

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147404.2 + phase: 0
(599 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
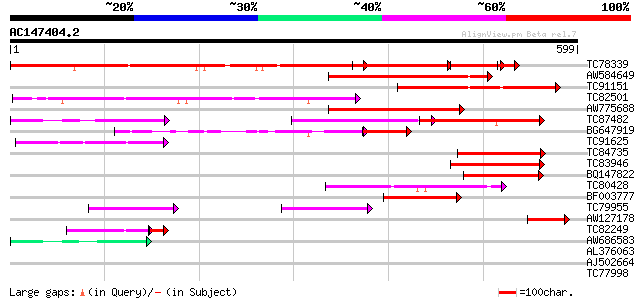
Score E
Sequences producing significant alignments: (bits) Value
TC78339 similar to PIR|T51937|T51937 probable inositol-1 4 5-tri... 376 e-104
AW584649 similar to GP|12323727|gb putative inositol polyphospha... 228 7e-60
TC91151 weakly similar to PIR|C86465|C86465 probable inositol po... 180 1e-45
TC82501 similar to PIR|D96739|D96739 hypothetical protein F14O23... 154 9e-38
AW775688 similar to PIR|E84430|E844 probable inositol polyphosph... 138 7e-33
TC87482 similar to PIR|E84430|E84430 probable inositol polyphosp... 123 2e-28
BG647919 similar to GP|20161480|db putative inositol-1 4 5-tri... 71 1e-27
TC91625 similar to PIR|D96739|D96739 hypothetical protein F14O23... 119 3e-27
TC84735 similar to GP|10178037|dbj|BAB11520. contains similarity... 110 2e-24
TC83946 similar to PIR|E84430|E84430 probable inositol polyphosp... 106 2e-23
BQ147822 similar to PIR|T48113|T481 inositol-1 4 5-trisphosphate... 102 5e-22
TC80428 similar to GP|4688596|emb|CAB41466.1 inositol 1 4 5-tris... 100 1e-21
BF003777 similar to GP|12323727|gb| putative inositol polyphosph... 86 5e-17
TC79955 similar to PIR|E84430|E84430 probable inositol polyphosp... 75 7e-14
AW127178 similar to GP|10444263|gb| inositol polyphosphate 5-pho... 64 2e-10
TC82249 similar to PIR|H84727|H84727 probable inositol polyphosp... 59 4e-10
AW686583 similar to PIR|E84430|E844 probable inositol polyphosph... 59 5e-09
AL376063 weakly similar to PIR|H85044|H85 hypothetical protein A... 33 0.41
AJ502664 32 0.91
TC77998 similar to PIR|T09254|T09254 isoflavone-7-O-methyltransf... 31 1.2
>TC78339 similar to PIR|T51937|T51937 probable inositol-1 4 5-trisphosphate
5-phosphatase (EC 3.1.3.56) At5P2 [imported] -
Arabidopsis, partial (53%)
Length = 1762
Score = 376 bits (966), Expect = e-104
Identities = 223/416 (53%), Positives = 273/416 (65%), Gaps = 38/416 (9%)
Frame = +1
Query: 2 KGKRSEVFWPSTVMKKWLNVKQKVYDFSEDEANTETDESEDDVKLIEIETNHTSKPQANT 61
+GKRSE FWPS VMKKWLN+K KV DFSEDE +TET ESEDDV + S
Sbjct: 52 RGKRSEAFWPSIVMKKWLNIKPKVNDFSEDEVDTET-ESEDDVCSPKQPRMQISDDSPFR 228
Query: 62 TKPTQL-------NTSCKDYEMKRHRRRKSETLRAQYINTKEVRVTIGTWNVAGKHPCND 114
T+ TQ +TS K RHRR KSETLRAQYINTKEVRV IG+WNVAG+HP D
Sbjct: 229 TQGTQSIFSSQISDTSFKKGCKTRHRRGKSETLRAQYINTKEVRVAIGSWNVAGRHPSED 408
Query: 115 LEIEGWLCTEEEPSDIYIIGFQEVVPLNAGNVFGAEDSKPIPKWDALIRRTLNKSSEPGT 174
L+I+ W+ EE PSDIYI GFQEVVPLNAGNV GAED+ PI KW+A+IRR+LNKSSEP +
Sbjct: 409 LDIDDWIYAEE-PSDIYIFGFQEVVPLNAGNVLGAEDNTPIQKWEAIIRRSLNKSSEPDS 585
Query: 175 KKKSNSAPPSPIRRISSFNTQI-------NPLDSALD-----------KKEEIKTIISIE 216
K KS+SAPPSP+ R SS + NP+D D ++ E IISI
Sbjct: 586 KHKSHSAPPSPVLRTSSAADVLADNIDAANPIDMLNDELMENVDKYDLQQLEESNIISIG 765
Query: 217 KNLQLSKIYDIDLQTILDWPELRLDPIHH-VDSSPKMRRVQSTS--------DSASLY-- 265
+L + K+Y ID LDWPE LD I VDS+PK+RRV S+S ++A LY
Sbjct: 766 NDLHVRKVYGID----LDWPERPLDAISQIVDSNPKLRRVLSSSARIGFDLNENAFLYGG 933
Query: 266 --GFEMKSLHQSSGNFSLLWSEKQQEIVPQVFDSHLDVSDMLSDEDNDTFSELANNEDAN 323
G +K H SSGN L K+Q+++P+V DS DVS+MLSD+ +D F EL N+D +
Sbjct: 934 GGGGGLKRTHHSSGNLGSLL--KEQQVIPKVVDSLDDVSEMLSDDGDDAFIELPENQDDD 1107
Query: 324 GIISVKSHPKYVRIVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKG 379
+ + KS +YVRI+SKQMVGIYVSVWVQR+LRRH+++LKVS VG G G +G
Sbjct: 1108ELGTTKSQARYVRIISKQMVGIYVSVWVQRRLRRHINNLKVSPVGSWSYGLYGKQG 1275
Score = 169 bits (429), Expect(3) = 2e-72
Identities = 83/105 (79%), Positives = 91/105 (86%)
Frame = +2
Query: 363 KVSQVGVGLMGYMGNKGSVSVSMSVFQSRMCFVCSHLASGQKDGAEQRRNSDVHEILQRT 422
K +GVGLMGYMGNKGSVS+SMS+FQSRMCFVCSHL SG KDGAEQRRNSDV+EIL+RT
Sbjct: 1226 KFHLLGVGLMGYMGNKGSVSISMSLFQSRMCFVCSHLTSGTKDGAEQRRNSDVNEILRRT 1405
Query: 423 RFSSVFDTDQPQKIPSHDKIFWFGDLNYRINMSDGEIRKLVDLKK 467
FSSVF TDQ IPSHD+IFWFGDLNYRI+M D E+RKLV KK
Sbjct: 1406 CFSSVFATDQALTIPSHDQIFWFGDLNYRISMLDSEVRKLVAQKK 1540
Score = 97.8 bits (242), Expect(3) = 2e-72
Identities = 43/58 (74%), Positives = 50/58 (86%)
Frame = +3
Query: 466 KKWNELMKFDQLSNELCKGHVFEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGEKRRA 523
KKWNEL+ +DQLSNEL GHVF+GWKEGLINF PTYKYE NSD++VG +EGEK+RA
Sbjct: 1536 KKWNELLNYDQLSNELRVGHVFDGWKEGLINFAPTYKYEINSDRYVGDIPKEGEKKRA 1709
Score = 45.1 bits (105), Expect(3) = 2e-72
Identities = 17/23 (73%), Positives = 20/23 (86%)
Frame = +1
Query: 516 QEGEKRRAPAWCDRILWLGKGIK 538
++G K+ PAWCDRILWLGKGIK
Sbjct: 1687 KKGRKKEPPAWCDRILWLGKGIK 1755
>AW584649 similar to GP|12323727|gb putative inositol polyphosphate
phosphatase 5' partial; 1-2276 {Arabidopsis thaliana},
partial (28%)
Length = 517
Score = 228 bits (580), Expect = 7e-60
Identities = 105/173 (60%), Positives = 137/173 (78%)
Frame = +1
Query: 338 VSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSRMCFVCS 397
VSKQMVGI+++VWV+R LR+H+H+LKVS VGVG+MGY+GNKGSVSVSMS++Q+ CF+C+
Sbjct: 1 VSKQMVGIFITVWVRRSLRKHIHNLKVSTVGVGIMGYIGNKGSVSVSMSIYQTLFCFICT 180
Query: 398 HLASGQKDGAEQRRNSDVHEILQRTRFSSVFDTDQPQKIPSHDKIFWFGDLNYRINMSDG 457
HL SG+K+G E +RNSDVHEIL+RT F S P+ I H++I W GDLNYRIN+S+
Sbjct: 181 HLTSGEKEGDELKRNSDVHEILRRTHFHSPSIIGLPKGILDHERIIWLGDLNYRINLSNV 360
Query: 458 EIRKLVDLKKWNELMKFDQLSNELCKGHVFEGWKEGLINFPPTYKYEFNSDKH 510
E + L+ K+W++L++ DQL EL G F GW EG +NFPPTYKYE NSDK+
Sbjct: 361 EAKALISKKQWSKLLEKDQLMRELKHG-AFGGWSEGALNFPPTYKYEVNSDKY 516
>TC91151 weakly similar to PIR|C86465|C86465 probable inositol polyphosphate
5-phosphatase [imported] - Arabidopsis thaliana, partial
(27%)
Length = 921
Score = 180 bits (457), Expect = 1e-45
Identities = 86/173 (49%), Positives = 123/173 (70%)
Frame = +2
Query: 410 RRNSDVHEILQRTRFSSVFDTDQPQKIPSHDKIFWFGDLNYRINMSDGEIRKLVDLKKWN 469
+RN+DV EI QRT F S+ D P+ I H++I WFGDLNYRI++ + R L+ K W+
Sbjct: 5 KRNADVREIHQRTHFYSLSDFGLPKSILDHERIIWFGDLNYRISLPYDKTRDLISKKHWS 184
Query: 470 ELMKFDQLSNELCKGHVFEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGEKRRAPAWCDR 529
+L++ DQL+ EL KG VF+GW EG +NFPPTYKYE NSDK++G + + RR PAWCDR
Sbjct: 185 KLVERDQLAKELEKG-VFDGWSEGKLNFPPTYKYEINSDKYIGEDPKVA--RRTPAWCDR 355
Query: 530 ILWLGKGIKQLKYQSAENQLSDHRPVSSIFLVDVEVIDHRKLERAIYFASAVV 582
IL G G++ L Y+ +E + SDHRPV++ ++ +VEV + +KL+RA+ + A +
Sbjct: 356 ILSHGNGMRLLSYKRSELKFSDHRPVTATYMAEVEVFNPKKLQRALNYTDAEI 514
>TC82501 similar to PIR|D96739|D96739 hypothetical protein F14O23.9
[imported] - Arabidopsis thaliana, partial (28%)
Length = 1406
Score = 154 bits (389), Expect = 9e-38
Identities = 123/410 (30%), Positives = 199/410 (48%), Gaps = 43/410 (10%)
Frame = +3
Query: 4 KRSEVFWPSTVMKKWLNVKQKVYDFSEDEANTETDESEDDVKLIEIETNHT---SKPQAN 60
K+ + W + VM+KWLN+K+K D+ +T+ D+ +DDV E ++++ S+ +
Sbjct: 210 KQQQNLWATMVMRKWLNIKRK----ESDDYSTDPDD-DDDVDDPETDSDNEEWGSRSRIR 374
Query: 61 TTKPTQLNTSCKDYEMKRHRRRKSETLRAQYINTKEVRVTIGTWNVAGKHPCNDLEIEGW 120
+ + ++ + RR+KS T+R+QYIN KE+RV +GTWNV GK P ND +I+ W
Sbjct: 375 DRREDEAPAESDEF-LPGLRRQKSLTVRSQYINKKELRVCVGTWNVGGKLPPNDFDIDDW 551
Query: 121 LCTEEEPSDIYIIGFQEVVPLNAGNVFGAE-DSKPIPK-WDALIRRTLNKSSEPGTKKK- 177
L P+DIY++G QE+VPLN N+F +P+P+ + LIR L SS K K
Sbjct: 552 L-DINHPADIYVLGLQEIVPLNTSNIFWLL*IPRPVPEVGENLIREALK*SSIKTFKDKI 728
Query: 178 ------SNSAPPSP------------------IRRISSFNTQINPLDSALDKKEEIKTII 213
N + +RR I L + K +K +
Sbjct: 729 LLVILHLNQSLNHQMMHQI*KKKFY*KVIVILVRRFIP*VKNIMYLMESQINKS*MKHLN 908
Query: 214 SIEKNLQLSKIYDIDLQTILDWPELRLDPIHHVDSSPKMRRVQSTSDSASLYGFEMKSLH 273
K+ S I + DLQ L + + +L+ ++H ++TS S ++ +
Sbjct: 909 ISLKDSNASDIAENDLQNQLSY-QRKLNRLNHFREEDSSENNETTS---SQQISKLSRMV 1076
Query: 274 QSSGNFSLLWSEKQQEIVPQ-VFDSHLDVSDMLSDEDNDTFS------------ELANNE 320
+ L W E ++PQ V + + S +F +L +
Sbjct: 1077SGTERIGLSWPEPPLHLLPQKVLERPTSFKPVKSFNKTKSFRACDSFKSKTDAIDLLADI 1256
Query: 321 DANGIISVKSHPKYVRIVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVG 370
D ++ K+ YV+IVSKQMVGI+++VWV+R LR+H+ ++KVS VGVG
Sbjct: 1257DLEALMKRKTRSSYVKIVSKQMVGIFITVWVRRSLRKHIQNMKVSTVGVG 1406
>AW775688 similar to PIR|E84430|E844 probable inositol
polyphosphate-5-phosphatase [imported] - Arabidopsis
thaliana, partial (21%)
Length = 581
Score = 138 bits (347), Expect = 7e-33
Identities = 61/144 (42%), Positives = 97/144 (67%)
Frame = +2
Query: 337 IVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSRMCFVC 396
I+S+Q+VG+++++W + L + + HL VS VG G++G + NKGS+S+ + ++ CF+C
Sbjct: 149 IISRQLVGMFITIWARCDLYQSIKHLNVSSVGCGVLGCLANKGSISIRFFLHETSFCFIC 328
Query: 397 SHLASGQKDGAEQRRNSDVHEILQRTRFSSVFDTDQPQKIPSHDKIFWFGDLNYRINMSD 456
SHLASG K+ +++RN++ +IL +T F D PQKI HD++ W GDLNYRI+MS
Sbjct: 329 SHLASGGKEEDKRQRNANAADILSQTNFPVGPLHDLPQKIIDHDRVVWLGDLNYRIDMSH 508
Query: 457 GEIRKLVDLKKWNELMKFDQLSNE 480
+ L+ ++W L+K DQL E
Sbjct: 509 SATQSLIKKREWETLLKHDQLKME 580
>TC87482 similar to PIR|E84430|E84430 probable inositol
polyphosphate-5-phosphatase [imported] - Arabidopsis
thaliana, partial (72%)
Length = 1567
Score = 123 bits (308), Expect = 2e-28
Identities = 59/136 (43%), Positives = 86/136 (62%), Gaps = 4/136 (2%)
Frame = +1
Query: 434 QKIPSHDKIFWFGDLNYRINMSDGEI-RKLVDLKKWNELMKFDQLSNELCKGHVFEGWKE 492
+KI HD + G + + E R LV+ + W+ L++ DQL EL G + GW E
Sbjct: 1021 EKILDHDHVILLGRFKLQEYLYQKETTRLLVEKRDWDSLLENDQLKMELESGQMLRGWHE 1200
Query: 493 GLINFPPTYKYEFNSDKHVG---GNTQEGEKRRAPAWCDRILWLGKGIKQLKYQSAENQL 549
G I F PTYKY NSD++ G ++ K+R+PAWCDRI+WLG G+KQ++Y +E++L
Sbjct: 1201 GTIKFAPTYKYFLNSDEYYGCCYHGMKKAAKKRSPAWCDRIIWLGNGLKQIEYARSESKL 1380
Query: 550 SDHRPVSSIFLVDVEV 565
SDHRPV ++F +V+V
Sbjct: 1381 SDHRPVKALFTAEVKV 1428
Score = 109 bits (272), Expect = 3e-24
Identities = 68/158 (43%), Positives = 94/158 (59%), Gaps = 4/158 (2%)
Frame = +2
Query: 298 HLDVSDMLSDEDNDTFSELANNEDAN-GIISVKSHPK-YVRIVSKQMVGIYVSVWVQRKL 355
H D+ D+D++ +L N N KS P+ + I+SKQMVGI +SVWV+ L
Sbjct: 605 HYCCKDIERDDDDNIEQDLKNICQGNIPAQQCKSAPQDFHCIISKQMVGILISVWVRSDL 784
Query: 356 RRHVHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSRMCFVCSHLASGQKDGAEQRRNSDV 415
+ H VS VG G+MG +GNKGSVSV + ++ CFVCSHLASG K+G E+ RNS+V
Sbjct: 785 SPFIRHPCVSCVGCGIMGCLGNKGSVSVRFLLHETSFCFVCSHLASGGKEGDEKHRNSNV 964
Query: 416 HEILQRTRF--SSVFDTDQPQKIPSHDKIFWFGDLNYR 451
EI RT F ++F+ + ++ + GDLNYR
Sbjct: 965 AEIFSRTSFPKGTIFEFTK-KRFLIMIM*YCLGDLNYR 1075
Score = 86.7 bits (213), Expect = 2e-17
Identities = 57/168 (33%), Positives = 78/168 (45%)
Frame = +2
Query: 1 MKGKRSEVFWPSTVMKKWLNVKQKVYDFSEDEANTETDESEDDVKLIEIETNHTSKPQAN 60
M K +EV WP+ V K LN + +F D + + P +
Sbjct: 191 MTRKLAEVMWPALVANKILNRRLGSRNFIADYPSY-------------------TDPLLS 313
Query: 61 TTKPTQLNTSCKDYEMKRHRRRKSETLRAQYINTKEVRVTIGTWNVAGKHPCNDLEIEGW 120
T QL+ S K + +T++ +V + TWN+ G P L IE
Sbjct: 314 TNNDDQLSLSTKSIN--------------DHSDTQKYKVFVSTWNIGGIAPDEGLNIEDL 451
Query: 121 LCTEEEPSDIYIIGFQEVVPLNAGNVFGAEDSKPIPKWDALIRRTLNK 168
L T + DIY+ GFQE+VPLNA NV G+EDSK KW++LIR LNK
Sbjct: 452 LETCSKSFDIYVFGFQEIVPLNASNVLGSEDSKISTKWNSLIRNALNK 595
>BG647919 similar to GP|20161480|db putative inositol-1 4 5-trisphosphate
5-phosphatase {Oryza sativa (japonica cultivar-group)},
partial (29%)
Length = 775
Score = 71.2 bits (173), Expect(2) = 1e-27
Identities = 69/275 (25%), Positives = 123/275 (44%), Gaps = 6/275 (2%)
Frame = +3
Query: 111 PCNDLEIEGWLCTEEEPSDIYIIGFQEVVPLNAGNVFGAEDSKPIPKWDALIRRTLNKSS 170
P +L++ +L EP D+Y++GFQE+VPLNAGNV ED +
Sbjct: 3 PSGNLDLSDFLQVRNEP-DMYVLGFQEIVPLNAGNVLVLED------------------N 125
Query: 171 EPGTKKKSNSAPPSPIRRISSFNTQIN-PLDSALDKKEEIKTIISIEKNLQLSKIYDIDL 229
EP K ++ N +N P D + +K +K S +L K
Sbjct: 126 EPAAKW------------LALINQSLNGPSDFSSNKG--LKPTASFGGSLYFQK------ 245
Query: 230 QTILDWPELRLDPIHHVDSSPKMRRVQSTSDSASLYGFEMKSLHQSSGNFSLLWSEKQ-- 287
P +++++ T L G +KS + +L E++
Sbjct: 246 --------------------PSLKKIKKTFKK--LNGKRLKSCN------CILEMERKAA 341
Query: 288 QEIVPQVFDSHLDVSDMLSDEDNDTFS---ELANNEDANGIISVKSHPKYVRIVSKQMVG 344
++ + +S++++ D ++E++D++ LA N+ KY + KQMVG
Sbjct: 342 KDFCFRCQESNVNLDDSSTEEEDDSYPISVALATNQ-----------MKYSLVTCKQMVG 488
Query: 345 IYVSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKG 379
I+VSVW++++L ++V HL++ G+MG +G +G
Sbjct: 489 IFVSVWMKKELIQYVGHLRICCTSRGIMGCLGTRG 593
Score = 70.5 bits (171), Expect(2) = 1e-27
Identities = 32/52 (61%), Positives = 41/52 (78%)
Frame = +2
Query: 373 GYMGNKGSVSVSMSVFQSRMCFVCSHLASGQKDGAEQRRNSDVHEILQRTRF 424
G NKG +SVSMS +Q++ CF+CSHLASG+K+G E RRN DV EIL+ T+F
Sbjct: 572 GMPRNKGCISVSMSFYQTKFCFICSHLASGEKEGDELRRNLDVIEILKNTQF 727
>TC91625 similar to PIR|D96739|D96739 hypothetical protein F14O23.9
[imported] - Arabidopsis thaliana, partial (12%)
Length = 621
Score = 119 bits (299), Expect = 3e-27
Identities = 65/161 (40%), Positives = 94/161 (58%)
Frame = +3
Query: 7 EVFWPSTVMKKWLNVKQKVYDFSEDEANTETDESEDDVKLIEIETNHTSKPQANTTKPTQ 66
++FW VM+KWLN+ D+S D + + + +DD E E K + + +
Sbjct: 147 QLFWARVVMRKWLNIGSNESDYSADPEDDDEFDEDDDEDDDEHEV-WGRKSRFMDNRGFE 323
Query: 67 LNTSCKDYEMKRHRRRKSETLRAQYINTKEVRVTIGTWNVAGKHPCNDLEIEGWLCTEEE 126
+ D+ K R++KS T R+QYINTKE+RV +GTWNV G+ P +DL+I+ WL E
Sbjct: 324 APSESNDFVPKL-RKQKSSTYRSQYINTKELRVCVGTWNVGGRLPPDDLDIDEWLGV-NE 497
Query: 127 PSDIYIIGFQEVVPLNAGNVFGAEDSKPIPKWDALIRRTLN 167
P+DIY++G QE+VPLNAGN+F KW +I LN
Sbjct: 498 PADIYVLGLQEIVPLNAGNIFWF*RYSSGSKWXNIIXEALN 620
>TC84735 similar to GP|10178037|dbj|BAB11520. contains similarity to
inositol polyphosphate 5'-phosphatase~gene_id:MUG13.16
{Arabidopsis thaliana}, partial (24%)
Length = 630
Score = 110 bits (274), Expect = 2e-24
Identities = 54/94 (57%), Positives = 64/94 (67%), Gaps = 1/94 (1%)
Frame = +2
Query: 474 FDQLSNELCKGHVFEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGE-KRRAPAWCDRILW 532
F QL E G VF+GWKEG I F PTYKY FNSD + + + KRR PAWCDRILW
Sbjct: 50 FSQLKMEREAGRVFKGWKEGKIYFAPTYKYAFNSDTYYAEGVKVSKNKRRTPAWCDRILW 229
Query: 533 LGKGIKQLKYQSAENQLSDHRPVSSIFLVDVEVI 566
G+GI+QL Y E + SDHRPV + FLV+VEV+
Sbjct: 230 HGRGIQQLSYVRKEFKFSDHRPVCATFLVEVEVM 331
>TC83946 similar to PIR|E84430|E84430 probable inositol
polyphosphate-5-phosphatase [imported] - Arabidopsis
thaliana, partial (11%)
Length = 652
Score = 106 bits (265), Expect = 2e-23
Identities = 54/102 (52%), Positives = 69/102 (66%), Gaps = 2/102 (1%)
Frame = +2
Query: 466 KKWNELMKFDQLSNELCKGHVFEGWKEGLINFPPTYKYEFNSDKHVGGNT--QEGEKRRA 523
K+ L++ DQL EL G+ GW EG I F PTYKY NSD + G ++ EKRRA
Sbjct: 5 KRVGSLLENDQLMMELMNGNNLRGWHEGPIKFAPTYKYCPNSDIYYGCCCPGKKIEKRRA 184
Query: 524 PAWCDRILWLGKGIKQLKYQSAENQLSDHRPVSSIFLVDVEV 565
PAWCDRI+W GK +KQL+Y +E++LSDHRPV +IF +V V
Sbjct: 185 PAWCDRIVWYGKDLKQLEYTRSESKLSDHRPVKAIFTAEVRV 310
>BQ147822 similar to PIR|T48113|T481 inositol-1 4 5-trisphosphate
5-Phosphatase-like protein - Arabidopsis thaliana,
partial (14%)
Length = 343
Score = 102 bits (253), Expect = 5e-22
Identities = 47/85 (55%), Positives = 54/85 (63%)
Frame = +2
Query: 480 ELCKGHVFEGWKEGLINFPPTYKYEFNSDKHVGGNTQEGEKRRAPAWCDRILWLGKGIKQ 539
E K VF GW EG I FPPTYKY NS + G + + +KR PAWCDRILW G G++Q
Sbjct: 86 ETGKAXVFTGWSEGKIYFPPTYKYSNNSYSYAGDDRRSKQKRXTPAWCDRILWXGSGLQQ 265
Query: 540 LKYQSAENQLSDHRPVSSIFLVDVE 564
L Y E+ SDH PV SIFL VE
Sbjct: 266 LSYVPGESXFSDHTPVCSIFLAXVE 340
>TC80428 similar to GP|4688596|emb|CAB41466.1 inositol 1 4 5-trisphosphate
5-phosphatase {Arabidopsis thaliana}, partial (22%)
Length = 757
Score = 100 bits (250), Expect = 1e-21
Identities = 71/216 (32%), Positives = 109/216 (49%), Gaps = 24/216 (11%)
Frame = +2
Query: 334 YVRIVSKQMVGIYVSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKGSVSVSMSVFQSRMC 393
+ RI S+Q+ G+ ++VWV+ + HV + + V G +GNKG+V++ + V+ MC
Sbjct: 128 FKRIGSRQLAGLVIAVWVKTNITLHVGDVDAAAVPCGFGRAIGNKGAVALRVRVYDRIMC 307
Query: 394 FVCSHLASGQKDGAEQRRNSDVHEILQRTRFSSVFD------------------TDQPQK 435
FV H A+ A RRNSD + + FS + T+ +
Sbjct: 308 FVNCHFAAHL--DAVGRRNSDFDYVYRTMSFSRPTNLLNTTPAGTSASIPMFRGTNPAEG 481
Query: 436 IP---SHDKIFWFGDLNYRI-NMSDGEIRKLVDLKKWNELMKFDQLSNELCKGHVFEGWK 491
IP D I + GDLNYR+ ++S E R V + ++ L + DQL E+ G+ F+G +
Sbjct: 482 IPELSEADMIVFLGDLNYRLDDISYDEARDFVSQRCFDWLRERDQLRAEMEAGNAFQGMR 661
Query: 492 EGLINFPPTYKYEFNSDKHVGG--NTQEGEKRRAPA 525
E +I FPPTYK+E +H G GEK+ PA
Sbjct: 662 EAVITFPPTYKFE----RHQAGLAGYDSGEKKXIPA 757
>BF003777 similar to GP|12323727|gb| putative inositol polyphosphate
phosphatase 5' partial; 1-2276 {Arabidopsis thaliana},
partial (14%)
Length = 487
Score = 85.5 bits (210), Expect = 5e-17
Identities = 38/82 (46%), Positives = 57/82 (69%)
Frame = +1
Query: 396 CSHLASGQKDGAEQRRNSDVHEILQRTRFSSVFDTDQPQKIPSHDKIFWFGDLNYRINMS 455
C+HL +G+K+ E +RN+DV EI QRT F S+ D P+ I H++I WFGDLNYRI++
Sbjct: 1 CTHLTAGEKEADEIKRNADVREIHQRTHFYSLSDFGLPKSILDHERIIWFGDLNYRISLP 180
Query: 456 DGEIRKLVDLKKWNELMKFDQL 477
+ R L+ K W++L++ DQ+
Sbjct: 181 YDKTRDLISKKHWSKLVERDQV 246
>TC79955 similar to PIR|E84430|E84430 probable inositol
polyphosphate-5-phosphatase [imported] - Arabidopsis
thaliana, partial (29%)
Length = 733
Score = 75.1 bits (183), Expect = 7e-14
Identities = 34/95 (35%), Positives = 56/95 (58%)
Frame = +3
Query: 84 SETLRAQYINTKEVRVTIGTWNVAGKHPCNDLEIEGWLCTEEEPSDIYIIGFQEVVPLNA 143
SET+ + + ++ + TWN+ G P DL ++ L + DIY++GFQE+VPL A
Sbjct: 282 SETVPNHPKDKHKYKIFVSTWNIGGIAPDEDLSVDDLLEAKNNSCDIYVLGFQEIVPLKA 461
Query: 144 GNVFGAEDSKPIPKWDALIRRTLNKSSEPGTKKKS 178
NV G+E+++ KW+++IR+ LNK +S
Sbjct: 462 SNVLGSENNEISNKWNSIIRKALNKDHRDNVHSES 566
Score = 68.2 bits (165), Expect = 9e-12
Identities = 37/97 (38%), Positives = 58/97 (59%), Gaps = 1/97 (1%)
Frame = +3
Query: 288 QEIVPQVFDSHL-DVSDMLSDEDNDTFSELANNEDANGIISVKSHPKYVRIVSKQMVGIY 346
QEIVP + L ++ +S++ N + N + + + S +H + I+SKQMVGI+
Sbjct: 438 QEIVPLKASNVLGSENNEISNKWNSIIRKALNKDHRDNVHSESTHDDFQCIISKQMVGIF 617
Query: 347 VSVWVQRKLRRHVHHLKVSQVGVGLMGYMGNKGSVSV 383
+SVW +R +R + H VS +G G+M + NKGSVSV
Sbjct: 618 ISVWTRRDIRPFIQHPSVSCIGCGIMRCLRNKGSVSV 728
>AW127178 similar to GP|10444263|gb| inositol polyphosphate 5-phosphatase II
{Arabidopsis thaliana}, partial (5%)
Length = 473
Score = 63.5 bits (153), Expect = 2e-10
Identities = 31/45 (68%), Positives = 37/45 (81%), Gaps = 1/45 (2%)
Frame = -1
Query: 548 QLSDHRPVSSIFLVDVEVIDHRKLERAIYFA-SAVVHPDVFLKED 591
+LSDHRPVSSIFLV+VEV DHRKL RA+ F +A VHP++F ED
Sbjct: 470 KLSDHRPVSSIFLVEVEVFDHRKLRRALNFTNTAAVHPEIFPDED 336
>TC82249 similar to PIR|H84727|H84727 probable inositol polyphosphate
5'-phosphatase [imported] - Arabidopsis thaliana,
partial (7%)
Length = 682
Score = 58.9 bits (141), Expect(2) = 4e-10
Identities = 34/96 (35%), Positives = 53/96 (54%), Gaps = 4/96 (4%)
Frame = +3
Query: 61 TTKPTQLNTSCKDYEMKRHRRRKSETLRA--QYINTKEVRVTIGTWNVAGKHPCNDL--E 116
TTK + S YE++ +S+ I+T ++R+ +GTWNVAG+ P L +
Sbjct: 171 TTKQKENKPSLSLYEIEDIVEDESDEYGELDSCISTNKLRIFVGTWNVAGRSPVGSLAVD 350
Query: 117 IEGWLCTEEEPSDIYIIGFQEVVPLNAGNVFGAEDS 152
++ WL + +DIY++GFQE+VPL V G S
Sbjct: 351 LDEWL-NLKNSADIYVLGFQEIVPLKTSTVIGGRRS 455
Score = 23.5 bits (49), Expect(2) = 4e-10
Identities = 10/20 (50%), Positives = 12/20 (60%)
Frame = +1
Query: 148 GAEDSKPIPKWDALIRRTLN 167
GAED W+ LI +TLN
Sbjct: 442 GAEDPSVATNWNNLIGKTLN 501
>AW686583 similar to PIR|E84430|E844 probable inositol
polyphosphate-5-phosphatase [imported] - Arabidopsis
thaliana, partial (18%)
Length = 605
Score = 58.9 bits (141), Expect = 5e-09
Identities = 46/150 (30%), Positives = 59/150 (38%)
Frame = +2
Query: 1 MKGKRSEVFWPSTVMKKWLNVKQKVYDFSEDEANTETDESEDDVKLIEIETNHTSKPQAN 60
M K +EV WP+ V K LN + +F D + + P +
Sbjct: 161 MTRKLAEVMWPALVANKILNRRLGSRNFIADYPSY-------------------TDPLLS 283
Query: 61 TTKPTQLNTSCKDYEMKRHRRRKSETLRAQYINTKEVRVTIGTWNVAGKHPCNDLEIEGW 120
T QL+ S K IN TWN+ G P L IE
Sbjct: 284 TNNDDQLSLSTKS------------------INDHSDTQKYNTWNIGGIAPDEGLNIEDL 409
Query: 121 LCTEEEPSDIYIIGFQEVVPLNAGNVFGAE 150
L T + DIY+ GFQE+VPLNA NV G+E
Sbjct: 410 LETCSKSFDIYVFGFQEIVPLNASNVLGSE 499
>AL376063 weakly similar to PIR|H85044|H85 hypothetical protein AT4g03540
[imported] - Arabidopsis thaliana, partial (73%)
Length = 504
Score = 32.7 bits (73), Expect = 0.41
Identities = 18/53 (33%), Positives = 30/53 (55%), Gaps = 6/53 (11%)
Frame = -2
Query: 48 EIETNHTSKPQANTTKPTQLNTSCKDYEMKRHRRRKSETL------RAQYINT 94
++E HTS+ Q N T+PT++++ +D +RR S T+ R +YI T
Sbjct: 452 DLEEEHTSRFQTNITRPTKVSSKVQDAAQVFLKRRGSRTV*MECNRRIKYIMT 294
>AJ502664
Length = 535
Score = 31.6 bits (70), Expect = 0.91
Identities = 13/20 (65%), Positives = 16/20 (80%)
Frame = -1
Query: 48 EIETNHTSKPQANTTKPTQL 67
+IE +HT K ANTTKPTQ+
Sbjct: 208 DIENDHTLKSPANTTKPTQI 149
>TC77998 similar to PIR|T09254|T09254 isoflavone-7-O-methyltransferase (EC
2.1.1.-) 9 - alfalfa, partial (36%)
Length = 758
Score = 31.2 bits (69), Expect = 1.2
Identities = 11/20 (55%), Positives = 17/20 (85%)
Frame = -3
Query: 48 EIETNHTSKPQANTTKPTQL 67
+++ HTS+ QANTT+PTQ+
Sbjct: 540 DLQKEHTSRSQANTTRPTQM 481
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.132 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,604,359
Number of Sequences: 36976
Number of extensions: 246493
Number of successful extensions: 1527
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 1483
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1504
length of query: 599
length of database: 9,014,727
effective HSP length: 102
effective length of query: 497
effective length of database: 5,243,175
effective search space: 2605857975
effective search space used: 2605857975
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147404.2


