYPSNPDQLYEV--SKSHGNKRKFLC 707
Query: 294 HTHGFKFLDRAAYI 307
HT +FL +AA I
Sbjct: 708 HTKKLRFL*QAAII 749
>BE997589 weakly similar to PIR|G86449|G86 hypothetical protein AAF81343.1
[imported] - Arabidopsis thaliana, partial (17%)
Length = 517
Score = 152 bits (383), Expect = 4e-37
Identities = 77/144 (53%), Positives = 99/144 (68%)
Frame = +2
Query: 156 ALGNGGYQPNIATFGADQFDEDHSKESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGL 215
A G GG+QP +ATFGADQ+DE + KE KVAFF YFY +LN+GSLFSNT+L Y+ED G
Sbjct: 2 AFGYGGHQPTLATFGADQYDERNPKERSLKVAFFCYFYFSLNVGSLFSNTVLVYYEDTGK 181
Query: 216 WALGFWASAGSAFLALVLFLVGTPKYRHFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDD 275
W +GF+ S SA +AL+ FL G+PKYR+ KP GNP+ R QVF AA+RK V + D
Sbjct: 182 WTMGFFISLISAIIALLTFLSGSPKYRYLKPSGNPVVRVAQVFTAAARKWDV-APAKADK 358
Query: 276 LYVIDEKESSNNSNRKILHTHGFK 299
L+ + S+ RKILH+ F+
Sbjct: 359 LFEVLGSRSAIKGCRKILHSDDFR 430
>AW694006 similar to PIR|F86358|F86 Similar to peptide transporter [imported]
- Arabidopsis thaliana, partial (33%)
Length = 641
Score = 148 bits (374), Expect = 5e-36
Identities = 77/220 (35%), Positives = 129/220 (58%)
Frame = +1
Query: 16 VDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDNAAAANNVS 75
VD+ G+P++R+ SG W + I+ + +++ G+ NL+ +LT L Q A AA NV+
Sbjct: 1 VDYRGQPAVRSKSGYWRSAWFIIGVEVAERISYIGIKGNLISYLTGPLKQSTATAAKNVN 180
Query: 76 KWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRPKGCGNGKL 135
W GT + L+GAF++DS+ GRY T + I++ GL L+++T L L
Sbjct: 181 IWAGTASLLPLLGAFVADSFLGRYHTIILASLIYILGLGLLTLSTILPSL---------A 333
Query: 136 ECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYSKVAFFSYFYLA 195
C S ++ +F++S+YL+A+G GG++P + FGADQFDE + +E ++ +FF+++Y
Sbjct: 334 SCSTQS--QVILFFISLYLVAIGQGGHKPCVQAFGADQFDEKYPEEHRARSSFFNWWYFT 507
Query: 196 LNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFL 235
+ G+ + IL Y +D W LGF +AL++FL
Sbjct: 508 MVAGATATLPILTYIQDNYSWVLGFGIPCVIMIIALIIFL 627
>TC85038 weakly similar to PIR|G86449|G86449 hypothetical protein AAF81343.1
[imported] - Arabidopsis thaliana, partial (14%)
Length = 652
Score = 146 bits (368), Expect = 2e-35
Identities = 70/127 (55%), Positives = 92/127 (72%)
Frame = +1
Query: 4 KEETEELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVL 63
+E T DGSVD++G+P+++ +SGRW + T++L+NQGL +AF GV NLV+F VL
Sbjct: 226 EEGINSCTKDGSVDFYGKPALKNSSGRWRSATLLLVNQGLVAVAFAGVEANLVIFCKLVL 405
Query: 64 GQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLA 123
Q N AAN S W GT Y FSL+GAFLSDSY GRY TC IFQ +F+ GLV+LS++T+L
Sbjct: 406 KQTNVEAANTFSIWMGTTYFFSLIGAFLSDSYLGRYLTCIIFQLVFIIGLVALSLSTHLF 585
Query: 124 LLRPKGC 130
LL+P C
Sbjct: 586 LLKPHRC 606
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.139 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,364,170
Number of Sequences: 36976
Number of extensions: 285552
Number of successful extensions: 1959
Number of sequences better than 10.0: 106
Number of HSP's better than 10.0 without gapping: 1858
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1885
length of query: 583
length of database: 9,014,727
effective HSP length: 101
effective length of query: 482
effective length of database: 5,280,151
effective search space: 2545032782
effective search space used: 2545032782
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147404.4


