

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147202.5 - phase: 0
(389 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
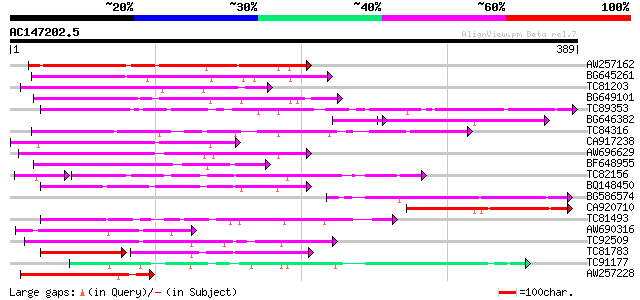
Score E
Sequences producing significant alignments: (bits) Value
AW257162 169 2e-42
BG645261 homologue to PIR|H90254|H902 sulfate ABC transporter p... 139 1e-33
TC81203 weakly similar to GP|10177464|dbj|BAB10855. gb|AAD21700.... 139 2e-33
BG649101 homologue to GP|6648199|gb|A unknown protein {Arabidops... 131 4e-31
TC89353 weakly similar to GP|9279704|dbj|BAB01261.1 gb|AAF25964.... 127 1e-29
BG646382 114 1e-29
TC84316 weakly similar to GP|11994612|dbj|BAB02749. gb|AAC24186.... 126 1e-29
CA917238 weakly similar to PIR|A84538|A845 hypothetical protein ... 125 3e-29
AW696629 105 2e-23
BF648955 similar to GP|18043667|gb| Unknown (protein for IMAGE:3... 101 4e-22
TC82156 similar to GP|20197985|gb|AAM15340.1 F-box protein famil... 91 1e-21
BQ148450 similar to GP|10177464|dbj gb|AAD21700.1~gene_id:MQB2.1... 95 4e-20
BG586574 94 9e-20
CA920710 similar to PIR|S64314|S643 probable membrane protein YG... 93 2e-19
TC81493 weakly similar to PIR|F86249|F86249 protein F25C20.23 [i... 86 2e-17
AW690316 86 3e-17
TC92509 weakly similar to PIR|T49129|T49129 hypothetical protein... 83 2e-16
TC81783 similar to GP|9294656|dbj|BAB03005.1 gb|AAF30317.1~gene_... 56 7e-16
TC91177 similar to GP|9294656|dbj|BAB03005.1 gb|AAF30317.1~gene_... 80 1e-15
AW257228 GP|17342711|gb 1-aminocyclopropanecarboxylic acid oxida... 79 4e-15
>AW257162
Length = 728
Score = 169 bits (427), Expect = 2e-42
Identities = 98/203 (48%), Positives = 128/203 (62%), Gaps = 9/203 (4%)
Frame = +1
Query: 14 NVASSPIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLA 73
+++SSP+ P D+I E+LS+L VK+L++ +CV SW LISDS FVK+HL RS ++ +
Sbjct: 106 SLSSSPVF-PDDIIAEILSWLTVKTLMKMKCVSKSWNTLISDSNFVKMHLNRSARHSQSY 282
Query: 74 VTQESRECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLCLLG 133
+ E R D P S+ L+ S TL DPYY L +KDC V GSCNGL+CL G
Sbjct: 283 LVSEHRG---DYNFVPFSVRGLMNGRS--ITLPKDPYYQLIEKDCPGVVGSCNGLVCLSG 447
Query: 134 ----LSEDREMWLRFWNPATRAISDKLGHYPADFTGGFEVAFGYDNSTDTYKVV--YLQK 187
+ E EMWLR WNPATR ISDKL ++ A+ +E FGYDN+T TYKVV Y
Sbjct: 448 CVADVEEFEEMWLRIWNPATRTISDKL-YFSANRLQCWEFMFGYDNTTQTYKVVALYPDS 624
Query: 188 GMT---GVFSLGDNVWRNIESFP 207
MT G+ +N+WRNI+SFP
Sbjct: 625 EMTTKVGIICFRNNIWRNIQSFP 693
>BG645261 homologue to PIR|H90254|H902 sulfate ABC transporter permease
protein SSO1032 [imported] - Sulfolobus solfataricus,
partial (4%)
Length = 771
Score = 139 bits (351), Expect = 1e-33
Identities = 94/229 (41%), Positives = 128/229 (55%), Gaps = 23/229 (10%)
Frame = +1
Query: 16 ASSPIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAVT 75
+++ ++ L LI L F + + R + + WK +I FVKLHL RS Q P L
Sbjct: 31 SATHLLFSLILILFLRLFKKSRQESKNRFINILWKSIIDHPTFVKLHLNRSSQKPDLTFV 210
Query: 76 QESRECSMDVIVSPTSLS----HLLENPSKPTTLTND-PYYSLNDKDCRSVAGSCNGLLC 130
+R ++ + S ++S LLENP L++D PYY L DKDC + GSCNGLLC
Sbjct: 211 SSAR-ITLWTLESTCTVSFTVFRLLENPPVIINLSDDDPYYQLKDKDCVHIIGSCNGLLC 387
Query: 131 LLGLSED-----REMWLRFWNPATRAISDKLGHY-------PADFTGG---FEVAFGYDN 175
L GL + ++ +RFWNPA+R IS K+G+ P+ F G + FGY N
Sbjct: 388 LFGLKINDSCGHMDISIRFWNPASRKISKKVGYCGDGVNNDPSCFPGKGDLLKFVFGYVN 567
Query: 176 STDTYKVVYLQKGMTG--VFSLGDNVWRNIESFPLG-YYLDNRVHLRDS 221
S+DTYKVVY G T VFSLGDNVWR IE+ P+ +++ VHL S
Sbjct: 568 SSDTYKVVYFIVGTTSARVFSLGDNVWRKIENSPVVIHHVSGFVHLSGS 714
>TC81203 weakly similar to GP|10177464|dbj|BAB10855.
gb|AAD21700.1~gene_id:MQB2.18~similar to unknown protein
{Arabidopsis thaliana}, partial (9%)
Length = 715
Score = 139 bits (349), Expect = 2e-33
Identities = 84/191 (43%), Positives = 107/191 (55%), Gaps = 18/191 (9%)
Frame = +1
Query: 8 SQQPPSNVASSPIILPLD-LITELLSFLPVKSLLQFRCVCMSWKILIS-DSFFVKLHLQR 65
+ PP + +P L LD LI ++LS LPVK+L+QF+CVC SWK LIS D F KLHLQR
Sbjct: 154 NSDPPQSNGGAPTSLLLDELIVDILSRLPVKTLMQFKCVCKSWKTLISHDPSFAKLHLQR 333
Query: 66 SIQNPRLA-VTQESRECSMDVIVSPTSLSHLLENPSKPT-----------TLTNDPYYSL 113
S +N L V+ S + + V P +SHL+E P T + +DPYY L
Sbjct: 334 SPRNTHLTLVSDRSSDDESNFSVVPFPVSHLIEAPLTITPFEPYHLLRNVAIPDDPYYVL 513
Query: 114 NDKDCRSVAGSCNGLLCL----LGLSEDREMWLRFWNPATRAISDKLGHYPADFTGGFEV 169
+ DC + GSCNGL CL L E+ W RFWNPAT +S++LG F +
Sbjct: 514 GNMDCCIIIGSCNGLYCLRCYSLIYEEEEHDWFRFWNPATNTLSEELG----CLNEFFRL 681
Query: 170 AFGYDNSTDTY 180
FGYD S DTY
Sbjct: 682 TFGYDISNDTY 714
>BG649101 homologue to GP|6648199|gb|A unknown protein {Arabidopsis
thaliana}, partial (3%)
Length = 799
Score = 131 bits (330), Expect = 4e-31
Identities = 90/231 (38%), Positives = 123/231 (52%), Gaps = 19/231 (8%)
Frame = +1
Query: 17 SSPIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAVTQ 76
S P+++P DLI +L+FLPVK++ Q + V SW LI+ F+K+HL +S QNP +T
Sbjct: 70 SVPVVIPSDLIAIILTFLPVKTITQLKLVSKSWNTLITSPSFIKIHLNQSSQNPNFILTP 249
Query: 77 ESRECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYS-LNDKDCRSVAGSCNGLLCLLGLS 135
++ S++ ++S + LL T++ D Y++ LN+ V GSCNGLLCLL S
Sbjct: 250 SRKQYSINNVLS-VPIPRLLTG----NTVSGDTYHNILNNDHHFRVVGSCNGLLCLLFKS 414
Query: 136 E---DREMWLRFWNPATRAISDKLG---HYPADFTGGFEVAFGYDNSTDTYKVVYLQKGM 189
E + R WNPATR IS++LG Y F G FG D TYK+V L
Sbjct: 415 EFITHLKFRFRIWNPATRTISEELGFFRKYKPLFGGVSRFTFGCDYLRGTYKLVALHTVE 594
Query: 190 TG--------VFSLG----DNVWRNIESFPLGYYLDNRVHLRDSLNWLGLR 228
G VF+LG D WRNI P + + VHL + NWL LR
Sbjct: 595 DGDVMRSNVRVFNLGNDDSDKCWRNI---PNPFVCADGVHLSGTGNWLSLR 738
>TC89353 weakly similar to GP|9279704|dbj|BAB01261.1
gb|AAF25964.1~gene_id:MYA6.2~similar to unknown protein
{Arabidopsis thaliana}, partial (10%)
Length = 1482
Score = 127 bits (318), Expect = 1e-29
Identities = 120/388 (30%), Positives = 179/388 (45%), Gaps = 20/388 (5%)
Frame = +3
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRL-AVTQESRE 80
+P ++I E+L LPV+SLLQFRCVC WK LISD F K H+ S P+L +V +
Sbjct: 105 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAK 284
Query: 81 CSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGLSEDREM 140
C++ + P L LL+NPS D + + ++ GSCNGLLCL +
Sbjct: 285 CNL--VSYP--LKPLLDNPSAHRVEPAD--FEMIHTTSMTIIGSCNGLLCLSDFYQ---- 434
Query: 141 WLRFWNPATRAISDKLGHYPADFTGGFEV---------AFGYDNSTDTYKVV-------Y 184
WNP S KL P+ F+ FGYD D YKV+
Sbjct: 435 -FTLWNP-----SIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYN 596
Query: 185 LQKGMTGVFSLGDNVWRNIESFPLGYYLDNRVHLRDSLNWLGLRSYVDDCDDYDCEYITS 244
L + T +++ G W I+ FP R L LG+ +V + +I S
Sbjct: 597 LDETKTLIYTFGGKDWTTIQKFPCD-------PSRCDLGRLGVGKFVSG----NLNWIVS 743
Query: 245 IEQFMIVALDLRTETCKELLLPRGFDE--VPCYEPSLCVLMDCICFSHLVKKTHLVIWKM 302
+ +IV D+ ET E+ LP+ + + Y S + + F H KTH V+W M
Sbjct: 744 --KKVIVFFDIEKETYGEMSLPQDYGDKNTVLYVSSNRIY---VSFDH-SNKTHWVVWMM 905
Query: 303 MDYGDDDSWTQLLEINLQILKKIDEWSAWVPLHLSKNYDTLILGSTLEHEFAVYNL-RDG 361
+YG +SWT+L+ I L + L +S++ ++L + +VYNL DG
Sbjct: 906 KEYGVVESWTKLMIIPQDKLTSPGPYCLSDALFISEH--GVLLMRPQHSKLSVYNLNNDG 1079
Query: 362 SIEKTRITNGESSWIYINDYVESLVFCH 389
++ +G+ + Y++ Y ESLV H
Sbjct: 1080GLDYCTTISGQFA-RYLHIYNESLVSPH 1160
>BG646382
Length = 603
Score = 114 bits (286), Expect(2) = 1e-29
Identities = 61/132 (46%), Positives = 79/132 (59%), Gaps = 14/132 (10%)
Frame = +3
Query: 253 LDLRTETCKELLLPRGFDEVPCYEPSLCVLMDCICFSHLVKKTHLVIWKMMDYGDDDSWT 312
L R ++ LLP GF EVPC EPSL VLMDC+CFSH K+T VIW+M ++G +SWT
Sbjct: 159 LIFRRRHTRQFLLPAGFKEVPCVEPSLRVLMDCLCFSHDFKRTEFVIWQMKEFGSQESWT 338
Query: 313 QLLEI--------------NLQILKKIDEWSAWVPLHLSKNYDTLILGSTLEHEFAVYNL 358
QL I NL +L ++ +PL+LS+N DTLIL + E +YN
Sbjct: 339 QLFRIKYINLQIHNLPINDNLDLLGYMECNIPLLPLYLSENGDTLILSNDEEEGVIIYNQ 518
Query: 359 RDGSIEKTRITN 370
RD +EKTRI+N
Sbjct: 519 RDKRVEKTRISN 554
Score = 33.1 bits (74), Expect(2) = 1e-29
Identities = 19/40 (47%), Positives = 24/40 (59%), Gaps = 2/40 (5%)
Frame = +2
Query: 222 LNWLGLRS-YVD-DCDDYDCEYITSIEQFMIVALDLRTET 259
L WL L +V DC + YI +EQ +IV+LDL TET
Sbjct: 59 LTWLALSGDFVSIDCGETSKAYIPLVEQLVIVSLDLSTET 178
>TC84316 weakly similar to GP|11994612|dbj|BAB02749.
gb|AAC24186.1~gene_id:MGL6.4~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (10%)
Length = 892
Score = 126 bits (317), Expect = 1e-29
Identities = 102/320 (31%), Positives = 149/320 (45%), Gaps = 18/320 (5%)
Frame = +3
Query: 16 ASSPI-ILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAV 74
A+ P+ LP +LI +L LPV+SLL+F+CVC SWK L SD+ F H S P+L V
Sbjct: 21 ANKPLPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQL-V 197
Query: 75 TQESRECSMDVIVSPTSLSHLLENPSK---PTTLTNDPYYSLNDKDCRSVAGSCNGLLCL 131
ES + + LLEN S P + T Y ++ GSCNG LCL
Sbjct: 198 ACESVSAYRTWEIKTYPIESLLENSSTTVIPVSNTGHQRY--------TILGSCNGFLCL 353
Query: 132 LGLSEDREMWLRFWNPATRAIS------DKLGHYPADFTGGFEVAFGYDNSTDTYKVV-- 183
++ + +R WNP+ S D+ +Y FGYD YK++
Sbjct: 354 Y---DNYQRCVRLWNPSINLKSKSSPTIDRFIYY----------GFGYDQVNHKYKLLAV 494
Query: 184 --YLQKGMTGVFSLGDNVWRNIESFPLGYYLDNRVHL----RDSLNWLGLRSYVDDCDDY 237
+ + T +++ G+N +N+E Y NR HL +LNW+ VD+ D
Sbjct: 495 KAFSRITETMIYTFGENSCKNVEVKDFPRYPPNRKHLGKFVSGTLNWI-----VDERDG- 656
Query: 238 DCEYITSIEQFMIVALDLRTETCKELLLPRGFDEVPCYEPSLCVLMDCICFSHLVKKTHL 297
+ I++ D+ ET +++LLP+ V Y P L VL +CIC T
Sbjct: 657 ---------RATILSFDIEKETYRQVLLPQHGYAV--YSPGLYVLSNCICVCTSFLDTRW 803
Query: 298 VIWKMMDYGDDDSWTQLLEI 317
+W M YG +SWT+L+ I
Sbjct: 804 QLWMMKKYGVAESWTKLMSI 863
>CA917238 weakly similar to PIR|A84538|A845 hypothetical protein At2g16220
[imported] - Arabidopsis thaliana, partial (6%)
Length = 603
Score = 125 bits (314), Expect = 3e-29
Identities = 74/165 (44%), Positives = 97/165 (57%), Gaps = 7/165 (4%)
Frame = +2
Query: 1 MNLSPPKSQQPPSNVASS----PIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDS 56
M+ P KS+ P A + P +LP +LITE+LS VKSL++ RCV W +ISD
Sbjct: 71 MSDPPAKSRVQPEAEAQAEAEPPSVLPDELITEVLSRGDVKSLMRMRCVSEYWNSMISDP 250
Query: 57 FFVKLHLQRSIQNPRLAVTQESRECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDK 116
FVKLH++RS +N L ++ D V P + L+EN TL +DPYY L DK
Sbjct: 251 RFVKLHMKRSARNAHLTLSLCKSGIDGDNNVVPYPVRGLIENGL--ITLPSDPYYRLKDK 424
Query: 117 DCRSVAGSCNGLLCLLGLSE---DREMWLRFWNPATRAISDKLGH 158
+C+ V GSCNG LCLLG S R +W RFWNPA ++ KLG+
Sbjct: 425 ECQYVIGSCNGWLCLLGFSSIGAYRHIWFRFWNPAMGKMTQKLGY 559
>AW696629
Length = 663
Score = 105 bits (263), Expect = 2e-23
Identities = 76/211 (36%), Positives = 103/211 (48%), Gaps = 10/211 (4%)
Frame = +2
Query: 7 KSQQPPSNVASSPIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRS 66
K ++ SSP ILP DLI ++LS+LPVK L++F V WK LI D F KLHLQ+S
Sbjct: 20 KKRESSEEATSSPPILPSDLIMQILSWLPVKLLIRFTSVSKHWKSLILDPNFAKLHLQKS 199
Query: 67 IQNPRLAVTQESRECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCN 126
+N + +T E V+ S LLE S +D D + GS N
Sbjct: 200 PKNTHMILTALDDEDDTWVVTPYPVRSLLLEQSS-----FSDEECCCFDYHSYFIVGSTN 364
Query: 127 GLLCLL--GLSEDR--EMWLRFWNPATRAISDKLGHYPADFTGGFEVAFGYDNSTDTYKV 182
GL+CL E+R E++++FWNP+ R S K G + FGYD+ DTYK
Sbjct: 365 GLVCLAVEKSLENRKYELFIKFWNPSLRLRSKKAPSLNIGLYGTARLGFGYDDLNDTYKA 544
Query: 183 V-----YLQKGMTG-VFSLGDNVWRNIESFP 207
V + M G V +GD+ W+ P
Sbjct: 545 VAVFWDHTTHKMEGRVHCMGDSCWKKDYRLP 637
>BF648955 similar to GP|18043667|gb| Unknown (protein for IMAGE:3661715) {Mus
musculus}, partial (3%)
Length = 514
Score = 101 bits (252), Expect = 4e-22
Identities = 62/166 (37%), Positives = 89/166 (53%), Gaps = 3/166 (1%)
Frame = +3
Query: 17 SSPIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAVTQ 76
+S LP +LI E++S+LPVK L+QFRCV +K LISD +FV++HL++S +NP LA+
Sbjct: 42 ASAAFLPSELIVEIISWLPVKYLMQFRCVSKFYKTLISDPYFVQMHLEKSARNPHLALMW 221
Query: 77 ESRECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGL-- 134
+ D + S+S LL N + +Y + GSCNGLLCL+
Sbjct: 222 QDDLLREDGSIIFLSVSRLLGNKYTTPPFQSGTFYH------SCIIGSCNGLLCLVDFHC 383
Query: 135 -SEDREMWLRFWNPATRAISDKLGHYPADFTGGFEVAFGYDNSTDT 179
+L FWNPATR K + + F+ +FGYD + T
Sbjct: 384 PDYYYYYYLYFWNPATRT---KFXNILITLSXDFKFSFGYDTLSKT 512
>TC82156 similar to GP|20197985|gb|AAM15340.1 F-box protein family AtFBX9
{Arabidopsis thaliana}, partial (4%)
Length = 1136
Score = 91.3 bits (225), Expect(2) = 1e-21
Identities = 76/251 (30%), Positives = 114/251 (45%), Gaps = 7/251 (2%)
Frame = +3
Query: 43 RCVCMSWKILISDSFFVKLHLQRSIQNPRLAVTQESRECSMDVIVSPTSLSHLLENPSKP 102
+ V SWK LISDS F K +L+ S + RL + ++ I + +LS L+
Sbjct: 459 KSVSKSWKSLISDSNFTKKNLRVSTTSHRLLFPKLTKG---QYIFNACTLSSLITTKGTA 629
Query: 103 TTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGLSEDREMWLRFWNPATRAISDKLGHYPAD 162
T + + LN + + GSC+G+LCL E + + WNP + L
Sbjct: 630 TAMQ----HPLNIRKFDKIRGSCHGILCL----ELHQRFAILWNPFINKYAS-LPPLEIP 782
Query: 163 FTGGFEVAFGYDNSTDTYKVVYLQKGM-------TGVFSLGDNVWRNIESFPLGYYLDNR 215
++ FGYD+STD+YKV K M T V ++G WR I+ FP YL +
Sbjct: 783 WSNTIYSCFGYDHSTDSYKVAAFIKWMPNSEIYKTYVHTMGTTSWRMIQDFPCTPYLKSG 962
Query: 216 VHLRDSLNWLGLRSYVDDCDDYDCEYITSIEQFMIVALDLRTETCKELLLPRGFDEVPCY 275
+ + NWL + D Y S+ ++V+L L E+ E+L P + V
Sbjct: 963 KFVSWTFNWLAYK------DKY------SVSSLLVVSLHLENESYGEILQP-DYGSVDVL 1103
Query: 276 EPSLCVLMDCI 286
SL VL DC+
Sbjct: 1104SISLWVLRDCL 1136
Score = 29.3 bits (64), Expect(2) = 1e-21
Identities = 17/42 (40%), Positives = 23/42 (54%), Gaps = 4/42 (9%)
Frame = +2
Query: 4 SPPKSQQPPSNVAS----SPIILPLDLITELLSFLPVKSLLQ 41
+PP+SQ + S LP DL+ ++L LPVKSL Q
Sbjct: 329 APPRSQMVSRTTVTKSRRSNPTLPFDLVEDILYRLPVKSLAQ 454
>BQ148450 similar to GP|10177464|dbj gb|AAD21700.1~gene_id:MQB2.18~similar to
unknown protein {Arabidopsis thaliana}, partial (9%)
Length = 666
Score = 95.1 bits (235), Expect = 4e-20
Identities = 70/202 (34%), Positives = 104/202 (50%), Gaps = 16/202 (7%)
Frame = +3
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAVTQESREC 81
LP ++ E+LS LPVK L+QF+CVC WK IS FVK HL+ + N R +
Sbjct: 3 LPFEIQVEILSRLPVKYLMQFQCVCKLWKSQISKPDFVKKHLR--VSNTRHLFLLTFSKL 176
Query: 82 SMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKD-CRSVAGSCNGLLCLLGLSEDREM 140
S ++++ LS + ++ T Y LN++D S+ GSC+G+LC+ +
Sbjct: 177 SPELVIKSYPLSSVF---TEMTPTFTQLEYPLNNRDESDSMVGSCHGILCI----QCNLS 335
Query: 141 WLRFWNPATRAISDKLG--HYPADFTGGFEVAFGYDNSTDTYKV------------VYLQ 186
+ WNP+ R + KL +P + AFGYD+S+DTYKV VY
Sbjct: 336 FPVLWNPSIRKFT-KLPSFEFPQNKFINPTYAFGYDHSSDTYKVVAVFCTSNIDNGVYQL 512
Query: 187 KGMTGVFSLGDNVWRNIES-FP 207
K + V ++G N WR I++ FP
Sbjct: 513 KTLVNVHTMGTNCWRRIQTEFP 578
>BG586574
Length = 732
Score = 94.0 bits (232), Expect = 9e-20
Identities = 63/176 (35%), Positives = 91/176 (50%), Gaps = 7/176 (3%)
Frame = +1
Query: 218 LRDSLNWLGLRSYVDDCDDYDCEYITSIEQFMIVALDLRTETCKELLLP-------RGFD 270
L +LNWLG ++ D D Q +I++LDL TET + L P
Sbjct: 1 LNSTLNWLG---FIQDGD--------LAPQLVIISLDLGTETYTQFLPPPISLDLSHVLQ 147
Query: 271 EVPCYEPSLCVLMDCICFSHLVKKTHLVIWKMMDYGDDDSWTQLLEINLQILKKIDEWSA 330
+V +P + +LMD +CF H + +T VIWKM +GD+ SW QLL+I+ L K++
Sbjct: 148 KVSHAKPGVSMLMDSLCFYHDLNETDFVIWKMTKFGDEKSWAQLLKISYHKL-KMNLKPG 324
Query: 331 WVPLHLSKNYDTLILGSTLEHEFAVYNLRDGSIEKTRITNGESSWIYINDYVESLV 386
+L N DTL+ + +YN R+ + KTR+ N + W IN YVESLV
Sbjct: 325 ISMFNLYVNGDTLVFVDDQKERAILYNWRNNRVVKTRV-NKKICWFSINHYVESLV 489
>CA920710 similar to PIR|S64314|S643 probable membrane protein YGR023w -
yeast (Saccharomyces cerevisiae), partial (7%)
Length = 774
Score = 92.8 bits (229), Expect = 2e-19
Identities = 52/123 (42%), Positives = 78/123 (63%), Gaps = 9/123 (7%)
Frame = -2
Query: 273 PCYEPSLCVLMDCICFSHLVKKTHLVIWKMMDYGDDDSWTQLLEI---NLQIL------K 323
PC EP L VL+DC+CF H + KT VIW+M ++G +SWT+L +I +LQ+L +
Sbjct: 773 PCAEPYLRVLLDCLCFLHDLGKTEFVIWQMKEFGVRESWTRLFKIPYVDLQMLNLPIDVQ 594
Query: 324 KIDEWSAWVPLHLSKNYDTLILGSTLEHEFAVYNLRDGSIEKTRITNGESSWIYINDYVE 383
++E+ +PL++SKN D +IL + + +YN RD +++ RI+N E W DYVE
Sbjct: 593 YLNEY-PMLPLYISKNGDKVILTNEKDDATIIYNKRDKRVDRARISN-EIHWFSAMDYVE 420
Query: 384 SLV 386
SLV
Sbjct: 419 SLV 411
>TC81493 weakly similar to PIR|F86249|F86249 protein F25C20.23 [imported] -
Arabidopsis thaliana, partial (9%)
Length = 728
Score = 85.9 bits (211), Expect = 2e-17
Identities = 73/268 (27%), Positives = 119/268 (44%), Gaps = 23/268 (8%)
Frame = +1
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAVTQESREC 81
LP ++ TE+LS +P K LL+ R C W+ LI + F+ LHL +S ++ + + Q SR
Sbjct: 16 LPTEVTTEILSRVPAKPLLRLRSTCKWWRNLIDSTDFIFLHLSKS-RDSVIILRQHSRLY 192
Query: 82 SMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAGSCNGLLCLLGLSEDREMW 141
+D+ S+ + E + P +++ V GSCNGLLC+ +++D
Sbjct: 193 ELDL----NSMDRVKE--------LDHPLMCYSNRI--KVLGSCNGLLCICNIADD---- 318
Query: 142 LRFWNPATR----AISDKL-------GHYPADFTGGFEVAFGYDNSTDTYKVVYLQK--- 187
+ FWNP R S+ L + FGYD++TD YK+V +
Sbjct: 319 IAFWNPTIRKHRIIPSEPLIRKETNENNTITTLLAAHVYGFGYDSATDDYKLVSISNFVD 498
Query: 188 -------GMTGVFSLGDNVWRNIESFPLGYYLDN--RVHLRDSLNWLGLRSYVDDCDDYD 238
++++G +VW + P V + +L+W+ R+ D D
Sbjct: 499 LHNRSYDSHVTIYTMGSDVWMPLPGVPYALCCARPMGVFVSGALHWVVPRALEPDSRD-- 672
Query: 239 CEYITSIEQFMIVALDLRTETCKELLLP 266
+IVA DLR E +E+ LP
Sbjct: 673 ----------LIVAFDLRFEVFREVALP 726
>AW690316
Length = 606
Score = 85.5 bits (210), Expect = 3e-17
Identities = 54/130 (41%), Positives = 71/130 (54%), Gaps = 6/130 (4%)
Frame = +2
Query: 5 PPKSQQPPSNVASSPIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQ 64
PP S PS V I LP D+I E L+FL VK L++ +CVC SW +ISD F K HL+
Sbjct: 185 PPPSAADPSAVV---IFLPDDVIIEFLTFLEVKDLIRMKCVCKSWNTIISDPIFAKTHLK 355
Query: 65 RS--IQNPRLAVTQESRECSMDVIVSPTSLSHLLENPSKPTT-LTNDP---YYSLNDKDC 118
+ + P LA + E S D P +S LL+ K ++ LT+D YY N K+
Sbjct: 356 KKKRTKKPHLAFLSDKSEGSGDCRAVP--ISRLLQEMKKSSSNLTHDDLPYYYRFNYKNY 529
Query: 119 RSVAGSCNGL 128
+ GSCNGL
Sbjct: 530 SDIVGSCNGL 559
>TC92509 weakly similar to PIR|T49129|T49129 hypothetical protein F26G5.80 -
Arabidopsis thaliana, partial (8%)
Length = 858
Score = 83.2 bits (204), Expect = 2e-16
Identities = 69/227 (30%), Positives = 102/227 (44%), Gaps = 12/227 (5%)
Frame = +3
Query: 11 PPSNVASSPIILPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNP 70
P S A LP +L+ E+L LPVK L Q RC+C S+ LISD F K HL S
Sbjct: 141 PQSRHAPPLPTLPFELVAEILCRLPVKLLXQLRCLCKSFNSLISDPKFAKKHLHSSTTPH 320
Query: 71 RLAVTQESRECSMDVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAG--SCNGL 128
L + + +IVSP + +L + P T Y + ++ S SC+G+
Sbjct: 321 HLILRSNNGSGRFALIVSP--IQSVLSTSTVPVPQTQLTYPTCLTEEFASPYEWCSCDGI 494
Query: 129 LCLLGLSEDREMWLRFWN-----PATRAISDKLGHYPADFTGGFEVAFGYDNSTDTYKVV 183
+CL +W F N P + IS L P+ FGYD D YKV
Sbjct: 495 ICLTTDYSSAVLWNPFINKFKTLPPLKYIS--LKRSPSCL-----FTFGYDPFADNYKVF 653
Query: 184 YL----QKGMTGVFSLGDNVWRNIESFPLGYYL-DNRVHLRDSLNWL 225
+ ++ V ++G + WR IE FP ++ D+ + + ++WL
Sbjct: 654 AITFCVKRTTVEVHTMGTSSWRRIEDFPSWSFIPDSGIFVAGYVHWL 794
>TC81783 similar to GP|9294656|dbj|BAB03005.1
gb|AAF30317.1~gene_id:F14O13.6~similar to unknown
protein {Arabidopsis thaliana}, partial (6%)
Length = 1225
Score = 56.2 bits (134), Expect(2) = 7e-16
Identities = 28/59 (47%), Positives = 37/59 (62%)
Frame = +2
Query: 22 LPLDLITELLSFLPVKSLLQFRCVCMSWKILISDSFFVKLHLQRSIQNPRLAVTQESRE 80
LP DL+ E+L LPVK L+Q RC+C + LISD F K H Q S ++ RL VT ++
Sbjct: 134 LPFDLLPEILCRLPVKLLIQLRCLCKFFNSLISDPNFAKKHFQFSAKHHRLMVTSRKQQ 310
Score = 45.1 bits (105), Expect(2) = 7e-16
Identities = 36/131 (27%), Positives = 57/131 (43%), Gaps = 6/131 (4%)
Frame = +3
Query: 84 DVIVSPTSLSHLLENPSKPTTLTNDPYYSLNDKDCRSVAG----SCNGLLCLLGLSEDRE 139
D ++ + + + + T DP ++L ++ + SC+G+ C E
Sbjct: 315 DFVLHDSPIPSVFSTSTSVTQTQLDPPFTLARRNYVNATAIAMCSCDGIFC----GELNL 482
Query: 140 MWLRFWNPATRAISDKLGHYPADFTGG-FEVAFGYDNSTDTYKVVYL-QKGMTGVFSLGD 197
+ WNP+ R L Y G F ++FGYD+ D YKVV + K V +LG
Sbjct: 483 GYCFVWNPSIRKFK-LLPPYKNPLEGDPFSISFGYDHFIDNYKVVAISSKYEVFVNALGT 659
Query: 198 NVWRNIESFPL 208
+ WR I PL
Sbjct: 660 DYWRRIREHPL 692
Score = 33.9 bits (76), Expect = 0.11
Identities = 22/90 (24%), Positives = 47/90 (51%), Gaps = 4/90 (4%)
Frame = +2
Query: 278 SLCVLMDCICFSHLVKKTHLVIWKMMDYGDDDSWTQLLEINLQILKKIDEWSAWVPLHLS 337
S+ +L DC+C + +W M +YG+ +SWT+L + ++ + A+ ++
Sbjct: 866 SVLMLRDCLCV-YEASDLFFNVWIMKEYGNQESWTKLYSVPKMQHRR---FKAYTISYIY 1033
Query: 338 KNYDTLILG----STLEHEFAVYNLRDGSI 363
++ D L+LG + + AVY+ + G++
Sbjct: 1034ED-DQLLLGIVDMQSSNTKLAVYDSKTGTL 1120
>TC91177 similar to GP|9294656|dbj|BAB03005.1
gb|AAF30317.1~gene_id:F14O13.6~similar to unknown
protein {Arabidopsis thaliana}, partial (9%)
Length = 1298
Score = 80.5 bits (197), Expect = 1e-15
Identities = 96/362 (26%), Positives = 137/362 (37%), Gaps = 46/362 (12%)
Frame = +2
Query: 42 FRCVCMSWKILISDSFFVKLHLQRSI---QNPRLAVTQESRECSMDVIVS--------PT 90
F+CVC W LIS F H Q + N + +T + S+D+ +S T
Sbjct: 74 FKCVCKLWLSLISQPHFANSHFQLTTATHTNRIMLITPYLQSLSIDLELSLNDDSAVYTT 253
Query: 91 SLSHLLENPSKPTTLTNDPYYSLNDKDCRSVA-----------GSCNGLLCLLGLSEDRE 139
+S L++ ++ YYS + D ++ GSC G + L S
Sbjct: 254 DISFLID---------DEDYYSSSSSDMDDLSPPKSFFILDFKGSCRGFILLNCYSS--- 397
Query: 140 MWLRFWNPATRAISDKLGHYPADFT-------GGFEVAFGYDNSTDTYKVVYL------- 185
L WNP+T H FT + FGYD STD Y V+ +
Sbjct: 398 --LCIWNPSTGF------HKRIPFTTIDSNPDANYFYGFGYDESTDDYLVISMSYEPSPS 553
Query: 186 QKGM---TGVFSLGDNVWRNIESFPLGYYLDNR-VHLRDSL-----NWLGLRSYVDDCDD 236
GM G+FSL NVW +E L Y N ++L +SL +WL R+
Sbjct: 554 SDGMLSHLGIFSLRANVWTRVEGGNLLLYSQNSLLNLVESLSNGAIHWLAFRN------- 712
Query: 237 YDCEYITSIEQFMIVALDLRTETCKELLLPRGFDEVPCYEPSLCVLMDCICFSHLV-KKT 295
I +IVA L EL LP P L V C+ H++ +
Sbjct: 713 -------DISMPVIVAFHLMERKLLELRLPNEIINGPSRAYDLWVYRGCLALWHILPDRV 871
Query: 296 HLVIWKMMDYGDDDSWTQLLEINLQILKKIDEWSAWVPLHLSKNYDTLILGSTLEHEFAV 355
IW M Y SWT+ L ++ W P + +K+ D I+G + A
Sbjct: 872 TFQIWVMEKYNVQSSWTKTLVLSFDGNPAHSFW----PKYYTKSGD--IVGRNMRCALAK 1033
Query: 356 YN 357
YN
Sbjct: 1034YN 1039
>AW257228 GP|17342711|gb 1-aminocyclopropanecarboxylic acid oxidase {Medicago
truncatula}, partial (11%)
Length = 702
Score = 78.6 bits (192), Expect = 4e-15
Identities = 48/100 (48%), Positives = 61/100 (61%), Gaps = 8/100 (8%)
Frame = +1
Query: 8 SQQPPSNVASSPIILPLD-LITELLSFLPVKSLLQFRCVCMSWKILISDS-FFVKLHLQR 65
S PP + +P L LD LI E+LS LPVK+L+QF+CVC SWK LISD F K HL R
Sbjct: 301 SDPPPHSNFGAPTSLLLDELIVEILSRLPVKTLMQFKCVCKSWKTLISDDPVFAKFHLHR 480
Query: 66 SIQNPRLA------VTQESRECSMDVIVSPTSLSHLLENP 99
S +N LA +T++ +CS V P ++HLLE P
Sbjct: 481 SPRNTHLAILSDRSITEDETDCS----VVPFPVTHLLEAP 588
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.139 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,868,469
Number of Sequences: 36976
Number of extensions: 269335
Number of successful extensions: 1630
Number of sequences better than 10.0: 123
Number of HSP's better than 10.0 without gapping: 1534
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1557
length of query: 389
length of database: 9,014,727
effective HSP length: 98
effective length of query: 291
effective length of database: 5,391,079
effective search space: 1568803989
effective search space used: 1568803989
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC147202.5


