

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147179.9 + phase: 1 /partial
(375 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
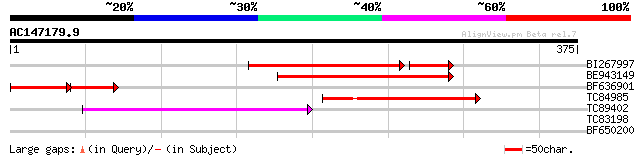
BI267997 similar to GP|13568392|em translation releasing factor2... 203 1e-57
BE943149 similar to GP|12321742|gb peptide chain release factor ... 120 9e-28
BF636901 similar to GP|13568392|emb translation releasing factor... 80 2e-27
TC84985 similar to GP|21928033|gb|AAM78045.1 AT3g62910/T20O10_10... 82 3e-16
TC89402 similar to GP|21928033|gb|AAM78045.1 AT3g62910/T20O10_10... 71 6e-13
TC83198 weakly similar to GP|13430502|gb|AAK25873.1 unknown prot... 38 0.007
BF650200 similar to GP|15215822|gb AT5g15330/F8M21_220 {Arabidop... 29 3.4
>BI267997 similar to GP|13568392|em translation releasing factor2
{Arabidopsis thaliana}, partial (28%)
Length = 409
Score = 203 bits (517), Expect(2) = 1e-57
Identities = 103/103 (100%), Positives = 103/103 (100%)
Frame = +2
Query: 159 RYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSG 218
RYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSG
Sbjct: 2 RYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSG 181
Query: 219 VEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAV 261
VEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAV
Sbjct: 182 VEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAV 310
Score = 38.1 bits (87), Expect(2) = 1e-57
Identities = 21/30 (70%), Positives = 24/30 (80%), Gaps = 1/30 (3%)
Frame = +1
Query: 265 HIPTGVTL-RCTEERSQLANKIKALSRLKA 293
HI V+L RCTEERSQLANKIKALS ++
Sbjct: 319 HISQLVSLFRCTEERSQLANKIKALSSFES 408
Score = 28.5 bits (62), Expect = 4.4
Identities = 13/26 (50%), Positives = 18/26 (69%)
Frame = +3
Query: 247 GGKGGQNVNKVETAVRITHIPTGVTL 272
G K + + +++ RITHIPTGVTL
Sbjct: 267 GEKEVRMLTRLKLPFRITHIPTGVTL 344
>BE943149 similar to GP|12321742|gb peptide chain release factor 2 putative
{Arabidopsis thaliana}, partial (22%)
Length = 411
Score = 120 bits (301), Expect = 9e-28
Identities = 62/116 (53%), Positives = 80/116 (68%)
Frame = +2
Query: 178 ATIEVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEE 237
A I+V+G +A+GY E G HR+VR SPF+S R TSF+ V V+P L +ES +V+I E
Sbjct: 62 AGIKVDGEFAFGYAKAEIGVHRLVRISPFDSNKRRHTSFAAVAVIPNLGDESSSVQINES 241
Query: 238 DLEISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKA 293
DL I RA G GGQ+VN E+AV+ITHIPTGVT C ERSQ NK A++ L++
Sbjct: 242 DLRIERFRASGAGGQHVNTTESAVKITHIPTGVTATCQNERSQHQNKASAMAVLQS 409
>BF636901 similar to GP|13568392|emb translation releasing factor2
{Arabidopsis thaliana}, partial (12%)
Length = 557
Score = 79.7 bits (195), Expect(2) = 2e-27
Identities = 41/41 (100%), Positives = 41/41 (100%)
Frame = +3
Query: 1 FYSLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCS 41
FYSLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCS
Sbjct: 339 FYSLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCS 461
Score = 60.5 bits (145), Expect(2) = 2e-27
Identities = 28/32 (87%), Positives = 30/32 (93%)
Frame = +1
Query: 41 SSFWDDRAKAQQTLSTLADVKEKIKLLNDYKT 72
+SFWDDRAKAQQTLSTLADVKEKIK +DYKT
Sbjct: 460 ASFWDDRAKAQQTLSTLADVKEKIKFAHDYKT 555
>TC84985 similar to GP|21928033|gb|AAM78045.1 AT3g62910/T20O10_10
{Arabidopsis thaliana}, partial (41%)
Length = 808
Score = 82.0 bits (201), Expect = 3e-16
Identities = 43/104 (41%), Positives = 66/104 (63%)
Frame = +1
Query: 208 SKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITHIP 267
++G TS + V +MP + E + V I +D+E++ +R+GG GGQNVNKVETA+ + H P
Sbjct: 25 TQGRVHTSTATVAIMPEVDE--VEVVIDPKDIEVTTARSGGAGGQNVNKVETAIDLVHKP 198
Query: 268 TGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIR 311
TG+ + CTE+R+Q+ NK A L+AKL I ++ + R
Sbjct: 199 TGIRIFCTEQRTQIQNKNLAFKLLRAKLYEIKVREQQESIRNQR 330
>TC89402 similar to GP|21928033|gb|AAM78045.1 AT3g62910/T20O10_10
{Arabidopsis thaliana}, partial (45%)
Length = 722
Score = 71.2 bits (173), Expect = 6e-13
Identities = 40/152 (26%), Positives = 75/152 (49%)
Frame = +1
Query: 49 KAQQTLSTLADVKEKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKS 108
K Q++S L V + D + +E+ + + + + + + E SL K+L++
Sbjct: 265 KLAQSVSELDVVVSLYQKFKDCEKTLEETQALAKDAGNDQDMTEMISYEIESLSKQLSEL 444
Query: 109 IDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKS 168
++ ++ L S P D+ ++ + AG GG +A WA L+RMY ++ E +K + S
Sbjct: 445 EEKLKVLLLPSDPLDERNIMLEVRAGTGGDEAGIWAGDLVRMYEKFSELNSWKHSAISCS 624
Query: 169 LGEEAGIKSATIEVEGRYAYGYLSGEKGTHRI 200
E+ G K+ +E++G Y L E HR+
Sbjct: 625 EAEKGGYKTYVMEIKGNRVYSKLKFESXVHRV 720
>TC83198 weakly similar to GP|13430502|gb|AAK25873.1 unknown protein
{Arabidopsis thaliana}, partial (2%)
Length = 609
Score = 37.7 bits (86), Expect = 0.007
Identities = 28/107 (26%), Positives = 54/107 (50%), Gaps = 3/107 (2%)
Frame = +2
Query: 6 KDVEIASQRVKEI---RESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKE 62
+DVE +V E +++ L ++EL +LEEEAS AQ+ TL D +
Sbjct: 260 RDVEGEKLKVAEASLEKQAMEWLLTQEELKRLEEEAS--------KHAQERSETLEDFRR 415
Query: 63 KIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSI 109
KLL+D ++++ ++ + + V +GL E+ + + + +S+
Sbjct: 416 VKKLLSDVRSELVSSQQSLASSRYKMQVQEGLLEQQLAELADQRESV 556
>BF650200 similar to GP|15215822|gb AT5g15330/F8M21_220 {Arabidopsis
thaliana}, partial (45%)
Length = 651
Score = 28.9 bits (63), Expect = 3.4
Identities = 26/101 (25%), Positives = 45/101 (43%)
Frame = +1
Query: 24 LQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVEDAETIVML 83
L++L QEL K + F+ D K ++ + ++KE+I+ L + +Q E +
Sbjct: 295 LRILNQELEKFND------FYVD--KEEEFVIRFQELKERIERLKEKSSQSEKYTSDCEF 450
Query: 84 TEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDK 124
+EEM + K L ++ N S F + YDK
Sbjct: 451 SEEMMDIRKDLVTIHGEMVLLKNYSSLNFAGLIKILKKYDK 573
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.131 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,423,607
Number of Sequences: 36976
Number of extensions: 88192
Number of successful extensions: 312
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 309
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 312
length of query: 375
length of database: 9,014,727
effective HSP length: 98
effective length of query: 277
effective length of database: 5,391,079
effective search space: 1493328883
effective search space used: 1493328883
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC147179.9


