

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147363.1 - phase: 0 /pseudo
(427 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
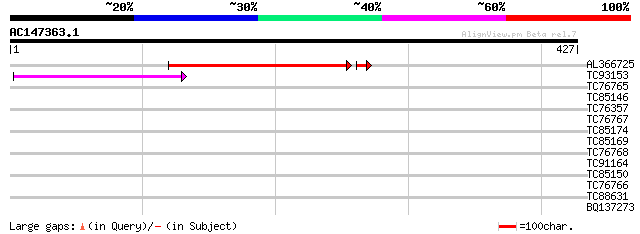
Score E
Sequences producing significant alignments: (bits) Value
AL366725 135 2e-33
TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Ci... 83 2e-16
TC76765 homologue to GP|12054977|emb|CAC20725. putative chalcone... 30 1.4
TC85146 homologue to SP|P30074|CHS2_MEDSA Chalcone synthase 2 (E... 30 1.4
TC76357 similar to SP|O04865|PLD_VIGUN Phospholipase D precursor... 29 3.0
TC76767 homologue to SP|P30074|CHS2_MEDSA Chalcone synthase 2 (E... 29 4.0
TC85174 homologue to SP|P30075|CHS4_MEDSA Chalcone synthase 4 (E... 29 4.0
TC85169 SP|P30077|CHS9_MEDSA Chalcone synthase 9 (EC 2.3.1.74) (... 28 5.2
TC76768 SP|P30075|CHS4_MEDSA Chalcone synthase 4 (EC 2.3.1.74) (... 28 6.8
TC91164 similar to GP|21554376|gb|AAM63483.1 unknown {Arabidopsi... 28 6.8
TC85150 homologue to SP|P30077|CHS9_MEDSA Chalcone synthase 9 (E... 28 6.8
TC76766 SP|P30075|CHS4_MEDSA Chalcone synthase 4 (EC 2.3.1.74) (... 28 6.8
TC88631 weakly similar to PIR|T00579|T00579 probable laccase [im... 28 8.8
BQ137273 28 8.8
>AL366725
Length = 485
Score = 135 bits (340), Expect(2) = 2e-33
Identities = 88/138 (63%), Positives = 98/138 (70%)
Frame = +3
Query: 120 IRITLQRLLSFRSVSSSRMD*GLRSRGQLDTNRSEFSLSW*TVAGYMRRIPRLITRL*MC 179
IR L RLLS R+ SSSRM *G SRGQ DTN SEFSL * A RRI RL+TRL*M
Sbjct: 9 IRTMLLRLLSSRNASSSRMV*GPTSRGQSDTNSSEFSLI**IPAESTRRILRLMTRL*MS 188
Query: 180 GRLRANRVALSPIMPLLIRANRGWSRIDGRGRRMLMLRLSVLIVVRKAIRAMLVLER*KD 239
GR RANRVALS I+PLLIRANR WS I RRML+ RLSV + RKA R M VL+R ++
Sbjct: 189 GRPRANRVALSLIVPLLIRANREWSMIGVLRRRMLLRRLSVSTMARKATRVMFVLKRSRN 368
Query: 240 VSDVVRRVMH*PIATVMI 257
VS V RRV+ * IA+ MI
Sbjct: 369 VSGVTRRVIL*LIASEMI 422
Score = 25.4 bits (54), Expect(2) = 2e-33
Identities = 9/11 (81%), Positives = 11/11 (99%)
Frame = +2
Query: 262 YIGSQCKQPKK 272
+IGSQCKQPK+
Sbjct: 452 HIGSQCKQPKR 484
>TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Cicer
arietinum}, partial (8%)
Length = 516
Score = 83.2 bits (204), Expect = 2e-16
Identities = 59/130 (45%), Positives = 75/130 (57%)
Frame = +3
Query: 4 GIILLLSREDMTPMEPRRGLRRSREFFVLCNALKFRRCDLVHIS*PRKLMIGGLVLYLP* 63
GII SR M M R G RR +E+ CN L+ RRC L * ++ MIGGLV L
Sbjct: 123 GIIHQHSRVGMLLMVLRSG*RRLKEYSESCNVLRHRRCSLGRTC*LKRPMIGGLVCCLFW 302
Query: 64 SKMVLL*LGLCLGGSS*ADTFRKMFAGRKRLSSLS*SKETCQ*LSMLPSLWSWQNFIRIT 123
S+M+L * G C G S A F++M RKRL+ S*++ TC SML SLW+ + IT
Sbjct: 303 SRMMLW*PGPCSGRSFWAGIFQRMSGVRKRLNFWS*NRVTCLSQSMLQSLWNLPHSTLIT 482
Query: 124 LQRLLSFRSV 133
++R LS SV
Sbjct: 483 VRRQLSSPSV 512
>TC76765 homologue to GP|12054977|emb|CAC20725. putative chalcone synthase
{Medicago truncatula}, complete
Length = 1418
Score = 30.4 bits (67), Expect = 1.4
Identities = 31/109 (28%), Positives = 47/109 (42%), Gaps = 7/109 (6%)
Frame = -3
Query: 302 NTPLVA------IIDIGATHCFIAFDCASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLK 355
+TPLV I D G H FIAF AS P+ G+ TT++ CL
Sbjct: 531 STPLVVQTIK*VIFDFG*PHSFIAFTAAS----------------FPSLGTSTTTMSCLA 400
Query: 356 SSL-SMFGRDFEMDLVCLPLSGMYVILGMNWFE*NHVHINYFSKSVYFS 403
S+ +M+ + + MY++L M+ +H+ N+ SV S
Sbjct: 399 SNEGAMYSQTLGFSFKISSVRYMYLLLIMDL---SHMR*NFSLSSVLCS 262
>TC85146 homologue to SP|P30074|CHS2_MEDSA Chalcone synthase 2 (EC 2.3.1.74)
(Naringenin-chalcone synthase 2). [Alfalfa] {Medicago
sativa}, complete
Length = 1406
Score = 30.4 bits (67), Expect = 1.4
Identities = 31/109 (28%), Positives = 47/109 (42%), Gaps = 7/109 (6%)
Frame = -2
Query: 302 NTPLVA------IIDIGATHCFIAFDCASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLK 355
+TPLV I D G H FIAF AS P+ G+ TT++ CL
Sbjct: 520 STPLVVQTIK*VIFDFG*PHSFIAFTAAS----------------FPSLGTSTTTMSCLA 389
Query: 356 SSL-SMFGRDFEMDLVCLPLSGMYVILGMNWFE*NHVHINYFSKSVYFS 403
S+ +M+ + + MY++L M+ +H+ N+ SV S
Sbjct: 388 SNEGAMYSQTLGFSFKISSVRYMYLLLIMDL---SHMR*NFSLSSVLCS 251
>TC76357 similar to SP|O04865|PLD_VIGUN Phospholipase D precursor (EC
3.1.4.4) (PLD) (Choline phosphatase), complete
Length = 2773
Score = 29.3 bits (64), Expect = 3.0
Identities = 11/26 (42%), Positives = 17/26 (65%)
Frame = -2
Query: 381 LGMNWFE*NHVHINYFSKSVYFSSVE 406
+ MNWF +HI YF+ S+YF + +
Sbjct: 486 ISMNWF---FIHIQYFNPSIYFLTTQ 418
>TC76767 homologue to SP|P30074|CHS2_MEDSA Chalcone synthase 2 (EC 2.3.1.74)
(Naringenin-chalcone synthase 2). [Alfalfa] {Medicago
sativa}, complete
Length = 1436
Score = 28.9 bits (63), Expect = 4.0
Identities = 30/109 (27%), Positives = 47/109 (42%), Gaps = 7/109 (6%)
Frame = -3
Query: 302 NTPLVA------IIDIGATHCFIAFDCASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLK 355
+TPLV I D G H FIAF AS P+ G+ TT++ CL
Sbjct: 534 STPLVVQTIK*VIFDFG*PHSFIAFTAAS----------------FPSLGTSTTTMSCLA 403
Query: 356 SSL-SMFGRDFEMDLVCLPLSGMYVILGMNWFE*NHVHINYFSKSVYFS 403
S+ +M+ + + MY++L ++ +H+ N+ SV S
Sbjct: 402 SNEGAMYSQTLGFSFKISSVRYMYLLLIIDL---SHMR*NFSLSSVLCS 265
>TC85174 homologue to SP|P30075|CHS4_MEDSA Chalcone synthase 4 (EC 2.3.1.74)
(Naringenin-chalcone synthase 4) (CHS12-1). [Alfalfa],
partial (82%)
Length = 1072
Score = 28.9 bits (63), Expect = 4.0
Identities = 30/109 (27%), Positives = 47/109 (42%), Gaps = 7/109 (6%)
Frame = -3
Query: 302 NTPLVA------IIDIGATHCFIAFDCASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLK 355
+TPLV I D G H FIAF AS P+ G+ TT++ CL
Sbjct: 509 STPLVVQTIK*VIFDFG*PHSFIAFTAAS----------------FPSLGTSTTTMSCLA 378
Query: 356 SSL-SMFGRDFEMDLVCLPLSGMYVILGMNWFE*NHVHINYFSKSVYFS 403
S+ +M+ + + MY++L ++ +H+ N+ SV S
Sbjct: 377 SNEGAMYSQTLGFSFKISSVRYMYLLLIIDL---SHMR*NFSLSSVLCS 240
>TC85169 SP|P30077|CHS9_MEDSA Chalcone synthase 9 (EC 2.3.1.74)
(Naringenin-chalcone synthase 9). [Alfalfa] {Medicago
sativa}, complete
Length = 1612
Score = 28.5 bits (62), Expect = 5.2
Identities = 26/90 (28%), Positives = 38/90 (41%), Gaps = 7/90 (7%)
Frame = -2
Query: 302 NTPLVA------IIDIGATHCFIAFDCASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLK 355
+TPLV I D G H FIAF AS P+ G+ TT++ CL
Sbjct: 717 STPLVVQKIKCVIFDFGWPHSFIAFTAAS----------------FPSLGTSTTTMSCLA 586
Query: 356 SS-LSMFGRDFEMDLVCLPLSGMYVILGMN 384
S+ +M+ + MY++L M+
Sbjct: 585 SNDGAMYSHTLGFSFKISSVRYMYLLLIMD 496
>TC76768 SP|P30075|CHS4_MEDSA Chalcone synthase 4 (EC 2.3.1.74)
(Naringenin-chalcone synthase 4) (CHS12-1). [Alfalfa],
partial (52%)
Length = 717
Score = 28.1 bits (61), Expect = 6.8
Identities = 25/87 (28%), Positives = 37/87 (41%), Gaps = 7/87 (8%)
Frame = -3
Query: 302 NTPLVA------IIDIGATHCFIAFDCASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLK 355
+TPLV I D G H FIAF AS P+ G+ TT++ CL
Sbjct: 514 STPLVVQTIK*VIFDFG*PHSFIAFTAAS----------------FPSLGTSTTTMSCLA 383
Query: 356 SSL-SMFGRDFEMDLVCLPLSGMYVIL 381
S+ +M+ + + MY++L
Sbjct: 382 SNEGAMYSQTLGFSFKISSVRYMYLLL 302
>TC91164 similar to GP|21554376|gb|AAM63483.1 unknown {Arabidopsis
thaliana}, partial (96%)
Length = 739
Score = 28.1 bits (61), Expect = 6.8
Identities = 22/61 (36%), Positives = 30/61 (49%), Gaps = 6/61 (9%)
Frame = +2
Query: 351 LVCLKSSLSMFGRDFEMDLVCLPLSGMYV------ILGMNWFE*NHVHINYFSKSVYFSS 404
L+ L +S +F+M L L L+G+Y+ ILG NW + NYF S F S
Sbjct: 236 LLLLAIIISRKNTNFQMCLFLLALAGVYLAERLNSILGENWK--SFSSQNYFDPSGVFMS 409
Query: 405 V 405
V
Sbjct: 410 V 412
>TC85150 homologue to SP|P30077|CHS9_MEDSA Chalcone synthase 9 (EC 2.3.1.74)
(Naringenin-chalcone synthase 9). [Alfalfa] {Medicago
sativa}, complete
Length = 1389
Score = 28.1 bits (61), Expect = 6.8
Identities = 25/87 (28%), Positives = 36/87 (40%), Gaps = 7/87 (8%)
Frame = -3
Query: 302 NTPLVA------IIDIGATHCFIAFDCASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLK 355
+TPLV I D G H FIAF AS P+ G+ TT++ CL
Sbjct: 520 STPLVVQNIKCVIFDFGWPHSFIAFTAAS----------------FPSLGTSTTTMSCLA 389
Query: 356 SSL-SMFGRDFEMDLVCLPLSGMYVIL 381
S+ +M+ + MY++L
Sbjct: 388 SNEGAMYSHTLGFSFKISSVRYMYLLL 308
>TC76766 SP|P30075|CHS4_MEDSA Chalcone synthase 4 (EC 2.3.1.74)
(Naringenin-chalcone synthase 4) (CHS12-1). [Alfalfa],
partial (50%)
Length = 731
Score = 28.1 bits (61), Expect = 6.8
Identities = 30/109 (27%), Positives = 47/109 (42%), Gaps = 7/109 (6%)
Frame = -1
Query: 302 NTPLVA------IIDIGATHCFIAFDCASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLK 355
+TPLV I D G H FIAF AS P+ G+ TT++ CL
Sbjct: 524 STPLVVQTIK*VIFDFGWPHSFIAFTAAS----------------FPSLGTSTTTMSCLA 393
Query: 356 SSL-SMFGRDFEMDLVCLPLSGMYVILGMNWFE*NHVHINYFSKSVYFS 403
S+ +M+ + + MY++L ++ +H+ N+ SV S
Sbjct: 392 SNEGAMYSQTLGFFFKISSVRYMYLLLIIDL---SHMR*NFSLSSVLCS 255
>TC88631 weakly similar to PIR|T00579|T00579 probable laccase [imported] -
Arabidopsis thaliana, partial (38%)
Length = 779
Score = 27.7 bits (60), Expect = 8.8
Identities = 15/44 (34%), Positives = 21/44 (47%), Gaps = 1/44 (2%)
Frame = +2
Query: 298 CYINNTPLVAIIDIGATHCFIAFDCASTL-GLDMSNMNGEMVVE 340
C+ N L+ + HCFI F C +L + SN N E V+
Sbjct: 17 CHNNEDYLMLLTWFTYYHCFICFSCRESLPSVCYSNCNSEEAVQ 148
>BQ137273
Length = 699
Score = 27.7 bits (60), Expect = 8.8
Identities = 14/42 (33%), Positives = 24/42 (56%), Gaps = 1/42 (2%)
Frame = +1
Query: 315 HCFIAFDCASTLGLDMS-NMNGEMVVETPAKGSVTTSLVCLK 355
HC + C+ LG + M +++ ETPA+ VT ++CL+
Sbjct: 388 HCVPSSMCSFMLGYACNCTMRIKLLQETPAQNCVTYGMLCLR 513
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.343 0.151 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,762,960
Number of Sequences: 36976
Number of extensions: 175862
Number of successful extensions: 1390
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 1039
Number of HSP's successfully gapped in prelim test: 35
Number of HSP's that attempted gapping in prelim test: 343
Number of HSP's gapped (non-prelim): 1089
length of query: 427
length of database: 9,014,727
effective HSP length: 99
effective length of query: 328
effective length of database: 5,354,103
effective search space: 1756145784
effective search space used: 1756145784
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 60 (27.7 bits)
Medicago: description of AC147363.1


